Abstract
Controlling plant disease has been a struggle for humankind since the advent of agriculture. Studies of plant immune mechanisms have led to strategies of engineering resistant crops through ectopic transcription of plants’ own defence genes, such as the master immune regulatory gene NPR1 (ref. 1). However, enhanced resistance obtained through such strategies is often associated with substantial penalties to fitness2, making the resulting products undesirable for agricultural applications. To remedy this problem, we sought more stringent mechanisms of expressing defence proteins. On the basis of our latest finding that translation of key immune regulators, such as TBF1 (ref. 3), is rapidly and transiently induced upon pathogen challenge (see accompanying paper4), we developed a ‘TBF1-cassette’ consisting of not only the immune-inducible promoter but also two pathogen-responsive upstream open reading frames (uORFsTBF1) of the TBF1 gene. Here we demonstrate that inclusion of uORFsTBF1-mediated translational control over the production of snc1-1 (an autoactivated immune receptor) in Arabidopsis thaliana and AtNPR1 in rice enables us to engineer broad-spectrum disease resistance without compromising plant fitness in the laboratory or in the field. This broadly applicable strategy may lead to decreased pesticide use and reduce the selective pressure for resistant pathogens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Fu, Z. Q. & Dong, X. Systemic acquired resistance: turning local infection into global defense. Annu. Rev. Plant Biol. 64, 839–863 (2013)
Gurr, S. J. & Rushton, P. J. Engineering plants with increased disease resistance: how are we going to express it? Trends Biotechnol. 23, 283–290 (2005)
Pajerowska-Mukhtar, K. M. et al. The HSF-like transcription factor TBF1 is a major molecular switch for plant growth-to-defense transition. Curr. Biol. 22, 103–112 (2012)
Xu, G. et al. Global translational reprogramming is a fundamental layer of immune regulation in plants. Nature http://dx.doi.org/10.1038/nature22371 (2017)
Gurr, S. J. & Rushton, P. J. Engineering plants with increased disease resistance: what are we going to express? Trends Biotechnol. 23, 275–282 (2005)
Boller, T. & Felix, G. A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 60, 379–406 (2009)
Schwessinger, B. et al. Transgenic expression of the dicotyledonous pattern recognition receptor EFR in rice leads to ligand-dependent activation of defense responses. PLoS Pathog. 11, e1004809 (2015)
Lacombe, S. et al. Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nature Biotechnol. 28, 365–369 (2010)
Bouwmeester, K. et al. The Arabidopsis lectin receptor kinase LecRK-I.9 enhances resistance to Phytophthora infestans in solanaceous plants. Plant Biotechnol. J. 12, 10–16 (2014)
Benedetti, M. et al. Plant immunity triggered by engineered in vivo release of oligogalacturonides, damage-associated molecular patterns. Proc. Natl Acad. Sci. USA 112, 5533–5538 (2015)
Jones, J. D. G. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006)
Dangl, J. L., Horvath, D. M. & Staskawicz, B. J. Pivoting the plant immune system from dissection to deployment. Science 341, 746–751 (2013)
Kim, S. H., Qi, D., Ashfield, T., Helm, M. & Innes, R. W. Using decoys to expand the recognition specificity of a plant disease resistance protein. Science 351, 684–687 (2016)
Chern, M. S. et al. Evidence for a disease-resistance pathway in rice similar to the NPR1-mediated signaling pathway in Arabidopsis. Plant J. 27, 101–113 (2001)
Quilis, J., Peñas, G., Messeguer, J., Brugidou, C. & San Segundo, B. The Arabidopsis AtNPR1 inversely modulates defense responses against fungal, bacterial, or viral pathogens while conferring hypersensitivity to abiotic stresses in transgenic rice. Mol. Plant Microbe Interact. 21, 1215–1231 (2008)
