Abstract
Rotavirus, a leading cause of severe gastroenteritis and diarrhoea in young children, accounts for around 215,000 deaths annually worldwide1. Rotavirus specifically infects the intestinal epithelial cells in the host small intestine and has evolved strategies to antagonize interferon and NF-κB signalling2,3,4,5, raising the question as to whether other host factors participate in antiviral responses in intestinal mucosa. The mechanism by which enteric viruses are sensed and restricted in vivo, especially by NOD-like receptor (NLR) inflammasomes, is largely unknown. Here we uncover and mechanistically characterize the NLR Nlrp9b that is specifically expressed in intestinal epithelial cells and restricts rotavirus infection. Our data show that, via RNA helicase Dhx9, Nlrp9b recognizes short double-stranded RNA stretches and forms inflammasome complexes with the adaptor proteins Asc and caspase-1 to promote the maturation of interleukin (Il)-18 and gasdermin D (Gsdmd)-induced pyroptosis. Conditional depletion of Nlrp9b or other inflammasome components in the intestine in vivo resulted in enhanced susceptibility of mice to rotavirus replication. Our study highlights an important innate immune signalling pathway that functions in intestinal epithelial cells and may present useful targets in the modulation of host defences against viral pathogens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Tate, J. E. et al. Effectiveness of pentavalent rotavirus vaccine under conditions of routine use in Rwanda. Clin. Infect. Dis. 62 (Suppl 2), S208–S212 (2016)
Sen, A., Rott, L., Phan, N., Mukherjee, G. & Greenberg, H. B. Rotavirus NSP1 protein inhibits interferon-mediated STAT1 activation. J. Virol. 88, 41–53 (2014)
Ding, S. et al. Comparative proteomics reveals strain-specific β-TrCP degradation via rotavirus NSP1 hijacking a host cullin-3–Rbx1 complex. PLoS Pathog. 12, e1005929 (2016)
Graff, J. W., Ettayebi, K. & Hardy, M. E. Rotavirus NSP1 inhibits NFkappaB activation by inducing proteasome-dependent degradation of beta-TrCP: a novel mechanism of IFN antagonism. PLoS Pathog. 5, e1000280 (2009)
Barro, M. & Patton, J. T. Rotavirus nonstructural protein 1 subverts innate immune response by inducing degradation of IFN regulatory factor 3. Proc. Natl Acad. Sci. USA 102, 4114–4119 (2005)
Uchiyama, R., Chassaing, B., Zhang, B. & Gewirtz, A. T. MyD88-mediated TLR signaling protects against acute rotavirus infection while inflammasome cytokines direct Ab response. Innate Immun. 21, 416–428 (2014)
Zhang, B. et al. Viral infection. Prevention and cure of rotavirus infection via TLR5/NLRC4-mediated production of IL-22 and IL-18. Science 346, 861–865 (2014)
Hirota, S. A. et al. NLRP3 inflammasome plays a key role in the regulation of intestinal homeostasis. Inflamm. Bowel Dis. 17, 1359–1372 (2011)
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011)
Nordlander, S., Pott, J. & Maloy, K. J. NLRC4 expression in intestinal epithelial cells mediates protection against an enteric pathogen. Mucosal Immunol. 7, 775–785 (2014)
Hu, B. et al. The DNA-sensing AIM2 inflammasome controls radiation-induced cell death and tissue injury. Science 354, 765–768 (2016)
Feng, N., Yasukawa, L. L., Sen, A. & Greenberg, H. B. Permissive replication of homologous murine rotavirus in the mouse intestine is primarily regulated by VP4 and NSP1. J. Virol. 87, 8307–8316 (2013)
Levy, M. et al. Microbiota-modulated metabolites shape the intestinal microenvironment by regulating NLRP6 inflammasome signaling. Cell 163, 1428–1443 (2015)
Miao, E. A. et al. Cytoplasmic flagellin activates caspase-1 and secretion of interleukin 1β via Ipaf. Nat. Immunol. 7, 569–575 (2006)
Zhao, Y. et al. The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus. Nature 477, 596–600 (2011)
