Abstract
The xa5 gene encodes a basal transcription factor (TFIIAγ) protein with wide spectrum resistance to bacterial blight caused by Xanthomonas oryzae pv. Oryzae (Xoo) in rice. It was only found in a few rice ecotypes, and the recessive characteristics limited its application in breeding. Here, we employed a TALEN-based technique to edit its dominant allelic TFIIAγ5 and obtained many mutant TFIIAγ5 genes. Most of them reduced rice susceptibility to varying degrees when the plants were challenged with the Xoo. In particular, the knocked-out TFIIAγ5 can reduce the rice susceptibility significantly, although it cannot reach the xa5-mediated resistance level, indicating TFIIAγ5 is a major component involved in disease susceptibility. In addition, the mutant encoding the protein with deletion of the 32nd amino acid or amino acid insertion between 32nd and 33rd site confers rice with the similar resistance to that of the knocked-out TFIIAγ5. Thus, the amino acids around 32nd site are also the important action sites of TFIIAγ5 besides the 39th amino acid previously reported. Moreover, the integration of xa5 into TFIIAγ5-knockout plants conferred them with a similar resistance as IRBB5, the rice variety containing the homozygous xa5 gene. Thus, TFIIAγ5 was not simply regarded as a resistant or a susceptible locus, as the substitution of amino acids might shift its functions.
Similar content being viewed by others
Introduction
Plant diseases caused by the bacterial from genus Xanthomonas can reduce the yield and quality in a variety of crops. During infection, the Xanthomonas bacterium delivers types of effectors into host cells through a type III secretion (T3S) pathway. Transcription activator-like effectors (TALEs) are major virulence factors in Xanthomonas. They modulate host transcription to facilitate pathogen growth or propagation1. TALEs can trans-activate host genes by directly binding to their effector binding elements (EBEs) in the DNA2. In addition, TALEs can combine other protein factors to control gene expression or to alert protein function, besides binding to DNA directly. Some truncated TALEs, also termed interfering TALEs or iTALEs, were used by the pathogen Xanthomonas oryzae and suppressed the disease resistance of plants by altering the action of resistant factors3,4. However, some TALEs such as AvrXa105, AvrXa276, and AvrXa237 can activate resistant genes in plants.
Bacterial blight caused by Xanthomonas oryzae pv. oryzae (Xoo) is a well-studied disease in rice. In the interaction between Xoo and rice, the TALEs were shown to play important roles in regulating the expressions of rice susceptible or resistant genes. To date, more than 40 bacterial blight resistant (R) genes in rice have been identified8 and eleven of them have been cloned and characterized5,6,7,9,10,11,12,13,14,15,16. Among these cloned genes, the dominant R genes Xa1, Xa10, Xa23, and Xa27 can be induced by TALEs to trigger hypersensitive response (HR) and cell death3,5,6,7. The recessive R genes such as xa13, xa25, and xa41 confer rice resistance because the TAL effectors cannot recognize their promoter to activate them16,17,18,19. However, the recessive R gene xa5 expresses constitutively and encodes a TFIIAγ-like protein, indicating that it behaves differently from other R genes in the Xoo-rice interaction.
TFIIAγ is a general transcription factor involved in RNA polymerase II-dependent transcription in higher eukaryotes20. In rice, the OsTFIIAγ5/Xa5 gene is showed to play an important role response to bacterial pathogens21. Xa5/TFIIAγ5 can work as a key component in the disease resistance mediated by some dominant R genes, such as Xa2722 and Xa105. On the other hand, it was shown to interact directly with TALEs to activate disease susceptibility genes. The attenuation or block of this interaction could eliminate or weaken the activation of rice susceptibility gene expression21. The two-nucleotide change in its second exon, resulting in the produce of recessive gene xa5, encoding a protein with the substitution of glutamic acid at 39th position to valine (E39V)13,23. The xa5 gene is also very important in rice breeding for its broad resistance spectrum to most Xoo strains, except a few strains employing the TALE of pthXo124. The pyramid lines of the xa5 gene and some other dominant genes such as Xa21 and Xa4 have a higher and wider spectrum resistance than the plants harboring only one BB resistance gene25. Nevertheless, the xa5 is a recessive gene and found only in a few rice ecotypes26, restricting its application on rice breeding. These previous studies may provide us some new ideas to use Xa5/TFIIAγ5 genes for breeding purposes.
Transcription activator-like effector nucleases (TALENs) are programmable nucleases in which a truncated TALE is linked to the catalytic domain of FokI for targeted genetic modifications. The target recognition is commanded by assembling TALEs using a code-like modularity of TALE-DNA recognition27. TALEs possess an N-terminal T3S signal mediating their translocation to host cells4,28, a C-terminal activation domain activating host gene transcription29, two nuclear localization signals (NLS) directing them into host nucleus30, and a central repeat domain recognizing the target sequence by one repeat recognizing one nucleotide31,32. The TALE repeats have a similar composition and construction. A typical repeat is 34 aa long, forming a right-handed superhelix that can wrap around a double-stranded DNA33,34. The specification of a repeat is conferred by a pair of neighbored amino acids at position 12 and 13, called the repeat-variable diresidue (RVD), which is situated at the loop linking two helices. The 12th residue stabilizes the loop and projects it into the major groove, while the 13th residue contacts the base deciding the specificity of target recognition35. The specificities and efficiencies of RVDs are variable. Each RVD contacts one or more DNA bases and efficient activation requires a well-balanced composition of “strong” and “weak” RVD36. The common correspondence between RVDs and their preferred nucleotides are interpreted as NI:A, HD:C, NG:T, NH:G, NN: G/A, NS:A/C/G/T30. TALENs could be designed to target almost any given DNA sequence in theory, but there are two limitations in the design of TALENs. One is the requirement for a thymine at the 5′ end of the target sequence, which will be recognized by the two amino-terminal cryptic repeat folds37. The other is the fussy and labored procedure to construct a powerful editing plasmid. In spite of these, many companies have chosen this method to edit target gene for its high precision38.
In view of the facts that the OsTFIIAγ5/Xa5 gene exists in the overwhelming majority rice varieties, the change of key amino acids in OsTFIIAγ5/Xa5 can break up its interaction with TALEs and the mutant OsTFIIAγ5/Xa5 can improve the disease resistance mediated by Xa21 or Xa4 gene, we selected TALEN to edit the TFIIAγ5 gene in popular breeding varieties and hope to improve their resistance to bacterial blight. Here we obtained many TFIIAγ5 mutations of long-fragment deletion, substitution and insertion, and found that the TFIIAγ5 plays a double role in pathogen invasion, and the amino acids between the 32nd and 40th are critical for its function.