Molla, K. A. et al. Tissue-specific expression of Arabidopsis NPR1 gene in rice for sheath blight resistance without compromising phenotypic cost. Plant Sci. 250, 105–114 (2016)
Huot, B., Yao, J., Montgomery, B. L. & He, S. Y. Growth-defense tradeoffs in plants: a balancing act to optimize fitness. Mol. Plant 7, 1267–1287 (2014)
Johnson, K. C. M., Dong, O. X., Huang, Y. & Li, X. A rolling stone gathers no moss, but resistant plants must gather their moses. Cold Spring Harb. Symp. Quant. Biol. 77, 259–268 (2012)
Rahmani, F. et al. Sucrose control of translation mediated by an upstream open reading frame-encoded peptide. Plant Physiol. 150, 1356–1367 (2009)
Li, X., Clarke, J. D., Zhang, Y. & Dong, X. Activation of an EDS1-mediated R-gene pathway in the snc1 mutant leads to constitutive, NPR1-independent pathogen resistance. Mol. Plant Microbe Interact. 14, 1131–1139 (2001)
Li, Y., Yang, S., Yang, H. & Hua, J. The TIR-NB-LRR gene SNC1 is regulated at the transcript level by multiple factors. Mol. Plant Microbe Interact. 20, 1449–1456 (2007)
Yi, H. & Richards, E. J. A cluster of disease resistance genes in Arabidopsis is coordinately regulated by transcriptional activation and RNA silencing. Plant Cell 19, 2929–2939 (2007)
Fitzgerald, H. A., Chern, M. S., Navarre, R. & Ronald, P. C. Overexpression of (At)NPR1 in rice leads to a BTH- and environment-induced lesion-mimic/cell death phenotype. Mol. Plant Microbe Interact. 17, 140–151 (2004)
Lawless, C. et al. Upstream sequence elements direct post-transcriptional regulation of gene expression under stress conditions in yeast. BMC Genomics 10, 7 (2009)
Calvo, S. E., Pagliarini, D. J. & Mootha, V. K. Upstream open reading frames cause widespread reduction of protein expression and are polymorphic among humans. Proc. Natl Acad. Sci. USA 106, 7507–7512 (2009)
Xu, G. et al. One-step, zero-background ligation-independent cloning intron-containing hairpin RNA constructs for RNAi in plants. New Phytol. 187, 240–250 (2010)
Nakagawa, T. et al. Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104, 34–41 (2007)
Xu, G. et al. Plant ERD2-like proteins function as endoplasmic reticulum luminal protein receptors and participate in programmed cell death during innate immunity. Plant J. 72, 57–69 (2012)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Lin, Y. J. & Zhang, Q. Optimising the tissue culture conditions for high efficiency transformation of indica rice. Plant Cell Rep. 23, 540–547 (2005)
Yuan, M. et al. A host basal transcription factor is a key component for infection of rice by TALE-carrying bacteria. eLife 5, e19605 (2016)
Yuan, Y. et al. Functional analysis of rice NPR1-like genes reveals that OsNPR1/NH1 is the rice orthologue conferring disease resistance with enhanced herbivore susceptibility. Plant Biotechnol. J. 5, 313–324 (2007)
Kang, H. et al. Dissection of the genetic architecture of rice resistance to the blast fungus Magnaporthe oryzae. Mol. Plant Pathol. 17, 959–972 (2016)
Acknowledgements
This study was supported by grants from National Institutes of Health 5R01 GM069594-11 and Howard Hughes Medical Institute and Gordon and Betty Moore Foundation (through grant GBMF3032) to X.D., National Natural Science Foundation of China (31371926) to M.Y., and the National Key Research and Development Program of China (2016YFD0100903) to S.W. We thank J. Motley, J. Marqués, P. Zwack and S. Zebell for comments on this manuscript.
Author information
Authors and Affiliations
Contributions
G.X. and X.D. designed the research. G.X. performed the Arabidopsis-related experiments with help from E.Z. for fitness tests. L.L. isolated snc1 genomic DNA. S.K. maintained Hpa Noco2 strain in the laboratory and helped with inoculation; M.Y., C.A. and G.X. performed and S.W. supervised the rice-related experiments. G.X. and X.D. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
A patent based on this study has been filed by Duke University with G.X. and X.D. as inventors.