Wu, C., Macleod, I. & Su, A. I. BioGPS and MyGene.info: organizing online, gene-centric information. Nucleic Acids Res. 41, D561–D565 (2013)
Halasz, P., Holloway, G. & Coulson, B. S. Death mechanisms in epithelial cells following rotavirus infection, exposure to inactivated rotavirus or genome transfection. J. Gen. Virol. 91, 2007–2018 (2010)
Kanneganti, T. D. et al. Critical role for Cryopyrin/Nalp3 in activation of caspase-1 in response to viral infection and double-stranded RNA. J. Biol. Chem. 281, 36560–36568 (2006)
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015)
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015)
Lin, J. D. et al. Distinct roles of type I and type III interferons in intestinal immunity to homologous and heterologous rotavirus infections. PLoS Pathog. 12, e1005600 (2016)
Deal, E. M., Jaimes, M. C., Crawford, S. E., Estes, M. K. & Greenberg, H. B. Rotavirus structural proteins and dsRNA are required for the human primary plasmacytoid dendritic cell IFNα response. PLoS Pathog. 6, e1000931 (2010)
Silvestri, L. S., Taraporewala, Z. F. & Patton, J. T. Rotavirus replication: plus-sense templates for double-stranded RNA synthesis are made in viroplasms. J. Virol. 78, 7763–7774 (2004)
Rojas, M., Arias, C. F. & López, S. Protein kinase R is responsible for the phosphorylation of eIF2α in rotavirus infection. J. Virol. 84, 10457–10466 (2010)
Wang, P. et al. Nlrp6 regulates intestinal antiviral innate immunity. Science 350, 826–830 (2015)
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006)
Kato, H. et al. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J. Exp. Med. 205, 1601–1610 (2008)
Pott, J. et al. Age-dependent TLR3 expression of the intestinal epithelium contributes to rotavirus susceptibility. PLoS Pathog. 8, e1002670 (2012)
Eydelloth, R. S., Vonderfecht, S. L., Sheridan, J. F., Enders, L. D. & Yolken, R. H. Kinetics of viral replication and local and systemic immune responses in experimental rotavirus infection. J. Virol. 50, 947–950 (1984)
Sheridan, J. F., Eydelloth, R. S., Vonderfecht, S. L. & Aurelian, L. Virus-specific immunity in neonatal and adult mouse rotavirus infection. Infect. Immun. 39, 917–927 (1983)
Sutterwala, F. S. et al. Critical role for NALP3/CIAS1/Cryopyrin in innate and adaptive immunity through its regulation of caspase-1. Immunity 24, 317–327 (2006)
Case, C. L. et al. Caspase-11 stimulates rapid flagellin-independent pyroptosis in response to Legionella pneumophila. Proc. Natl Acad. Sci. USA 110, 1851–1856 (2013)
Wlodarska, M. et al. NLRP6 inflammasome orchestrates the colonic host-microbial interface by regulating goblet cell mucus secretion. Cell 156, 1045–1059 (2014)
Lara-Tejero, M. et al. Role of the caspase-1 inflammasome in Salmonella typhimurium pathogenesis. J. Exp. Med. 203, 1407–1412 (2006)
Rathinam, V. A. et al. The AIM2 inflammasome is essential for host defense against cytosolic bacteria and DNA viruses. Nat. Immunol. 11, 395–402 (2010)
Feng, N. et al. Role of interferon in homologous and heterologous rotavirus infection in the intestines and extraintestinal organs of suckling mice. J. Virol. 82, 7578–7590 (2008)
Burns, J. W. et al. Analyses of homologous rotavirus infection in the mouse model. Virology 207, 143–153 (1995)
Feng, N., Burns, J. W., Bracy, L. & Greenberg, H. B. Comparison of mucosal and systemic humoral immune responses and subsequent protection in mice orally inoculated with a homologous or a heterologous rotavirus. J. Virol. 68, 7766–7773 (1994)
Sen, A., Pruijssers, A. J., Dermody, T. S., García-Sastre, A. & Greenberg, H. B. The early interferon response to rotavirus is regulated by PKR and depends on MAVS/IPS-1, RIG-I, MDA-5, and IRF3. J. Virol. 85, 3717–3732 (2011)