Results
TFIIAγ5 plays a role in the host infection of Xoo
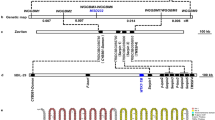
To assess whether the host TFIIAγ5 is required for the invasion of Xoo, we edited the TFIIAγ5 using the TALEN method. The target sequence of our TALEN was designed from the first exon of TFIIAγ5 to enhance the incidence of knock-out mutants (Fig. 1). The space sequence between the couple of target sequences is more mutable than the flanking regions, and enzyme restriction sites are normally designed here to detect the presence of mutations. Here, two restriction sites for BbvCI and SacI in the space sequence were employed to detect mutations by PCR restriction enzyme digestion assay (PCR-RE).
Structure of the gene TFIIAγ5 (Xa5andxa5) displaying the discrepancy between the allelic Xa5 and xa5 that was involved in recognition by the TALEN-Xa5. The rice gene TFIIAγ5 has two natural alleles, Xa5 and xa5. The gene structure of both genes is the same and is shown above. The target region of the TALEN-Xa5 locates in the exon 1 and marked by a red strip. The underlined target sequence of TALEN-Xa5 in Xa5 and xa5 is showed in the bottom of the figure. The discrepancy of the two nucleotides between Xa5 and xa5 was also showed in red letters.
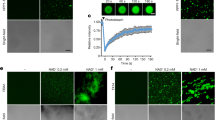
Considering the two nucleotides difference between xa5 and Xa5 (Fig. 1), the target sequence of the TALEN-Xa5 was designed to cover these two sites. The transient expression of the TALEN plasmid was first conducted in rice protoplasts containing Xa5 (TP309) and xa5 (IRBB5), respectively. Then the genome DNA was isolated from these protoplasts and subjected to the PCR-RE analysis. Sequence analyses of these PCR productions showed that the fragments from the xa5 gene all kept the initial characteristics of BbvCI and SacI digestion (Supp. Fig. S1). The result showed that the TALEN-Xa5 plasmid specializes in the Xa5 edition.
Subsequently, the TALEN-Xa5 plasmid was transformed into a rice callus by Agrobacterium-mediated genetic transformation and yielded many transgenic lines. We analyzed these transgenic seedlings through PCR and a sequencing method, and finally obtained T1 seedlings with homozygous frame-shift mutations of Xa5. Firstly, the FokI gene was detected in these transgenic plants (see the PCR primer showed in Supp. Table S1). The absence of FokI in T1 generation plants meant that their Xa5 gene was not edited later. Secondly, the Xa5 mutants in these plants without FokI were selected by PCR/RE analyses (see PCR primer showed in Supp. Table S1) and sequencing. Finally, the T2 transgenic plants with the homozygous frame-shift mutant Xa5 were obtained and inoculated with the Xoo strain PXO86. Two of these (also referred to as the knock-out mutants), line 119 and line 123 deriving from the Japonica variety TP309, were checked for two consecutive generations in Beijing in 2015 and in Hainan in 2016 (Fig. 2a). The blank control was TP309 and the transgenic control was a line of the positive transgenic TP309 (TP), which did not contain the FokI gene in the T1 generation and thus had an intact Xa5 gene in the T2 generation. The lesion length of TP309 was 9.82 ± 1.90 cm in the experiment in Beijing and 6.51 ± 0.67 cm in the Hainan experiment. The lesion length of the TP plant was 8.20 ± 1.20 cm in the Beijing experiment, and the lesion length of the three T3 lines from the T2 transgenic control was 5.38 ± 0.82, 5.36 ± 0.93, 5.84 ± 0.97 cm, respectively, in the Hainan experiment. Apparently, although each line had a longer lesion length in the 2015 experiment, all mutation lines had a shorter lesion length than the controls. Similar results were obtained from the knock-out mutants of Indica varieties, MH86 (Fig. 2b) and D62B (Fig. 2c). All these results showed that the knock-out of TFIIAγ5 alleviates the bacterial leaf blight disease in rice (Fig. 2), and TFIIAγ5 might facilitate Xoo in the host infection
The lesions length of rice leaves in the knock-out mutants 14 days after inoculation with PXO86. (a) The T2 plants and their derivative T3 plants were inoculated by PXO86 in Beijing in August, 2015 and 2016 respectively. (b,c) The T3 plants in MH86 or D62B background were examined in Hainan in February, 2016. (d) The disease symptoms of the T3 plants were photographed 14 days after inoculation. (e) The targeted sequences of TALEN-Xa5 in transgenic plants. WT is the initial target sequence of Xa5, and the others are the mutant sequences. TP309, MH86, and D62B. TP309, MH86, and D62B are blank controls; TP, 1TP, 2TP and 3T were transgenic controls in TP309 background with pCAMBIA1300 vector; 1MH and 2MH are transgenic controls in MH86 background; 1D6 is the transgenic control in D62B background. All controls had the same target sequence as the WT. The other rice lines were knock-out mutants with their target sequences shown in e. TP, 119, 123, 278, 23 and 179 are the homozygous T2 mutant plants. 1TP, 2TP and 3TP are the random selected T3 plants derived from the TP; 119-1, 119-2 and 119-3 are the T3 plants from the 119; 123-1, 123-2, 123-3, and 123-4 are the T3 plants from the 123. The 278-1 and 278-2 are homozygous mutant T3 plants from the 278. 23-1 and 179-1 are homozygous mutant T3 plants from 23 and 179 respectively. Bars represent the average ± SD of three biological repeats. Different letters above columns indicate significant differences at P < 0.05 as determined by a one-way ANOVA followed by post hoc Tukey honest significant difference (HSD) analysis.
TFIIAγ5 can be used by different Xoo strains to infect rice
When infecting a plant, bacteria induce and recruit host genes by secreting effectors into the host cells through the type III system. These effectors can interact with the host components to exert their function. To explore the function of TFIIAγ5 in the infection process, we analyzed the disease symptoms of these knock-out mutants by inoculating them with different Xoo strains (Fig. 3).