Additional information
Reviewer Information Nature thanks J. Bailey-Serres, P. Rushton and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Conservation of uORF2TBF1 nucleotide and peptide sequences in plant species.
a, Schematic of TBF1 mRNA structure. The 5′ leader sequence contains two uORFs, uORF1 and uORF2. CDS, coding sequence. b–d, Alignment of uORF2 nucleotide sequences (b) and alignment (c) and phylogeny (d) of uORF2 peptide sequences in different plant species. The corresponding triplets encoding the conserved amino acids among these species are underlined. Identical residues (black background), similar residues (grey background), and missing residues (dashes) were identified using Clustlw2. At (Arabidopsis thaliana; AT4G36988), Pv (Phaseolus vulgaris; XP_007155927), Gm (Glycine max; XP_006600987), Gr (Gossypium raimondii; CO115325), Nb (Nicotiana benthamiana; CK286574), Ca (Cicer arietinum; XP_004509145), Pd (Phoenix dactylifera; XP_008797266), Ma (Musa acuminata subsp. Malaccensis; XP_009410098), Os (O. sativa; Os09g28354).
Extended Data Figure 2 Characterization of uORFsTBF1 and uORFsbZIP11 in translational control.
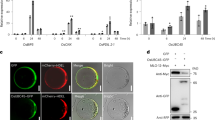
Related to Fig. 1. a, b, Subcellular localization of the LUC–YFP fusion (a) and GFPER (b). SP, signal peptide from Arabidopsis basic chitinase; HDEL, ER retention signal. Representative of eight images. Scale bar, 10 μm. c–e, mRNA levels of LUC in (Fig. 1b; n = 3), GFPER in (Fig. 1c; n = 4), and TBF1–YFP in (Fig. 1d; n = 3) 2 d.p.i. before cell death was observed in plants expressing TBF1. f, Schematics of the 5′ leader sequences used in studying the translational activities of WT uORFsbZIP11, mutant uorf2abZIP11 (ATG to CTG), or uorf2bbZIP11 (ATG to TAG). g–i, uORFsbZIP11-mediated translational control of cytosol-synthesized LUC (g; chemiluminescence with pseudo-colour); ER-synthesized GFPER (h; fluorescence under ultraviolet light); and cell death induced by overexpression of TBF1–YFP fusion (i; cleared using ethanol) after transient expression in N. benthamiana for 2 days (g, h) and 3 days (i), respectively. Representative of four images. j–l, mRNA levels of LUC in (g; n = 2 experiments with three technical replicates), GFPER in (h; n = 3 experiments with three technical replicates), and TBF1–YFP in (i; n = 3 experiments with three technical replicates). m, Translational efficiency changes in LUC controlled by the 5′ leader sequence containing WT uORFsbZIP11, mutant uorf2abZIP11, or uorf2bbZIP11 in response to elf18 in N. benthamiana. Mean of the LUC/RLUC activity ratios (n = 12). n, LUC/RLUC mRNA changes in m. Mean of LUC/RLUC mRNA normalized to Mock from two experiments with three technical replicates. Bar with solid circles, mean with individual biological replicates.
Extended Data Figure 3 Three developmental phenotypes observed in primary Arabidopsis transformants expressing snc1.
The three developmental phenotypes observed in T1 (that is, the first generation) Arabidopsis transgenic lines carrying 35S:uorfsTBF1-snc1, 35S:uORFsTBF1-snc1, TBF1p:uorfsTBF1-snc1, and TBF1p:uORFsTBF1-snc1 (above). Representative of five images. Fisher’s exact test was used for the pairwise statistical analysis (below). Different letters in ‘Total’ indicate significant differences between type III versus type I + type II (P < 0.01).
Extended Data Figure 4 Effects of controlling transcription and translation of snc1 on defence and fitness in Arabidopsis.
Related to Fig. 2. a, b, Psm ES4326 growth in WT, snc1, transgenic line numbers 1–4 after inoculation by spray (a) or infiltration (b). Mean ± s.e.m. c, Hpa Noco2 growth as measured by spore counts 7 d.p.i. Mean ± s.e.m. d–g, Analyses of plant radius (d), fresh weight (e), silique number (f), and total seed weight (g). Mean ± s.e.m. h, i, Relative levels of Psm ES4326-induced snc1 protein (h; numbers below immunoblots; see Supplementary Fig. 1 for gel source data) and mRNA (i; mean from two experiments with three technical replicates). Solid circles, individual biological replicates. Numbers 1–4, four independent transgenic lines carrying TBF1p:uORFsTBF1-snc1 with 1 and 2 shown in Fig. 2. h.p.i., hours after Psm ES4326 infection; CBB, Coomassie brilliant blue. See Source Data for sample size (n). Different letters above bars indicate significant differences (P < 0.05).