Ball, J. M., Tian, P., Zeng, C. Q., Morris, A. P. & Estes, M. K. Age-dependent diarrhea induced by a rotaviral nonstructural glycoprotein. Science 272, 101–104 (1996)
Saxena, K. et al. Human intestinal enteroids: a new model to study human rotavirus infection, host restriction, and pathophysiology. J. Virol. 90, 43–56 (2015)
Acknowledgements
We would like to thank C. Jin, B. Hu, A. Solis, P. Bielecki, R. Nowarski, R. Jackson, W. Bailis, A. Rongvaux, J. Henao-Mejia, A. Williams, E. Elinav, T. Strowig, J. Alderman, C. Lieber, J. Stein, C. Hughes, L. Evangelisti, L. Borelli, P. Ranney and other members of Flavell laboratory for technical help and helpful discussion. S.Z. is supported by a fellowship from Helen Hay Whitney Foundation-Howard Hughes Medical Institute. S.D. is supported by the Walter V. and Idun Berry Postdoctoral Fellowship and an Early Career Award from the Thrasher Research Fund. P.W. is supported by R21AI103807. C.A. is supported by R01DK099097, R01DK106593 and R01AI120369. H.B.G. is supported by VA Merit Review grant I01BX000158, and NIH grants R01AI021362 and U19AI116484. R.A.F. and E.F. are investigators of the Howard Hughes Medical Institute, and are supported by U19AI089992.
Author information
Authors and Affiliations
Contributions
S.Z. wrote the manuscript, designed, performed and interpreted experiments with technical assistance from S.D. and P.H.W; S.D. and H.B.G. provided rotavirus related reagents, S.D. performed the rotavirus fecal shedding assay, qPCR and immunostaining experiments; P.H.W, W.P and Y.Z performed RNA binding experiments; Z.W. and H.C. performed infections in intestinal organoids; Y.Y. and H.Y. performed TUNEL staining; H.-B.L. and K.A.J. helped with DHX9 functional studies; G.W., X.L. and Y.Z. assisted in the characterization of Nlrp9b mice; N.P., M.Z., and J.Z. performed 16s sequencing and PcoA analysis; N.F. performed the human IEC organoid experiment; C.A. provided human gut biopsies; S.D., P.H.W., N.W.P., W.P., L.S., Y.K., J.L., C.A., E.F., H.B.G. and R.A.F. provided critical comments and suggestions; R.A.F. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks I. Goodfellow, C. Reis e Sousa and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Increase of rotavirus infection in Casp1-deficient but not NLR-deficient mice.
a, qPCR detection of viral loads in ileum by measuring different virus transcripts, non-structural protein 5 (NSP5), viral protein 6 (VP6) or non-structural protein 3 (NSP3) (mean ± s.e.m., Student’s t-test, ***P < 0.001) from wild-type and Casp1−/− suckling mice which were orally inoculated with rotavirus EW strain with diarrhoea dose 1 (DD)50. b, c, Viral loads in ileum (mean ± s.e.m., Student’s t-test, *P < 0.05) (b) or diarrhoea duration and percentage of diarrhoea (c) from wild-type and different inflammasome-forming NLRs or other receptor-knockout mice, which were orally inoculated with rotavirus EW strain with DD50. b, c, Representative of two independent experiments.
Extended Data Figure 2 Tissue-specific Nlrp9b expression and characterization of Nlrp9b-deficient mice.
a, Nlrp9b expression in different mouse tissues as per BioGPS. b, qPCR verification of Nlrp9. a–c, Expression in different mouse tissues. c, Illustration of the CRISPR–Cas9 strategy to generate Nlrp9b-deficient mice. d, qPCR validation of Nlrp9b expression in the intestine tissues from littermate wild-type and Nlrp9b−/− mice (mean ± s.e.m., Student’s t-test, ***P < 0.001). e, Haematoxylin-and-eosin staining of ileal sections from wild-type and Nlrp9b-knockout mice in steady state. f, Unweighted UniFrac PCoA analysis followed by bacterial 16S rRNA sequencing of fecal microbiota harvested from littermate wild-type mice and Nlrp9b-knockout mice separated by indicated time points. b, Representative of three independent experiments.
