Abstract
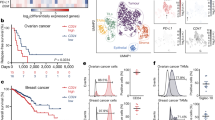
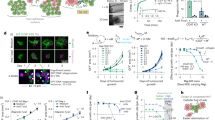
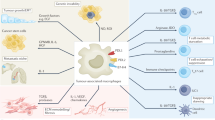
Programmed cell death protein 1 (PD-1) is an immune checkpoint receptor that is upregulated on activated T cells for the induction of immune tolerance1,2. Tumour cells frequently overexpress the ligand for PD-1, programmed cell death ligand 1 (PD-L1), facilitating their escape from the immune system3,4. Monoclonal antibodies that block the interaction between PD-1 and PD-L1, by binding to either the ligand or receptor, have shown notable clinical efficacy in patients with a variety of cancers, including melanoma, colorectal cancer, non-small-cell lung cancer and Hodgkin’s lymphoma5,6,7,8,9. Although it is well established that PD-1–PD-L1 blockade activates T cells, little is known about the role that this pathway may have in tumour-associated macrophages (TAMs). Here we show that both mouse and human TAMs express PD-1. TAM PD-1 expression increases over time in mouse models of cancer and with increasing disease stage in primary human cancers. TAM PD-1 expression correlates negatively with phagocytic potency against tumour cells, and blockade of PD-1–PD-L1 in vivo increases macrophage phagocytosis, reduces tumour growth and lengthens the survival of mice in mouse models of cancer in a macrophage-dependent fashion. This suggests that PD-1–PD-L1 therapies may also function through a direct effect on macrophages, with substantial implications for the treatment of cancer with these agents.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Nishimura, H., Nose, M., Hiai, H., Minato, N. & Honjo, T. Development of lupus-like autoimmune diseases by disruption of the PD-1 gene encoding an ITIM motif-carrying immunoreceptor. Immunity 11, 141–151 (1999)
Freeman, G. J. et al. Engagement of the PD-1 immunoinhibitory receptor by a novel B7 family member leads to negative regulation of lymphocyte activation. J. Exp. Med. 192, 1027–1034 (2000)
Okazaki, T. & Honjo, T. The PD-1–PD-L pathway in immunological tolerance. Trends Immunol. 27, 195–201 (2006)
Keir, M. E., Butte, M. J., Freeman, G. J. & Sharpe, A. H. PD-1 and its ligands in tolerance and immunity. Annu. Rev. Immunol. 26, 677–704 (2008)
Zou, W., Wolchok, J. D. & Chen, L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: Mechanisms, response biomarkers, and combinations. Sci. Transl. Med. 8, 328rv4 (2016)
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012)
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012)
Hamid, O. et al. Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N. Engl. J. Med. 369, 134–144 (2013)
Topalian, S. L. et al. Survival, durable tumor remission, and long-term safety in patients with advanced melanoma receiving nivolumab. J. Clin. Oncol. 32, 1020–1030 (2014)
Pollard, J. W. Tumour-educated macrophages promote tumour progression and metastasis. Nat. Rev. Cancer 4, 71–78 (2004)
Chao, M. P. et al. Anti-CD47 antibody synergizes with rituximab to promote phagocytosis and eradicate non-Hodgkin lymphoma. Cell 142, 699–713 (2010)
Forty Seven Inc. Phase 1 trial of Hu5F9-G4, a CD47-targeting antibody. https://clinicaltrials.gov/ct2/show/NCT02216409?term=cd47&rank=8 (2014)
Celgene. A Phase 1, dose finding study of CC-90002 in subjects with advanced solid and hematologic cancers. https://clinicaltrials.gov/ct2/show/NCT02367196?term=cd47&rank=7 (2015)
Huang, X. et al. PD-1 expression by macrophages plays a pathologic role in altering microbial clearance and the innate inflammatory response to sepsis. Proc. Natl Acad. Sci. USA 106, 6303–6308 (2009)
Bally, A. P. et al. NF-κB regulates PD-1 expression in macrophages. J. Immunol. 194, 4545–4554 (2015)
