Abstract
The adult mammalian heart is non-regenerative owing to the post-mitotic nature of cardiomyocytes. The neonatal mouse heart can regenerate, but only during the first week of life. Here we show that changes in the composition of the extracellular matrix during this week can affect cardiomyocyte growth and differentiation in mice. We identify agrin, a component of neonatal extracellular matrix, as required for the full regenerative capacity of neonatal mouse hearts. In vitro, recombinant agrin promotes the division of cardiomyocytes that are derived from mouse and human induced pluripotent stem cells through a mechanism that involves the disassembly of the dystrophin–glycoprotein complex, and Yap- and ERK-mediated signalling. In vivo, a single administration of agrin promotes cardiac regeneration in adult mice after myocardial infarction, although the degree of cardiomyocyte proliferation observed in this model suggests that there are additional therapeutic mechanisms. Together, our results uncover a new inducer of mammalian heart regeneration and highlight fundamental roles of the extracellular matrix in cardiac repair.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Soonpaa, M. H., Kim, K. K., Pajak, L., Franklin, M. & Field, L. J. Cardiomyocyte DNA synthesis and binucleation during murine development. Am. J. Physiol. Heart Circ. Physiol. 271, H2183–H2189 (1996)
Bergmann, O. et al. Evidence for cardiomyocyte renewal in humans. Science 324, 98–102 (2009)
Tzahor, E. & Poss, K. D. Cardiac regeneration strategies: staying young at heart. Science 356, 1035–1039 (2017)
Porrello, E. R. et al. Transient regenerative potential of the neonatal mouse heart. Science 331, 1078–1080 (2011)
Rupp, F., Payan, D. G., Magill-Solc, C., Cowan, D. M. & Scheller, R. H. Structure and expression of a rat agrin. Neuron 6, 811–823 (1991)
Glass, D. J. et al. Agrin acts via a MuSK receptor complex. Cell 85, 513–523 (1996)
Burden, S. J., Yumoto, N. & Zhang, W. The role of MuSK in synapse formation and neuromuscular disease. Cold Spring Harb. Perspect. Biol. 5, a009167 (2013)
Bowe, M. A., Deyst, K. A., Leszyk, J. D. & Fallon, J. R. Identification and purification of an agrin receptor from Torpedo postsynaptic membranes: a heteromeric complex related to the dystroglycans. Neuron 12, 1173–1180 (1994)
Henry, M. D. & Campbell, K. P. Dystroglycan: an extracellular matrix receptor linked to the cytoskeleton. Curr. Opin. Cell Biol. 8, 625–631 (1996)
Campbell, K. P. & Kahl, S. D. Association of dystrophin and an integral membrane glycoprotein. Nature 338, 259–262 (1989)
Richardson, G. D., Laval, S. & Owens, W. A. Cardiomyocyte regeneration in the mdx mouse model of nonischemic cardiomyopathy. Stem Cells Dev. 24, 1672–1679 (2015)
Morikawa, Y. et al. Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice. Sci. Signal. 8, ra41 (2015)
Kuhn, B. et al. Periostin induces proliferation of differentiated cardiomyocytes and promotes cardiac repair. Nat. Med. 13, 962–969 (2007)
Bian, W., Badie, N., Himel, H. D., IV & Bursac, N. Robust T-tubulation and maturation of cardiomyocytes using tissue-engineered epicardial mimetics. Biomaterials 35, 3819–3828 (2014)
D’Uva, G. et al. ERBB2 triggers mammalian heart regeneration by promoting cardiomyocyte dedifferentiation and proliferation. Nat. Cell Biol. 17, 627–638 (2015)
O’Meara, C. C. et al. Transcriptional reversion of cardiac myocyte fate during mammalian cardiac regeneration. Circ. Res. 116, 804–815 (2015)
Mazzon, C. et al. Agrin is required for survival and function of monocytic cells. Blood 119, 5502–5511 (2012)
Yahalom-Ronen, Y., Rajchman, D., Sarig, R., Geiger, B. & Tzahor, E. Reduced matrix rigidity promotes neonatal cardiomyocyte dedifferentiation, proliferation and clonal expansion. eLife 4, e07455 (2015)