Extended Data Figure 3 Specific Nlrp9b expression in IECs and generation of Nlrp9b IEC conditional knockout mice.
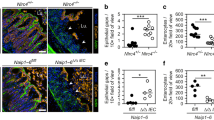
a, qPCR analysis of NLR inflammasome components in IEC fraction and IEL fraction isolated by EDTA. Vil1 is an IEC marker and Cd45 is a haematopoietic marker. b, Method to isolate IEC, IEL and LPL from intestine of Villin-Cre-tomato mice for experiment in Fig. 2f. c, qPCR analysis of NLR inflammasome components in intestinal epithelial cells (IECs), intraepithelial lymphocytes (IELs), lamina propria lymphocytes (LPLs), isolated according to the method showed in b. d, Workflow to generate Nlrp9b IEC conditional knockout mice. e, qPCR validation of Nlrp9b expression in IECs from Nlrp9b IEC conditional knockout and the littermate control (mean ± s.e.m., Student’s t-test, ***P < 0.001). SI, small intestine; LI, large intestine. a, c, Representative of three independent experiments.
Extended Data Figure 4 Rotavirus induction of cell death in the small intestine.
a, Representative cell death staining in IECs from wild-type mice orally inoculated with PBS or rotavirus. b, Summary of cell death and survival from the experiment in a (mean ± s.e.m., Student’s t-test, *P < 0.05). c, Western blots of different cell death indicators, Casp1 p10 (pyroptosis), Casp3 p17/p19 (apoptosis), and Lc3 I/II (autophagy), in the IECs from rotavirus infected ileum. a–c, Representative of two independent experiments.
Extended Data Figure 5 Establishment of intestinal organoid culture and the role of Nlrp9b in cytokine secretion.
a, Single organoid (mini-gut) culture. Crypts are isolated from the small intestine of indicated mouse strains and cultured in Matrigel with IntestiCult growth medium from Stem Cell Technology for 3 weeks. b, ELISA measurement of cytokines and interferons (IFNs) production or c. QPCR analysis of cytokine, IFNs and anti-microbial peptides in the explants of wild-type and Nlrp9b−/− mice treated with rotavirus. (mean ± s.e.m., Student’s t-test, **P < 0.01). b, c, Representative of three independent experiments.
Extended Data Figure 6 Gsdmd expression in different tissues and generation of Gsdmd deficient mice.
a, Illustration of the CRISPR–Cas9 strategy to generate Nlrp9b-deficient mice and primer design used in b. b, Genotyping of the Gsdmd mutant pups. The mice chosen for experiments shown in Fig. 3c, d are shown in red, with a deletion in one allele sequenced by primer 2, as well as mutations that result in frame shift in another allele sequenced by primer 1. c, Sequencing of red-labelled mice with primer 1. d, qPCR validation of Gsdmd expression in the intestine tissues from these five red-labelled Gsdmd mutant mice, comparing with wild-type mice and Il18−/− mice (mean ± s.e.m., Student’s t-test, ***P < 0.001). e, Gsdmd expression in different mouse tissues as per BioGPS. f, RNA sequencing data reveals the expression of gasdermin family members and housekeeping gene Hprt in the ileal epithelial cells of wild-type mice. b–d, Representative of two experiments and the data was combined in Fig. 3c and d.
Extended Data Figure 7 Rotavirus RNA distribution in the cytoplasm, and role of Nlrp9b in anti-EMCV response in the intestine.
a, Human intestinal epithelial cell line HT-29 cells were seeded into chamber slides and infected with RRV at MOI = 1 for 24 h. Infected HT-29 cells were stained sequentially with anti-dsRNA (J2 mouse monoclonal Ab, 1:60), anti-mouse Alexa-594 secondary antibody (1:200) and Alexa-488-conjugated anti-NSP2 (clone 191, mouse monoclonal Ab, 1:250). Cells were mounted with anti-fade DAPI and processed for confocal microscopy (LSM710). b, Survival curve (Log-rank (Mantel–Cox) test, **P < 0.01) of different mouse strains challenged by EMCV (105 pfu) orally. b, Representative of two independent experiments.