Resistance spectrum analyses of the Xa5 knockout mutants. The spectrum analysis was carried out in TP309 (a), D62B (b), and MH86 (c) genetic backgrounds. (d) The targeted sequences of TALEN-Xa5 in transgenic plants. The target sequence in TP309, D62B, and MH86 are same and refer as WT; 192, 294 and 23 are the homozygous T2 knockout mutants from TP309, D62B and MH86 respectively. PXO86 (P2), PXO71 (P4), PXO112 (P5), PXO145 (P7) and PXO280 (P8) are the Philippines Xoo strains; HN01 is a newly isolated Xoo strain from Hainan, China. Bars represent the average ± SD of three biological repeats. Asterisks signs indicate a statistically significant difference compared with the control plants (*P < 0.05 and **P < 0.01).
The TP309 mutant line 192 had a 5-nucleotide deletion in the first exon of TFIIAγ5 (Fig. 3a), the D62B mutant line 294 had a 1-nucleotide deletion in the first exon (Fig. 3b), and the MH86 mutant line 23 had a 25-nucleotide deletion in the first exon (Fig. 3c).
Six Xoo strains were used to examine whether the TFIIAγ5 was recruited during infection. Five of the strains were isolated from the Philippines, namely PXO86 (P2), PXO71 (P4), PXO112 (P5), PXO145 (P7) and PXO280 (P8). The other strain was HN01, a new isolate from Hainan, which can overcome the resistance of Xa21 in rice. The statistical analysis showed that the Xa5 depletion decreased the rice’s susceptibility to Xoo with different genetic backgrounds (Fig. 3a–c). These indicated that the TFIIAγ5 can be used by different Xoo strains to infect rice.
Xa5 and xa5 function differently in response to the invasion of Xoo
The severity of bacterial leaf blight depends on many factors, such as the growing environment and genetic background. However, the inoculation analyses in 2015 and 2016 year at two breeding bases in Beijing and Hainan exhibited a similar trend, namely that the knock-out mutant plants of Xa5 showed enhanced resistance to Xoo than the control plants (Fig. 2). IRBB5 and IR24 are near-isogenic lines, and they respectively had xa5 and Xa5 in its TFIIAγ5 loci (Fig. 1). It is a pity that we were not able to obtain the knock-out mutant from IR24, due to its poor capacity for regeneration after transformation. Thus, it is difficult to determine whether the high resistance of IRBB5 was brought about by xa5 or merely by the genetic background.
To further clarify the function of TFIIAγ5 in pathogen infection, xa5 was transformed into two knock-out mutants of TFIIAγ5, CX1 and CX5 by the agrobacterium-mediated method. CX1 and CX5 are from the rice breed CX6221B, which carries the resistant gene Xa21 in the genetic background of the rice D62B39. The TFIIAγ5 lacks 11 and 38 nucleotides in the first exon respectively in these two mutants (see the bottom of the Fig. 4). At least 20 T0 positive transgenic lines for each transformation were obtained. These transgenic plants carrying the xa5 gene were named CX1-xa5 or CX5-xa5. In 2015, they were inoculated with the Xoo strain HN01, which was able to overcome the resistance of Xa21 in rice. All these plants showed significantly enhanced disease resistance, and have shorter lesions of leaves than the two wild type plants and the TFIIAγ5 knock-out mutants (Fig. 4). These indicated that the Xa5 and xa5 function differently in response to the invasion of pathogens.
Resistance analysis of Xa5 and xa5 in the plants containing Xa21 gene. (a)The lesions length of rice leaves in the knock-out mutants 14 days after inoculation with HN01. (b) The targeted sequences of TALEN-Xa5 in transgenic plants. CX1 and CX5 are T3 knock-out mutants from CX6221B (a stable hybrid line with Xa21 gene in D62B background), CX1-xa5 and CX5-xa5 are the T0 transgenic plants with xa5 gene in CX1 and CX5 background respectively. CX6221B contains Xa5 and Xa21 gene, while IRBB5 only has the gene xa5. The HN01 was able to overcome the resistance of Xa21 gene, but not that of xa5 gene. The standard deviation (STDEV) is indicated in each column. Bars represent the average ± SD of three biological repeats. Different letters above columns indicate significant differences at P < 0.05 as determined by a one-way ANOVA followed by post hoc Tukey honest significant difference (HSD) analysis.
The amino acids around 32nd site are important for TFIIAγ5 in the response to Xoo
In the TALEN-Xa5 transformation, eight types of in-frame mutants were generated besides the knocked-out mutants of Xa5 (Table 1). All the edited sites in these mutants took place in the region between the 26th and the 40th amino acids of TFIIAγ5. These mutations were then tested for their resistance to Xoo in the T2 or T3 homozygous plants lacking intact TALEN-Xa5 proteins. For example, the type 1 mutation lacked the 32nd amino acid and the type 8 was inserted by a glutamic acid (E) between the 32nd and the 33rd amino acid. The lesion length of the mutants of type 1 was significantly shorter than that of their wild type varieties (Fig. 5a). The lesion length of leaves in the mutant 3 from MH86 was 6.00 ± 1.09 cm; the length of the lesions on leaves in the mutants O23 and O32 from TP309 were 4.61 ± 0.72 and 4.14 ± 0.79 cm, respectively. However, the lesion length of leaves in MH86 was 9.47 ± 1.78 cm, and that of TP309 was 9.82 ± 1.90 cm. Moreover, the lesion length of leaves in these type 1 mutant plants were similar to that in the other types of in-frame mutant plants in each background. The length of the lesions on leaves in the type 7 mutant 217 and type 8 mutant 235 from MH86 were 5.24 ± 0.84 and 5.15 ± 1.65 cm, respectively, similar to that of mutant 3. The lesion length of the leaves in the type 3 mutants 133 was 5.36 ± 0.86 cm, similar with that of the mutants of O23 and O32 (Fig. 5a). The knockout lines 119 and 123 were 5.65 ± 1.32 and 5.32 ± 1.35 cm, respectively, similar to the in-frame lines with the TP309 background (Supp. Fig. S2a). The type 8 mutant 260-1 from D62B harbored an amino acid insertion between the 32nd and 33rd amino acids. The lesion length of its leaves was 3.9 ± 0.73 cm, which was shorter than that of D62B, which was 6.95 ± 1.25 cm (Fig. 5b). However, it was slightly longer than that in the type 4 mutant 180-1 (2.97 ± 0.49 cm) and the knocked-out mutant 179-1 (2.17 ± 0.73 cm) with the same background (Supp. Fig. S2b). The type 4 mutant 180-1 lacked the three amino acids from the 32nd to the 34th amino acid of the Xa5 (Table 1). All these results indicated that the amino acids around the 32nd site might be crucial to TFIIAγ5 in governing rice’s susceptibility to Xoo.