Chen, W., Wang, J., Jia, L., Liu, J. & Tian, Y. Attenuation of the programmed cell death-1 pathway increases the M1 polarization of macrophages induced by zymosan. Cell Death Dis. 7, e2115 (2016)
Shen, L. et al. PD-1/PD-L pathway inhibits M.tb-specific CD4+ T-cell functions and phagocytosis of macrophages in active tuberculosis. Sci. Rep. 6, 38362 (2016)
Sica, A., Schioppa, T., Mantovani, A. & Allavena, P. Tumour-associated macrophages are a distinct M2 polarised population promoting tumour progression: potential targets of anti-cancer therapy. Eur. J. Cancer 42, 717–727 (2006)
Esashi, E., Sekiguchi, T., Ito, H., Koyasu, S. & Miyajima, A. Cutting edge: a possible role for CD4+ thymic macrophages as professional scavengers of apoptotic thymocytes. J. Immunol. 171, 2773–2777 (2003)
Baba, T. et al. CD4+/CD8+ macrophages infiltrating at inflammatory sites: a population of monocytes/macrophages with a cytotoxic phenotype. Blood 107, 2004–2012 (2006)
Zhen, A. et al. CD4 ligation on human blood monocytes triggers macrophage differentiation and enhances HIV infection. J. Virol. 88, 9934–9946 (2014)
Maute, R. L. et al. Engineering high-affinity PD-1 variants for optimized immunotherapy and immuno-PET imaging. Proc. Natl Acad. Sci. USA 112, E6506–E6514 (2015)
Kleffel, S. et al. Melanoma cell-intrinsic PD-1 receptor functions promote tumor growth. Cell 162, 1242–1256 (2015)
Lin, D. Y. et al. The PD-1/PD-L1 complex resembles the antigen-binding Fv domains of antibodies and T cell receptors. Proc. Natl Acad. Sci. USA 105, 3011–3016 (2008)
Dahan, R. et al. FcγRs modulate the anti-tumor activity of antibodies targeting the PD-1/PD-L1 axis. Cancer Cell 28, 285–295 (2015)
Agata, Y. et al. Expression of the PD-1 antigen on the surface of stimulated mouse T and B lymphocytes. Int. Immunol. 8, 765–772 (1996)
Benson, D. M. Jr et al. The PD-1/PD-L1 axis modulates the natural killer cell versus multiple myeloma effect: a therapeutic target for CT-011, a novel monoclonal anti-PD-1 antibody. Blood 116, 2286–2294 (2010)
Karyampudi, L. et al. PD-1 blunts the function of ovarian tumor-infiltrating dendritic cells by inactivating NF-κB. Cancer Res. 76, 239–250 (2016)
Ansell, S. M. et al. PD-1 blockade with nivolumab in relapsed or refractory Hodgkin’s lymphoma. N. Engl. J. Med. 372, 311–319 (2015)
Reichel, J. et al. Flow sorting and exome sequencing reveal the oncogenome of primary Hodgkin and Reed–Sternberg cells. Blood 125, 1061–1072 (2015)
Saederup, N. et al. Selective chemokine receptor usage by central nervous system myeloid cells in CCR2-red fluorescent protein knock-in mice. PLoS One 5, e13693 (2010)
Maruyama, C. et al. Genotyping the mouse severe combined immunodeficiency mutation using the polymerase chain reaction with confronting two-pair primers (PCR-CTPP). Exp. Anim. 51, 391–393 (2002)
Takenaka, K. et al. Polymorphism in Sirpa modulates engraftment of human hematopoietic stem cells. Nat. Immunol. 8, 1313–1323 (2007)
MacDonald, K. P. et al. An antibody against the colony-stimulating factor 1 receptor depletes the resident subset of monocytes and tissue- and tumor-associated macrophages but does not inhibit inflammation. Blood 116, 3955–3963 (2010)
Acknowledgements
The authors thank S. Karten for assistance in editing the manuscript; and A. McCarty, T. Storm and T. Naik for technical support. Research reported in this publication was supported by the D. K. Ludwig Fund for Cancer Research (to I.L.W.); the A.P. Giannini Foundation and the Stanford Dean’s Fellowship (to M.N.M.); the Stanford Medical Scientist Training Program NIH-GM07365 (to B.M.G., B.W.D. and J.M.T.); a Cancer Research Institute Irvington Fellowship (to R.L.M.); and a Swiss National Science Foundation fellowship P300P3_155336 (to G.H.). The project described was supported, in apart, by ARRA Award Number 1S10RR026780-01 from the National Center for Research Resources (NCRR). Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the NCRR or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