Extended Data Figure 8 Role of Dhx9 in Nlrp9b-mediated anti-rotavirus response.
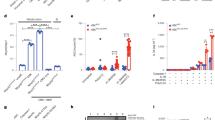
a, Biotin-labelled low molecular weight poly(I:C) were incubated with whole-cell lysates from HEK293T cells transfected with Flag–NLRP9 and indicated siRNA. b, HEK293T cells were transfected with DHX9-targeting siRNA or control siRNA for 24 h, co-transfected with Flag–NLRP9 and HA–ASC for another 24h, and infected with rotavirus human rotavirus WA strain (estimated MOI = 2) for 6 h. Cell lysates were co-immunoprecipitated with anti-Flag antibody and analysed by western blot with the indicated antibodies. c, Validation of Dhx9 conditional knockout mice. Dhx9 mRNA expression in intestinal epithelial cells from indicated mouse strains (mean ± s.e.m., Student’s t-test, **P < 0.01, ***P < 0.001). d, Cytotoxicity measured by lactate dehydrogenase (LDH) release in organoid cells from the indicated mouse strains infected with rotavirus EW strain (5 × 105 DD50, estimated MOI = 5) for 3 h (mean ± s.e.m., Student’s t-test, *P < 0.05). e, ELISA measurement of Il-18 secretion upon treating intestinal organoid cells from the indicated mouse strains with rotavirus for 3 h (mean ± s.e.m., Student’s t-test, **P < 0.01). f, qPCR detection of viral loads by measuring virus transcript VP6 in organoid cells from the indicated mouse strains infected with rotavirus for 6 h (mean ± s.e.m., Student’s t-test, *P < 0.05).
Extended Data Figure 9 Nlrp9b regulation during rotavirus infection and murine intestine.
a, Workflow to generate Flag–Nlrp9 -knock-in mice. b, Immunoprecipitation and western blots detection of Flag–Nlrp9b protein, c, qPCR analysis of Nlrp9b, Asc and rotavirus transcripts, in IECs samples from Flag–Nlrp9b-knock-in mice that were infected with 100 DD50 of rotavirus by the indicated time points. d, Expression of select innate immune receptors or adaptors were measured by qPCR in different intestine sections isolated from mice with different ages as indicated in the figure. (mean ± s.e.m., Student’s t-test compare to expression levels in day 0, *P < 0.05, **P < 0.01.) Experiments in b and c were replicated twice.
Extended Data Figure 10 Human NLRP9 expression in intestinal epithelial cells.
a, Clinical information of patients from whom the gut biopsies were collected. b, Method to isolate IEC fraction, lamina propria fraction and LPL from human gut biopsies. c, qPCR analysis of human NLRP9 inflammasome components in IEC fraction, lamina propria fraction, LPL and PBMC, which are isolated according to b. VILLIN1 is an IEC marker, and CD45 is a haematopoietic marker. d, Expression of indicated genes were measured by qPCR in human IEC organoids and select cell lines (mean ± s.e.m., Student’s t-test, ***P < 0.001).
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1, the uncropped gels. (PDF 2581 kb)
Source data
Rights and permissions
About this article
Cite this article
Zhu, S., Ding, S., Wang, P. et al. Nlrp9b inflammasome restricts rotavirus infection in intestinal epithelial cells. Nature 546, 667–670 (2017). https://doi.org/10.1038/nature22967
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22967
This article is cited by
-
NLRP inflammasomes in health and disease
Molecular Biomedicine (2024)
-
Multiomics characterization of pyroptosis in the tumor microenvironment and therapeutic relevance in metastatic melanoma
BMC Medicine (2024)
-
Biological and clinical roles of IL-18 in inflammatory diseases
Nature Reviews Rheumatology (2024)
-
The gasdermin family: emerging therapeutic targets in diseases
Signal Transduction and Targeted Therapy (2024)
-
The role of inflammasomes in human diseases and their potential as therapeutic targets
Signal Transduction and Targeted Therapy (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.