Extended Data Figure 5 Functionality of uORFsTBF1 in rice.
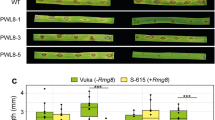
a, b, LUC activity (a) and mRNA levels (b) in three independent primary transgenic rice lines (called ‘T0’ in rice research) carrying 35S:uorfsTBF1-LUC and 35S:uORFsTBF1-LUC. Mean of LUC activities (RLU, relative light unit) of three biological replicates. Solid circles, individual biological replicates; and mean of LUC mRNA levels of three technical replicates after normalization to the 35S:uorfsTBF1-LUC line 1. c, Representative lesion mimic disease (LMD) phenotypes (above) and percentage of AtNPR1-transgenic rice plants showing lesion mimic disease in the second generation (T1) grown in the growth chamber (below).
Extended Data Figure 6 Effects of controlling transcription and translation of AtNPR1 on defence in T0 rice.
Related to Fig. 3. a–d, Lesion length measurements after infection by Xoo strain PXO347 in primary transformants (T0) for 35S:uorfsTBF1-AtNPR1 (a), 35S:uORFsTBF1-AtNPR1 (b), TBF1p:uorfsTBF1-AtNPR1 (c), and TBF1p:uORFsTBF1-AtNPR1 (d). Lines further analysed in T1 and T2 are circled. e, Average leaf lesion lengths. WT, recipient O. sativa cultivar ZH11. Mean ± s.e.m. Different letters above bars indicate significant differences (P < 0.05). See Source Data for sample size (n).
Extended Data Figure 7 Effects of controlling transcription and translation of AtNPR1 on defence in T1 rice.
Related to Fig. 3. a, b, Representative symptoms observed in T1 AtNPR1-transgenic rice plants grown in the greenhouse (a) after Xoo inoculation and corresponding leaf lesion length measurements (b). PCR was performed to detect the presence (+) or the absence (−) of the transgene gene. c, Quantification of leaf lesion length of four lines for Xoo inoculation in field-grown T1 AtNPR1-transgenic rice plants. Mean ± s.e.m. See Source Data for sample size (n). Different letters above bars indicate significant differences (P < 0.05). d, e, Relative levels of AtNPR1 mRNA (d) and protein (e; numbers below immunoblots; see Supplementary Fig. 1 for gel source data) in response to Xoo infection. Mean of AtNPR1 mRNA levels of three technical replicates after normalization to 0 h.p.i. (d). Solid circles, individual biological replicates.
Extended Data Figure 8 Effects of controlling transcription and translation of AtNPR1 on fitness in T1 rice under field conditions.
Related to Fig. 3. Mean ± s.e.m. See Source Data for sample size (n). Different letters above bars indicate significant differences among constructs (P < 0.05).
Supplementary information
Supplementary Figure
This file contains the uncropped immunoblots. (PDF 225 kb)
Supplementary Table 1
This file lists the plasmids, primers and antibodies in this study. (XLSX 14 kb)
Rights and permissions
About this article
Cite this article
Xu, G., Yuan, M., Ai, C. et al. uORF-mediated translation allows engineered plant disease resistance without fitness costs. Nature 545, 491–494 (2017). https://doi.org/10.1038/nature22372
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22372
This article is cited by
-
Synthetic modulation of ROS scavenging during host—Sclerotinia sclerotiorum interaction: a new strategy for the development of highly resistant plants
Phytopathology Research (2024)
-
Unravelling molecular mechanisms involved in resistance priming against downy mildew (Plasmopara viticola) in grapevine (Vitis vinifera L.)
Scientific Reports (2023)
-
Pervasive downstream RNA hairpins dynamically dictate start-codon selection
Nature (2023)
-
Plant HEM1 specifies a condensation domain to control immune gene translation
Nature Plants (2023)
-
A tripartite rheostat controls self-regulated host plant resistance to insects
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.