Zhang, D. et al. Tissue-engineered cardiac patch for advanced functional maturation of human ESC-derived cardiomyocytes. Biomaterials 34, 5813–5820 (2013)
Gesemann, M. et al. Alternative splicing of agrin alters its binding to heparin, dystroglycan, and the putative agrin receptor. Neuron 16, 755–767 (1996)
Cohen, S., Zhai, B., Gygi, S. P. & Goldberg, A. L. Ubiquitylation by Trim32 causes coupled loss of desmin, Z-bands, and thin filaments in muscle atrophy. J. Cell Biol. 198, 575–589 (2012)
Cohen, S., Nathan, J. A. & Goldberg, A. L. Muscle wasting in disease: molecular mechanisms and promising therapies. Nat. Rev. Drug Discov. 14, 58–74 (2015)
Constantin, B. Dystrophin complex functions as a scaffold for signalling proteins. Biochim. Biophys. Acta 1838, 635–642 (2013)
Heallen, T. et al. Hippo signaling impedes adult heart regeneration. Development 140, 4683–4690 (2013)
Xin, M. et al. Hippo pathway effector Yap promotes cardiac regeneration. Proc. Natl Acad. Sci. USA 110, 13839–13844 (2013)
von Gise, A. et al. YAP1, the nuclear target of Hippo signaling, stimulates heart growth through cardiomyocyte proliferation but not hypertrophy. Proc. Natl Acad. Sci. USA 109, 2394–2399 (2012)
Reischauer, S., Arnaout, R., Ramadass, R. & Stainier, D. Y. R. Actin binding GFP allows 4D in vivo imaging of myofilament dynamics in the zebrafish heart and the identification of Erbb2 signaling as a remodeling factor of myofibril architecture. Circ. Res. 115, 845–856 (2014)
Liu-Chittenden, Y. et al. Genetic and pharmacological disruption of the TEAD–YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 26, 1300–1305 (2012)
Wang, C. et al. Verteporfin inhibits YAP function through up-regulating 14-3-3σ sequestering YAP in the cytoplasm. Am. J. Cancer Res. 6, 27–37 (2015)
Jackman, C. P., Carlson, A. L. & Bursac, N. Dynamic culture yields engineered myocardium with near-adult functional output. Biomaterials 111, 66–79 (2016)
Acknowledgements
This work was supported by grants to E.T. from the European Research Council, Israel Science Foundation and the Britain Israel Research and Academic Exchange (BIRAX). E.T., J.F.M. and N.B. are supported by Foundation LeDucq and NIH funding. We thank R. Burgess for providing the Agrnfl/fl mice. We thank Y. Levin, head of de Botton Institute for Protein Profiling, The Nancy and Stephen Grand Israel National Center for Personalized Medicine, for protein profiling. We thank E.T. and I.S. laboratory members, A. Navon, K. Yaniv and P Riley for fruitful discussions, O. Goresh and B. Siani for animal husbandry, and N. Akron, D. Gorelik, R. Levine, S. Goldsmith, C. Raanan and M. Osin for genotyping and histology.
Author information
Authors and Affiliations
Contributions
E.B., with help from A.G., performed most of the experiments. Y.E.M. and S.C. performed glycerol-gradient experiments. I.Y.S. and N.B. performed 3D iPSC–CM culture experiments, E.B., K.B.U., D.K., J.L. and J.F.M. performed in vivo animal experiments, O.Y. performed in vitro experiments, D.R.B. performed western blots, D.R. and R.S. performed hiPS–CM experiments, Y.U. and I.S. performed in situ zymography. E.T. supervised the project and with E.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks K. Yutzey and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 P1 and P7 ECM explants contain increased gelatinase activity.
a, b, Cell removal assessment of heart sections by DAPI or scanning electron microscopy. Scale bars, 50 μm (a) and 20 μm (b). c, A schematic diagram of the in situ zymography assay. d, Immunofluorescence evaluation of Col1, Col4 and gelatin degradation in response to P1 and P7 ECM. e, Quantification of the in situ zymography assay. n = 2 samples.
Extended Data Figure 2 Mass spectrometry results and validation.
a, Venn diagram representing the LC–MS results. b, qPCR analysis of genes obtained from the LC–MS in P1 and P7 whole-heart lysates. n = 3 P1 hearts and 3 P7 hearts.