Lesion length in rice leaves of in-frame mutants inoculated with Xoo. (a) The homozygous T2 plants of in-frame mutants were inoculated by PXO86 in Beijing in August, 2015. TP309 and MH86 were the control plants containing the gene Xa5; TP is the T2 transgenic control plant from TP309 with pCAMBIA1300 vector. 133 is the T2 plant from TP309 with mutant Xa5 protein, in which two amino acids were replaced with six amino acids in the exon 1 (type 3 in the Table 1). The mutant Xa5 proteins in O23, O32 and 3 all have an amino acid deletion in the exon 1 (type 1 in Table 1). 217 expresses a mutant Xa5 protein with four amino acid-deletion in the exon 1 (type 5 in Table 1). 235 expresses the mutant Xa5 protein, in which five amino acids were replaced with nine amino acids in the exon 1 (type 6 in the Table 1). IRBB5 contains the homozygous resistant gene xa5, and functions as the resistant control here. (b) The homozygous T3 plants of in-frame mutants from D62B and IR24 were inoculated with PXO86 in Hainan in February 2016. D62B and IR24 are the rice varieties with homozygous Xa5 gene. 1D6 and 1IR are the T3 transgenic control plants with pCAMBIA1300 plasmid derived from D62B and IR24 respectively. 180-1 expresses a mutant Xa5 protein with three amino acid-deletion in the exon 1 (type 4 in Table 1). 260-1 expresses a mutant Xa5 protein with an amino acid insertion in the exon 1 (type 8 in Table 1). 38-1 and 38-2 are the T2 plants derived from 38 that have a nine amino acid-deletion in the exon 1 of the Xa5 protein (type 2 in Table 1). Bars represent the average ± SD of three biological repeats. Different letters above columns indicate significant differences at P < 0.05 as determined by a one-way ANOVA followed by post hoc Tukey honest significant difference (HSD) analysis.
In a previous study, the 39th amino acid was characterized as a critical site of Xa513. The 39th amino acid substitution of Xa5 leads it to be a recessive resistance protein xa5. Here we obtained two in-frame mutants 38-1 and 38-2 from IR24, they both had the deletion from the 26th to the 34th amino acids (type 2 in Table 1). IR24 containing the Xa5 gene was the near isogenic line of resistant variety IRBB5 that harbors the homozygous xa5 gene. The two mutant plants were inoculated with the Xoo strain PXO86 and investigated after two weeks. The lesion length of leaves in IR24 was 5.91 ± 0.93 cm and in IRBB5 this was 2.36 ± 0.47 cm. The length of the lesions on leaves in the mutants 38-1 and 38-2 were 4.36 ± 0.82 and 4.44 ± 0.58 cm, respectively, and were shorter than that of IR24 but longer than that of IRBB5 (Fig. 5b), indicating that the mutant Xa5 of type 2 enhance rice resistance, though the resistance is not as strong as that conferred by xa5. All these suggested that the amino acids around 32nd site are the critical amino acids determining TFIIAγ5’s action.
TALEN-Xa5 can mediate off-target modification to the TFIIAγ1 gene
There are two TFIIAγ-like genes, TFIIAγ1 and TFIIAγ5 in the rice genome. Sequence alignment showed that the sequence identity between TFIIAγ1 and TFIIAγ5 was 85.8% at the protein level and 81.3% on the cDNA level (Supp. Fig. S3). The possible off-target sequence in TFIIAγ1 was determined through analyzing the sequences of TFIIAγ1 and TFIIAγ5. Four mismatched sites were found in each of the two target regions (Fig. 6). Because the experiment of transiently expressing TALEN-Xa5 in rice protoplasts showed that the two nucleotides of difference in the target sequence led to undetectable edition in the gene Xa5 (Supp. Fig. S1), the off-target detection of TALEN-Xa5 on the TFIIAγ1 was only conducted in the Xa5 mutant lines for the definite editing activity. The cutting site (GAGCTC) for the restriction enzyme SacI in the two target sequences was used to identify the off-target in TFIIAγ1 through PCR/RE analysis (Fig. 6). The PCR product for the potential target sequence of TFIIAγ1 can be digested by SacI into a band of 302 bp and a band of 511 bp. In the 49 T0 lines containing edited TFIIAγ5, only four lines were detected to have edited TFIIAγ1 by the PCR/RE analysis, indicating that the high degree of nucleotide sequence similarity between TFIIAγ5 and TFIIAγ1 can lead to the off-target of TALEN-Xa5 on the TFIIAγ1 though the off-target efficiency is very low (Fig. 6). In addition, the four TFIIAγ1 mutants were all chimeras or heterozygotes and had many various mutations besides the wild type of TFIIAγ1 (Supp. Table S2). It is regrettable that none of the four TFIIAγ1 mutants survived for the T1 generation; one died in the seedling stage and the others were sterile.
The off-target analysis of TALEN-Xa5 in the rice TFIIAγ1 gene. (a) The possible target region of TALEN-Xa5 in the rice TFIIAγ1 gene. The target sequence in the gene Xa5 and the possible target sequence in the gene TFIIAγ1 were underlined. The restriction enzyme cutting site of SacI in the space region used to detect the mutations was marked with a blue box. The SacI can cut the PCR product into two fragments of 511 bp and 302 bp in theory. (b) The T0 generation plants of TALEN-Xa5-transformed lines in TP309 background were examined by PCR/RE. 148T0, 207T0 and 221T0 are the plants with the mutant TFIIAγ1 gene. CK1 is TP309 plant. (c) The T0 generation plants of TALEN-Xa5-transformed lines in MH86 background were examined by PCR/RE. 237 T0 is the plant with the mutant TFIIAγ1 gene. CK2 is MH86 plant. The primers of TFIIA-gamma-1F and TFIIA-gamma-1Rwere used to examine off-target activity in TFIIAγ1 were shown in Supp. Table S2.
The disease resistance mediated by Xa21 was independent of TFIIAγ5
Xa21, encoding a receptor kinase-like protein, confers on rice a broad-spectrum resistance to multiple pathogen isolates of Xanthomonas oryzae pv. oryzae. It is a dominant gene and is usually used together with other resistant genes such as xa5 in breeding for permanent and stable disease resistance. In this study, we also edited Xa5 in the plants with the Xa21 gene, CX6221 and CX862139. They were derived from the two widely used restorer lines D62B and MH86 by integrating a single copy of Xa21. Four homozygous mutant Xa5 T2 lines and three homozygous Xa5 mutant T3 lines were obtained, respectively, in CX6221 and CX8621. Most of these mutant lines were knocked-out mutants except for the line 250-1 from CX8621 (the type 9 mutation in Table 1) that lacked 13 amino acids from the 19th to the 32nd of Xa5. To analyze the resistance phenotypes, these mutants were cultured and inoculated with the Xoo strain HN01 at the tillering stage in Beijing or Hainan (Fig. 7a,b). The HN01 was isolated from Hainan and can overcome the disease resistance conferred by Xa21, but not xa5.