S.R.G. wrote the manuscript. S.R.G., R.L.M., M.N.M., A.M.R. and I.L.W. conceived and designed all experiments. S.R.G. performed TAM staining, made the HAC protein and conducted all in vivo studies, phagocytosis assays and analysis. B.W.D. and R.S. helped with FACS gating and TAM analysis. G.H. generated NSG Ccr2−/− mice. B.M.G. conducted bone marrow transplants. S.R.G., R.L.M. and M.N.M. generated cell lines. R.G. acquired human colon cancer samples. J.M.T. taught the immunofluorescence protocol. D.C. and A.J.C. characterized foamy TAMs. R.L.M. and I.L.W. supervised the research and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.R.G., R.L.M., M.N.M., A.M.R. and I.L.W. are inventors on a patent (15/502,439) that is related to the HAC protein. S.R.G. and M.N.M. provide paid consulting services to Ab Initio Biotherapeutics Inc., which licensed this patent. R.L.M. and A.M.R. are founders of Ab Initio Biotherapeutics Inc.
Additional information
Reviewer Information Nature thanks V. A. Boussiotis, M. De Palma and A. Mantovani for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 FACS gating strategy for TAMs.
Debris and doublets were removed, then TAMs were assessed as Hoechst−CD45+CD8a−CD19−Ter119−TCRβ−CD11b+F4/80+. TAM PD-1 gating is shown as well, based on the PD-1 isotype control. All other gates were determined on the basis of FMOs. T cells, gated as Hoechst−CD45+TCRβ+CD8a+, are shown as PD-1+ positive control.
Extended Data Figure 2 TAM characterization.
a, No primary control for immunofluorescence images is shown. Cytospinned TAMs were stained with fluorescently conjugated secondary antibodies only. n = 2, two experimental repeats. 20× magnification; scale bar, 20 μm. Red, 594; green, 488; blue, Hoechst. b, Mouse PD-1− TAMs trend towards an M1 (CD206−MHC IIhigh) expression profile, rather than M2 (CD206+MHC IIlow or negative). TAMs that did not adhere to either of these expression profiles were not classified as M1 or M2. n = 5, experiment conducted once. Paired one-tailed t-test. c, Human PD-1− TAMs are predominantly M1 (CD206−CD64+) rather than M2 (CD206+CD64−). n = 10, two experimental repeats. Paired one-tailed t-test. d, In mice, there is a highly significant correlation between tumour volume and the percentage of PD-1+ TAMs. n = 20, two experimental repeats. An exponential growth equation is shown. e, Donor chimaerism six weeks after bone marrow transplantation. Granulocytes (Gr1high), 99%; myeloid cells (CD11b+), 92%; B cells (CD19+), 97%; T cells (TCRβ+), 74%. n = 8, experiment conducted once. Data are mean ± s.e.m.; **P < 0.01; n.s., not significant.
Extended Data Figure 3 Ex vivo phagocytosis assay with FACS-sorted TAMs.
Sorted PD-1− and PD-1+ TAMs from CT26 tumours were assayed with pHrodo green Staphylococcus aureus bioparticles. These particles are GFPlow at neutral pH, and GFPhigh in the acidic phagosome. a, Representative histogram showing difference in GFP fluorescence of PD-1− versus PD-1+ TAMs in the phagocytosis assay, and in comparison to S. aureus bioparticles alone. Bioparticles alone are clearly GFPlow, but have an obvious upshift in fluorescence when they are phagocytosed. b, Representative histograms showing the flow cytometry gating strategy for phagocytosis by PD-1− and PD-1+ TAMs. GFPhigh TAMs were considered to be phagocytosing. c, Analysis of phagocytosis shows that PD-1+ TAMs phagocytosed significantly less than PD-1− TAMS. n = 4, two experimental repeats. Paired one-tailed t-test. Data are mean ± s.e.m.; ****P < 0.0001.