Extended Data Figure 3 P1 and P8 cardiac cell separation.
qPCR of mRNA from P1 and P8 heart lysates. qPCR analysis of six cell populations (fibroblasts, non-fibroblasts, cardiomyocytes, non-cardiomyocytes, endothelial cells, non-endothelial cells) for CD90 (also known as Thy1, a fibroblast marker), Myh6 (a cardiomyocyte marker) and Pecam1 (Pecam, an endothelial cell marker) (for P1, n = 4 cardiomyocyte, 4 non-cardiomyocyte, 4 fibroblast, 4 non-fibroblasts, 7 endothelial cell, 7 non-endothelial cell samples and for P8 n = 2 cardiomyocyte and 2 non-cardiomyocyte samples). Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001; statistical significance was calculated using a one-tailed t-test.
Extended Data Figure 4 Agrin induces cardiomyocyte proliferation in vitro.
a–d, Immunofluorescence evaluation of P1 and P7 cardiomyocytes (cTnT+ or Myh6CretdTomato+, see Methods) cell number (a), cell-cycle activity (Ki67; b), mitosis (pH3; c) or cytokinesis (Aurkb, also known as Aim1; d) in response to agrin dosage in vitro. White arrowheads indicate proliferating cells. Analysis of five individual wells per treatment (a); n = 1,855 P1 and 1,328 P7 cardiomyocytes pooled from the analysis of three P1 and four P7 samples (b); n = 11,450 P1 and 13,469 P7 cardiomyocytes pooled from the analysis of four P1 and six P7 samples (c); n = 7,971 P1 and 3,856 P7 cardiomyocytes pooled from the analysis of three P1 and three P7 samples (d). Scale bars, 30 μm (b) and 100 μm (c). Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001; statistical significance was calculated using an ANOVA followed by Dunnett’s post hoc test relative to the control group(a–c) or using a one-tailed t-test (d).
Extended Data Figure 5 Agrin-cKO cardiac characterization.
a, Tnni3:Tnni1 protein ratio in P8 agrin-cKO and wild-type mice. n = 4 wild-type and 7 cKO mice. b, Immunofluorescence analysis and Pearsons’s correlation coefficent analysis of P14 T-tubules by Cav3 colocalizion with α-actinin labelled z-lines. n = 5 wild-type and 6 cKO samples. c, Immunofluorescence analysis of WGA membrane staining in P1 wild-type and agrin-cKO depicting changes in cell size. n = 2 wild-type and 3 cKO samples. d, qPCR analysis of pathological hypertrophic marker Acta1 in P1 wild-type and agrin-cKO heart lysates. n = 21 wild-type and 26 cKO samples. e, f, Serial echocardiographic measurements of ejection fraction (EF) and wall thickness of 1-, 3- and 14-month-old agrin cKO and wild-type mice. n = 8 mice per group at 1 month, 4 wild-type and 5 cKO at 3 months and 4 wild-type and 3 cKO at 14 months. g, Heart to body weight ratio of 1-, 3- and 14-month-old agrin-cKO and wild-type mice. n = 9 wild-type and 4 cKO at 1 month, 4 wild-type and 5 cKO at 3 months and 6 wild-type and 5 cKO at 14 months. h, Histological sections of 1- and 3-month-old wild-type and agrin-cKO stained with Masson’s trichrome. i, Scheme of P1 heart resection. j, Representative images of agrin-cKO and wild-type mice 28 days after resection at P1. k, Functional cardiac recovery measurement (cardiac output), 28 days after resection. n = 5 wild-type and 3 cKO mice (j, k). Scale bars, 20 μm (b), 10 μm (c) and 1 mm (h). Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01; Statistical significance was calculated using a one-tailed t-test.
Extended Data Figure 6 Agrin induces cardiac regeneration in juvenile mice.
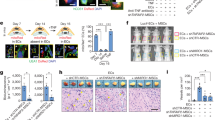
a, A schematic diagram depicting LAD ligation in juvenile and adult mice. b, c, In vivo evaluation of cardiomyocytes cell-cycle re-entry by Ki67 (b) or Aurkb (c) in heart sections seven days after myocardial infarction. n = 1,934 cardiomyocytes pooled from the analysis of three PBS-treated mice and three agrin-treated mice (b); n = 1,450 cardiomyocytes pooled from the analysis of three PBS- and six agrin-treated samples (c). d, Scar quantification based on Masson’s trichrome staining of heart sections of PBS- and agrin-treated juvenile mice. Representative pictures are provided. n = 4 PBS- and 8 agrin-treated mice. e–g, Serial echocardiographic measurements of ejection fraction (EF), fractional shortening (FS) and wall thickness of uninjured and injured PBS- and agrin-treated juvenile mice following myocardial infarction, according to the schema in a. n = 3 uninjured, 3 PBS- and 3 agrin-treated mice at day 13; and n = 4 uninjured, 5 PBS- and 9 agrin-treated mice at day 43. Scale bars, 10 μm (b), 20 μm (c) and 1 mm (d). Data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001; statistical significance was calculated using a one-tailed t-test(b–d, g) or ANOVA followed by Dunnett’s post hoc test relative to the control group (e, f).