Lesion lengths in the leaves of TFIIAγ5/Xa5 mutant plants with Xa21 gene. (a) Inoculation analysis of the T2 mutant plants from CX6221B was conducted in Beijing in August, 2015. (b) Inoculation analysis of the T3 mutant plants from CX8621 was carried out in Hainan in February, 2016. 248-1, 250-1 and 252-1 are the T3 homozygous mutants in CX8621 background from the T2 plants 248, 250 and 252, respectively. (c) The targeted sequences of TALEN-Xa5 in transgenic plants. CX6221B and CX8621 are the plants contain the homozygous resistant gene Xa21 derived from the varieties D62B and MH86 respectively. 143, 144, 146, and 147 are the T2 mutants in CX6221B background. D62B, CX6221B and CX8621 have the wild type Xa5, shown as WT in the bottom table. All the plants were inoculated with Xoo strain, HN01. Bars represent the average ± SD of three biological repeats. Asterisks signs indicate a statistically significant difference compared with the control plants (*P < 0.05 and **P < 0.01).
The length of the lesions on the leaves in the four mutants from CX6221B ranged from 5.98 ± 1.32 to 6.75 ± 1.24 cm, indicating that they have a uniform phenotype (P > 0.05) and are significantly shorter (P < 0.01) than that of the control plant CX6221B (about 10.41 ± 1.37 cm). The same results were achieved from the mutant CX8621 plants. The length of the lesions on the leaves in 248-1, 250-1 and 252-1 were 2.80 ± 0.75, 2.65 ± 0.52, 2.95 ± 0.59 cm, respectively, and are shorter than that of CX8621(4.81 ± 0.87 cm). These results were similar to the experiment on the knocked-out mutants of Xa5 inoculated by PXO86 (Fig. 2). In the same way, these mutant plants were inoculated with other Xa21-incompatible Xoo strains, PXO99 (P6), PXO71 (P4), PXO112 (P5), PXO145 (P7), and PXO280 (P8). These mutant plants showed a similar phenotype to the control plants, and the lengths of the lesions on their leaves were all less than 1 cm. This indicated that the knocked-out mutants of Xa5 all displayed a similar reduced effect on the disease symptoms of rice plants with or without Xa21, though climate and environment may have some influence on disease development (Fig. 7a,b).
Discussion
Bacterial blight caused by Xoo is one of the most destructive rice diseases throughout the world, and stands out in the top 10 bacterial diseases (Mansfield et al., 2012). In some areas of Asia, this disease can reduce crop yield by up to 50%. The most effective approach to combat it is the use of resistant varieties (Khush et al. 1989). More than 40 bacterial blight resistance genes have been identified in rice and ten of them have been cloned and characterized (Kim et al. 2015; Hutin et al. 2015). DNA sequence and function studies have showed that the encoding proteins of these cloned genes are structurally diverse, indicating that their resistance mechanisms to bacterial blight are very complicated (Song et al. 1995). To date, only a few of these genes have been used for resistance breeding (Hutin et al. 2015). Generally, the dominant resistance genes have been paid the most attention in breeding. For example, numerous resistant breeding lines harboring the dominant resistance gene Xa4 have been developed since the first resistant varieties, IR20 and IR22, were released in 1969 (Khush GS, 1977). In China, the Xa4 gene has also been widely integrated into the parental lines of hybrid rice since 1980 (Zhang, 2009). The first map-based cloned broad-spectrum resistance gene, Xa21, was introduced into many rice varieties and other plants through agrobacterium-mediated or marker-assisted breeding method (Datta et al. 2002; Huang et al. 1997; Kottapalli et al. 2010; Luo and Yin 2013; Luo et al. 2014; Perez et al. 2008; Rajpourohit et al. 2011; Singh et al. 2001; Zhang et al. 2006; Zhai et al. 2000; Tripathi et al. 2014). However, these dominant genes are vulnerable to counter the evolution of pathogens (Lee et al. 1999). Comparatively speaking, the resistance conferred by recessive genes is much more stable, but the recessive properties limit their application in rice breeding through conventional methods or plant biotechnology approaches. In this study, we used TALEN-based techniques to edit the dominant allelic of the xa5 gene and obtained a serial of mutant Xa5/ TFIIAγ5 genes with different rice backgrounds. In the T0 or T1 transgenic plants, most were able to reduce rice susceptibility to varying degrees, indicating that the TALEN-based editing technique is very efficient in accelerating the application of the xa5 gene in rice breeding.
The xa5 gene was first characterized in the rice varieties IR1545-284 and RP291-7 (Petpisit et al. 1977). It confers rice a broad-spectrum resistance to Xoo strains, including the Philippine races 1–3 and all Japanese races. Previous studies showed that the pyramid lines of the xa5 gene and some other dominant genes such as Xa21 and Xa4 have a higher and wider spectrum resistance than the plants harboring only one resistance gene (Huang et al. 1997). However, there is an exceptional case, where the resistance mediated by Xa27 depends on the Xa5 (OsTFIIAγ5) instead of xa5 (OsTFIIAγ5V39E)5,22,24. The current model of Xa5 is that it interacted directly with the TAL effector of Xoo or Xoc to cause bacterial blight or streak21. The xa5 is correlated with the reduced expressions of OsSWEET genes, which led to a virulence effector-dependent quantitative trait for bacterial blight24. To probe the particular function of Xa5, we edited it through the TALEN-based technique and obtained many mutants including in-frame and knock-out mutants. These edited Xa5 mutants enhanced rice resistance to Xoo (Fig. 2 and Supplementary Fig. S2), but their resistance was inferior to that conferred by the resistant gene xa5 (Figs. 2 and 4). When these mutant plants were transformed with the xa5 gene, they were similar to IRBB5 in disease resistance. These results indicated that the mutant Xa5 with 39th amino acid replacement has a different effect from that with the amino acid deletion or insertion around 32nd site in response to Xoo. However, the previously study showed that the amino acids between 31st and 50th form the domain of alpha-helices 3 (H3) in Xa513, the amino acids in the domain are logical to have the same effects. We speculated that the different effects are due to the amino acids deletion or insertion of Xa5 can destroy H3 domain, while the amino acid replacement just change the conformation or binding property of Xa5.