Extended Data Figure 4 Immunocompromised mice also exhibit tumour-specific expression of PD-1 on macrophages.
a, Analysis of PD-L1-overexpressing CT26/YFP+ tumours in BALB/c Rag2−/−γc−/− mice shows that TAMs specifically express PD-1. n = 4, two experimental repeats. Paired one-way ANOVA with multiple comparisons correction. b, There is a highly significant correlation between BALB/c Rag2−/−γc−/− tumour volume and the percentage of PD-1+ TAMs. n = 9, two experimental repeats. Best fit line is shown. c, Analysis of DLD-tg(hPD-L1)-GFP-luc+ tumours shows that TAMs specifically express PD-1. n = 5, two experimental repeats. Paired one-way ANOVA with multiple comparisons correction. d, There is a highly significant correlation between NSG tumour volume and the percentage of PD-1+ TAMs. n = 1, two experimental repeats. Best fit line is shown. Data are mean ± s.e.m.; *P < 0.05; **P < 0.01; ***P < 0.001; n.s., not significant.
Extended Data Figure 5 In vivo phagocytosis analysis.
a, Representative FACS plots showing gating strategy for in vivo phagocytosis. Here, total phagocytosis was analysed by first gating on TAMs, and then gating on YFP+ cells. Total TAM PD-1 expression from the same tumour sample is shown side by side to demonstrate that high PD-1 expression inversely correlates with phagocytosis. b, Analysis of PD-1− TAM phagocytosis shows that the presence or absence of PD-L1 does not affect PD-1− TAM phagocytosis. PD-L1 overexpression, n = 7; PD-L1 knockout, n = 9, two experimental repeats. Paired one-tailed t-test. c, TAM PD-1 expression is not affected by the presence or absence of PD-L1. PD-L1 overexpression, n = 7; PD-L1 knockout, n = 9, two experimental repeats. Paired one-tailed t-test. Data are mean ± s.e.m.; n.s., not significant.
Extended Data Figure 6 In vivo TAM depletion.
TAMs were depleted with anti-CSF1R treatment in NSG-Ccr2−/− mice. a, TAM depletion protocol does not affect the number of granulocytes (Gr1high) in DLD-tg(PD-L1)-GFP-luc+ tumours. n = 10, experiment conducted once. Unpaired one-tailed t-test. b, TAM depletion protocol eliminates almost all TAMs in tumours. n = 10, experiment conducted once. Unpaired one-tailed t-test. Data are mean ± s.e.m.; ****P < 0.0001; n.s., not significant.
Supplementary information
Supplementary Information
This file contains a Supplementary Figure and Supplementary Tables 1-3. (PDF 221 kb)
Rights and permissions
About this article
Cite this article
Gordon, S., Maute, R., Dulken, B. et al. PD-1 expression by tumour-associated macrophages inhibits phagocytosis and tumour immunity. Nature 545, 495–499 (2017). https://doi.org/10.1038/nature22396
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22396
This article is cited by
-
Unravelling immune microenvironment features underlying tumor progression in the single-cell era
Cancer Cell International (2024)
-
Emerging mechanisms of the unfolded protein response in therapeutic resistance: from chemotherapy to Immunotherapy
Cell Communication and Signaling (2024)
-
Targeting the macrophage immunocheckpoint: a novel insight into solid tumor immunotherapy
Cell Communication and Signaling (2024)
-
Fine particulate matter 2.5 induces susceptibility to Pseudomonas aeruginosa infection via expansion of PD-L1high neutrophils in mice
Respiratory Research (2024)
-
Nanoparticles in tumor microenvironment remodeling and cancer immunotherapy
Journal of Hematology & Oncology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.