Extended Data Figure 7 Pharmacokinetics of agrin injection in injured adult hearts.
Mice were subjected to LAD ligation (as shown in Fig. 4a), and received either PBS or agrin treatment. Hearts were collected at respective time points after treatment, and were subjected to protein extraction and anti-His-tag immunopreciptation using Ni-NTA resin. a, Representative western blot of agrin from relevant immunoprecipitation samples. Anti-tubulin western blot of respective total extracts, as immunoprecipitation loading control is shown at the bottom. b, The kinetics of recombinant agrin up to 96 h after treatment. The amount of agrin is presented as a percentage of the first time point (t = 0), and normalized according to tubulin. n = 3 repeats.
Extended Data Figure 8 Dag1 expression in P1 and P7 whole hearts and cardiac cell separation.
a, qPCR of Dag1 mRNA from P1 and P7 heart lysates. n = 5 P1 and 3 P7 samples. b, qPCR analysis of Dag1 mRNA in distinct cardiac populations from P1 mice (fibroblasts, non-fibroblasts, cardiomyocytes, non-cardiomyocytes, endothelial cells, non-endothelial cells). n = 4 cardiomyocyte, 4 non-cardiomyocyte, 4 fibroblast, 4 non-fibroblast and 7 endothelial and 7 non-endothelial cell samples. c, Serial immunofluorecence counting of Myh6-lineage-derived tdTomato-labelled cardiomyocytes treated with ouabain and digoxin which inhibit Na+K+ pumps. Quantification was made using acuman (see Methods). n = 31 control, 16 agrin-, 11 ouabain- and 11 digoxin-treated individual wells per treatment. d, quantification of cardiomyocyte sarcomeric status by cTnT and Myh6-linage immunofluorescence analysis of P7 agrin-treated cardiomyocytes cultured in vitro for three days. n = 2,222 cardiomyocytes pooled from the analysis of three samples per group. Representative images of cardiomyocytes with intact, partially disassembled and severely disassembled sarcomeres are provided. Scale bars, 40 μm. Data are presented as mean ± s.e.m. *P < 0.05, ***P < 0.001; statistical significance was calculated using a one-tailed t-test (a, b, d) or ANOVA followed by Dunnett’s post hoc test relative to control group (c).
Extended Data Figure 9 A model for the agrin–DGC–Yap signalling axis in cardiomyocyte maturation and cardiac regeneration.
Agrin triggers mild cardiomyocyte dedifferentiation and proliferation by modulation of DGC integrity and signalling. During neonatal stages agrin suppresses the maturation of the DGC. At P7, agrin levels are reduced and ECM–DGC interaction through Dag1 promotes cardiomyocyte differentiation and maturation. Yap is tethered to the DGC upon cardiomyocyte maturation while upon agrin treatment, it translocates to the nucleus to facilitate cardiomyocyte cell-cycle re-entry.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3 and the uncropped western blots. (PDF 1635 kb)
Rights and permissions
About this article
Cite this article
Bassat, E., Mutlak, Y., Genzelinakh, A. et al. The extracellular matrix protein agrin promotes heart regeneration in mice. Nature 547, 179–184 (2017). https://doi.org/10.1038/nature22978
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22978
This article is cited by
-
Protein glycosylation in cardiovascular health and disease
Nature Reviews Cardiology (2024)
-
Animal models to study cardiac regeneration
Nature Reviews Cardiology (2024)
-
N-Acetyltransferase 10 represses Uqcr11 and Uqcrb independently of ac4C modification to promote heart regeneration
Nature Communications (2024)
-
Deficiency of skeletal muscle Agrin contributes to the pathogenesis of age-related sarcopenia in mice
Cell Death & Disease (2024)
-
Targeting cardiomyocyte cell cycle regulation in heart failure
Basic Research in Cardiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.