Previous studies showed that TALEN-mediated genome modification is accompanied by very rare off-target effects40,41. In this study, among the 49 T0 transgenic lines containing edited TFIIAγ5, four lines were detected to have edited TFIIAγ1 by the PCR/RE analysis, showing that TALEN can mediate off-target modification though the efficiency is very low. TFIIAγ1 and TFIIAγ5 were speculated to arise from whole genome duplication in the common ancestor of grasses, and share a high degree of nucleotide sequence similarity with each other13,42. The high degree of nucleotide sequence similarity between TFIIAγ5 and TFIIAγ1 is the main reason for causing the off-target of TALEN-Xa5 on the TFIIAγ1. In addition, unlike the RNAi and Cas9-edited TFIIAγ1 transgenic plants21,43, none of the four TALEN-edited TFIIAγ1 plants survived for the T1 generation; one died in the seedling stage and the others were sterile. The lower off-target efficiency and the random insertion of T-DNA may account for these. The lower off-target efficiency leads to the lower yield of TALEN-edited TFIIAγ1 plants, and the insertion of T-DNA can destroy some important genes to result in plant sterile. So the smaller sample capacity can not reflect the real effect of TALEN-edited TFIIAγ1. We will probe the TALEN-edited TFIIAγ1 based on its own sequence in the future.
Methods
Construction of the TALEN-Xa5 expression vectors
The target selection of TALEN-Xa5 considered some qualifications described by a series of researchers and employed a web-based tool called TAL Effector-Nucleotide Targeter 2.0 (TALE-NT 2.0; https://boglab.plp.iastate.edu/)44. In addition, we restricted the pair of target sequences to the first exon of the Xa5 gene.
The central repeat domain of TALEN was constructed and then inserted into a commercial TALEN skeleton (Sidansai, Shanghai, China) of clone plasmid using enzyme digestion and a ligation reaction. The two TALEN-Xa5 monomers were respectively generated using the different promoters 35S and Ubiquitin, while they employed the same nopaline synthase (NOS) terminator. Two cassettes of monomer genes were tandemly linked up by an AscI site, and then were inserted into the expression vector of pCAMBIA1300 (GenBank accession number AF234296.1) after the KpnI/SacI digestion. Generally, the construction of the TALEN-Xa5 vectors followed the instructions of the Sidansai commercial kit. Finally, the accuracy of the TALEN-Xa5 vector was examined by Sanger sequencing at each construction step.
Agrobacterium-mediated transformation of rice
The binary plasmid containing the TALEN-Xa5 in the pCAMBIA1300 was introduced into Agrobacterium by electroporation and then the Agrobacterium infection of calluses transmitted the expression vectors into rice.
Calluses of Japonica and Indica cultivar were generated from seeds and transformed by Agrobacterium tumefaciens LBA4404 and EHA105, respectively45. Four recipient rice cultivars were used in the experiment. TP309 belongs to O. sativa Japonica while MH86, D62B and IR24 belong to O. sativa Indica. Moreover, two transgenic lines of the gene Xa21 created by our lab were used for a combination test of the resistance loci of the gene Xa21 and xa5. The transgenic lines of CX8621 and CX6221 were the F4 or F5 generations of O. sativa cultivar MH86 and D62B, respectively. The primers Hpt-F and Hpt-R for the HPT gene were used to select the positive seedlings in the T0 generation (Table S1).
The detection of mutations by PCR/RE assay and sequencing in transgenic rice
The transgenic seedlings were maintained in sterile conditions and checked for mutations before transplantation to the soil. The mutation detection was carried out using the PCR/RE assay and was further proved using gene sequencing as described27. The genomic DNA of transgenic seedlings was extracted following the CTAB-DNA precipitation method, providing the template for amplifying the fragment containing the TALEN-Xa5 target site in the PCR/RE assay. The PCR primers were showed in Table S1 as Xa5F and Xa5R. The amplification product was then digested by two restriction endonucleases BbvCI and SacI. The sites of BbvCI and SacI were respectively located at each end of the space lying between the TALEN-Xa5 target pair. If the TALEN-Xa5 edited the two endonuclease sites, distinct gel electrophoresis strips would appear, indicating the occurrence of mutations. In some cases, however, mutations occurred beyond the space region, leading to the loss of mutants in the PCR/RE assay. Nevertheless, missing a mutant is rare, and the mutation rate dropped down sharply away from the TALEN cut site in the space region. The mutations proved by PCR/RE were then verified by sequencing.
Several types of mutations frequently existed in one seedling, especially for the first transgenic generation. In this case, each mutation band from the PCR/RE digestion was purified and cloned to into the pEASY-T1 vector, and then heat shock transformed competent cells of E. Coli before sequencing plasmids were conducted and five or six positive clones of each transformation were randomly selected for the Sanger sequencing. The occurrence of mutations indicated gene editing on the gene Xa5 in the transgenic seedlings. Only Xa5 mutants in the T0 seedlings were reserved to produce T1 generation.
Screening the separation of transgenic fragments in T1 generation
The genomic DNA from individual rice seedlings was extracted using the CTAB-DNA precipitation method. FokI genes in each monomer of the TALEN-Xa5 and the HPT gene in the expressing vector skeleton were screened using PCR to examine whether they had segregated. The detection primers for each gene were shown as Fok-F and Fok-R, Hpt-F and Hpt-R in Table S1. Notably, the mutations of gene Xa5 were checked by the PCR/RE assay in the seedlings of the T1 generation that tested negative for the FokI and HPT genes. Only Xa5 mutants negative for the FokI and HPT genes were maintained to produce T2 generation.
Screening homozygous mutants of the gene Xa5 in T2 generation
The PCR/RE assay and Sanger sequencing were used to recheck the Xa5 mutants within T2 generation. Five or more seedlings of every T2 line were randomly selected. Homozygous mutants identified by the PCR/RE assay were then to be sequenced. The PCR product from the homozygous mutant was firstly purified and cloned to into the pEASY-T1 vector, and then transformed into competent cells of E. Coli. Five or six positive clones from each transformation were selected randomly for the Sanger sequencing. All positive clones showing the same type of mutations indicated that a seedling was a homozygous mutant. Only T2 homozygotes for the gene Xa5 mutation were used to generate T3 homozygous lines. Additionally, some T2 lines from the homozygotes of the T1 mutants were already homozygous. It was important that all samplings from the same line showed the same mutation.
Inoculation test in homozygous lines of gene Xa5 mutations
Only homozygous lines were used in the inoculation test of the bacterial blight pathogen to assess the effect of gene Xa5 mutations on rice resistance, which may be exerted in the T2 or T3 generation. The inoculation test was carried out two times. The first time, rice was grown during the normal growing season, from April to October in 2015 in Beijing (39°N latitude), China, then inoculated in July, with data collection in August. The second time, rice was grown from November, 2015 to April, 2016 under field conditions in Lingshui (18°N latitude), Hainan, China, then inoculated in January with data collection in February.
For the resistance evaluation, the Xoo strains PXO86 and HN01 (isolated in Hainan) were cultivated on PSA medium (30% potato filtrate, 0.05% Ca(NO3)2 • 4H2O, 0.2% Na2HPO4 • 12H2O, 1.5% sucrose, and 1.5% agar) at 28 °C for 2 days and then re-suspended in sterile water with a dilution of OD600 nm = 0.5 for inoculation. The cryopreserved strains should be activated on the PSA medium at 28 °C for 2 days and subsequently transferred to a new PSA medium. At least 10 plants of each line were examined. For the resistance spectrum assay, the Xoo strains PXO86, PXO71, PXO112, PXO99, PXO145, PXO280 and HN01 were employed. At least five plants from each line were examined. Five or ten leaves of each plant were inoculated using the leaf-clipping method for the booting seedlings (during the panicle development). Disease severity was determined on the basis of lesion length on the 14th day after inoculation. Significant differences between wild type controls and transgenic plants as determined by a one-way ANOVA (p < 0.05) followed by post hoc Tukey HSD analysis. All of the experiments were repeated two times with similar results.
Detecting off-target effects on the homologous gene TFIIAγ1
Off-target effects on TFIIAγ1, another TFIIAγ gene in rice, were examined by PCR/RE. TFIIAγ1 is a paralog of rice TFIIAγ5 with 86% identity at the amino acid sequence level. The homologous off-target loci of the gene Xa5 were blasted in National Center for Biotechnology in formation (NCBI)(https://blast.ncbi.nlm.nih.gov/Blast.cgi). PCR was performed using the primer TFIIA-gamma-1F and TFIIA-gamma-1R (Table S1). The restriction endonuclease SacI was used to digest the amplification product. The normal product was digested into two fragments, one of 511 bp and the other 302 bp. The off-target effect of TALEN-Xa5 on TFIIAγ1 might cause abnormal digestion fragments for the disappearance of the digestion site within the loop between the two target recognition sequences, providing a convenient way to evaluate the accuracy of the editing of TALEN-Xa5.
References
Boch, J. & Bonas, U. Xanthomonas AvrBs3 Family-Type III Effectors: Discovery and Function. Annu. Rev. Phytopathology 48, 419–436 (2010).
Doyle, E. L., Stoddard, B. L., Voytas, D. F. & Bogdanove, A. J. TAL effectors: highly adaptable phytobacterial virulence factors and readily engineered DNA-targeting proteins. Trends Cell Biol. 23, 390–398 (2013).
Ji, Z. Y. et al. Interfering TAL effectors of Xanthomonas oryzae neutralize R-gene-mediated plant disease resistance. Nat. Commun. 7, 13435, https://doi.org/10.1038/ncomms13435 (2016).
Read, A. C. et al. Suppression of Xo1-Mediated Disease Resistance in Rice by a Truncated, Non-DNA-Binding TAL Effector of Xanthomonas oryzae. Front. Plant. Sci. 7, 1516, https://doi.org/10.3389/fpls.2016.01516 (2016).
Tian, D. S. et al. The Rice TAL Effector-Dependent Resistance Protein XA10 Triggers Cell Death and Calcium Depletion in the Endoplasmic Reticulum. Plant. Cell 26, 497–515 (2014).
Gu, K. Y. et al. R gene expression induced by a type-III effector triggers disease resistance in rice. Nat. 435, 1122–1125 (2005).
Wang, C. L. et al. XA23 Is an Executor R Protein and Confers Broad-Spectrum Disease Resistance in Rice. Mol. Plant. 8, 290–302 (2015).
Kim, S. M. et al. Identification and fine-mapping of a new resistance gene, Xa40, conferring resistance to bacterial blight races in rice (Oryza sativa L.). Theor. Appl. Genet. 128, 1933–1943 (2015).
Song, W. Y. et al. A Receptor Kinase-Like Protein Encoded by the Rice Disease Resistance Gene, Xa21. Sci. 270, 1804–1806 (1995).
Yoshimura, S. et al. Expression of Xa1, a bacterial blight-resistance gene in rice, is induced by bacterial inoculation. Proc. Natl. Acad. Sci. USA 95, 1663–1668 (1998).
Sun, X. L. et al. Xa26, a gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an LRR receptor kinase-like protein. Plant. J. 37, 517–527 (2004).
Iyer-Pascuzzi, A. S., Jiang, H., Huang, L. & McCouch, S. R. Genetic and functional characterization of the rice bacterial blight disease resistance gene xa5. Phytopathology 98, 289–295 (2008).
Jiang, G. H. et al. Testifying the rice bacterial blight resistance gene xa5 by genetic complementation and further analyzing xa5 (Xa5) in comparison with its homolog TFIIAgamma1. Mol. Genet. Genomics 275, 354–366 (2006).
Chu, Z. et al. Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes. Dev. 20, 1250–1255 (2006).
Hu, K. et al. Improvement of multiple agronomic traits by a disease resistance gene via cell wall reinforcement. Nat. Plants 3, 17009, https://doi.org/10.1038/nplants.2017.9 (2017).
Hutin, M., Sabot, F., Ghesquiere, A., Koebnik, R. & Szurek, B. A knowledge-based molecular screen uncovers a broad-spectrum OsSWEET14 resistance allele to bacterial blight from wild rice. Plant. J. 84, 694–703 (2015).
Yang, B., Sugio, A. & White, F. F. Os8N3 is a host disease-susceptibility gene for bacterial blight of rice. Proc. Natl Acad. Sci. U S Am. 103, 10503–10508 (2006).
Liu, Q. S. et al. A paralog of the MtN3/saliva family recessively confers race-specific resistance to Xanthomonas oryzae in rice. Plant. Cell Environ. 34, 1958–1969 (2011).
Zhou, J. H. et al. Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice. Plant. J. 82, 632–643 (2015).
Li, Y. F. et al. Characterization and functional analysis of Arabidopsis TFIIA reveal that the evolutionarily unconserved region of the large subunit has a transcription activation domain. Plant. Mol. Biol. 39, 515–525 (1999).
Yuan, M. et al. A host basal transcription factor is a key component for infection of rice by TALE-carrying bacteria. Elife. 5, e19605, https://doi.org/10.7554/eLife.19605 (2016).
Gu, K. Y., Tian, D. S., Qiu, C. X. & Yin, Z. C. Transcription activator-like type III effector AvrXa27 depends on OsTFIIA gamma 5 for the activation of Xa27 transcription in rice that triggers disease resistance to Xanthomonas oryzae pv. oryzae. Mol. Plant. Pathol. 10, 829–835 (2009).
Iyer, A. S. & McCouch, S. R. The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol. Plant. Microbe Interact. 17, 1348–1354 (2004).
Huang, S. et al. The broadly effective recessive resistance gene xa5 of rice is a virulence effector-dependent quantitative trait for bacterial blight. Plant. J. 86, 186–194 (2016).
Huang, N. et al. Pyramiding of bacterial blight resistance genes in rice: marker-aided selection using RFLP and PCR. Theor. Appl. Genet. 95, 313–320 (1997).
Garris, A. J., McCouch, S. R. & Kresovich, S. Population structure and its effect on haplotype diversity and linkage disequilibrium surrounding the xa5 locus of rice (Oryza sativa L.). Genet. 165, 759–769 (2003).
Miller, J. C. et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotechnol. 29, 143–148 (2011).
Szurek, B., Rossier, O., Hause, G. & Bonas, U. Type III-dependent translocation of the Xanthomonas AvrBs3 protein into the plant cell. Mol. Microbiology 46, 13–23 (2002).
Schornack, S., Moscou, M. J., Ward, E. R. & Horvath, D. M. Engineering Plant Disease Resistance Based on TAL Effectors. Annu. Rev. Phytopathology 51, 383–406 (2013).
Streubel, J., Baum, H., Grau, J., Stuttman, J. & Boch, J. Dissection of TALE-dependent gene activation reveals that they induce transcription cooperatively and in both orientations. PLoS One 12, e0173580, https://doi.org/10.1371/journal.pone.0173580 (2017).
Boch, J. et al. Breaking the Code of DNA Binding Specificity of TAL-Type III Effectors. Sci. 326, 1509–1512 (2009).
Moscou, M. J. & Bogdanove, A. J. A Simple Cipher Governs DNA Recognition by TAL Effectors. Sci. 326, 1501–1501 (2009).
Deng, D. et al. Structural Basis for Sequence-Specific Recognition of DNA by TAL Effectors. Sci. 335, 720–723 (2012).
Mak, A. N. S., Bradley, P., Cernadas, R. A., Bogdanove, A. J. & Stoddard, B. L. The Crystal Structure of TAL Effector PthXo1 Bound to Its DNA Target. Sci. 335, 716–719 (2012).
de Lange, O., Binder, A. & Lahaye, T. From dead leaf, to new life: TAL effectors as tools for synthetic biology. Plant. J. 78, 753–771 (2014).
Streubel, J., Blucher, C., Landgraf, A. & Boch, J. TAL effector RVD specificities and efficiencies. Nat. Biotechnol. 30, 593–595 (2012).
Kim, H. & Kim, J. S. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 15, 321–334 (2014).
Sedeek, K. E. M., Mahas, A. & Mahfouz, M. Plant Genome Engineering for Targeted Improvement of Crop Traits. Front. Plant. Sci. 10, 114, https://doi.org/10.3389/fpls.2019.00114 (2019).
Gao, L. et al. Do transgenesis and marker-assisted backcross breeding produce substantially equivalent plants? A comparative study of transgenic and backcross rice carrying bacterial blight resistant gene Xa21. BMC Genomics 14, 738, https://doi.org/10.1186/1471-2164-14-738 (2013).
Li, T. et al. Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes. Nucleic Acids Res. 39, 6315–6325 (2011).
Ding, Q. et al. A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell 12, 238–251 (2013).
Sun, H. Z. & Ge, S. Molecular evolution of the duplicated TFIIAgamma genes in Oryzeae and its relatives. BMC Evol. Biol. 10, 128, https://doi.org/10.1186/1471-2148-10-128 (2010).
Ma, W. et al. Xanthomonas oryzae pv. oryzae TALE proteins recruit OsTFIIAgamma1 to compensate for the absence of OsTFIIAgamma5 in bacterial blight in rice. Mol. plant. Pathol. 19, 2248–2262 (2018).
Doyle, E. L. et al. TAL Effector-Nucleotide Targeter (TALE-NT) 2.0: tools for TAL effector design and target prediction. Nucleic Acids Res. 40, W117–W122 (2012).
Nishimura, A., Aichi, I. & Matsuoka, M. A protocol for Agrobacterium-mediated transformation in rice. Nat. Protocols. 1, 2796–2802 (2006).
Acknowledgements
This research was supported by the grants from the Ministry of Science and Technology of China (No. 2016YFD0101801), the Ministry of Agriculture of China (No. 2016ZX08001002 and 2016ZX08009003), the National Natural Science Foundation of China (No. 31571245 and 31900383), and the Postdoctoral Science Foundation of China (No. 2019M660853).
Author information
Authors and Affiliations
Contributions
W. Zhai, G. Jiang and Z. Xia designed the experiments. J. Han, P. Liu, C. Li, Y. Wang and L. Guo performed the experiments and analyzed the dates. J. Han, G. Jiang and W. Zhai wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Dr. Xia’s work has been funded by the National Natural Science Foundation of China (No. 31071173). He has received compensation as a member of Hainan University of China. Dr. Han, Dr. Jiang and Prof. Zhai declare no potential conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, J., Xia, Z., Liu, P. et al. TALEN-based editing of TFIIAy5 changes rice response to Xanthomonas oryzae pv. Oryzae. Sci Rep 10, 2036 (2020). https://doi.org/10.1038/s41598-020-59052-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-59052-w
This article is cited by
-
Rapid generation of fragrant thermo-sensitive genic male sterile rice with enhanced disease resistance via CRISPR/Cas9
Planta (2024)
-
A review of approaches to control bacterial leaf blight in rice
World Journal of Microbiology and Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.