Abstract
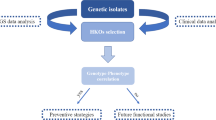
A major goal of biomedicine is to understand the function of every gene in the human genome1. Loss-of-function mutations can disrupt both copies of a given gene in humans and phenotypic analysis of such ‘human knockouts’ can provide insight into gene function. Consanguineous unions are more likely to result in offspring carrying homozygous loss-of-function mutations. In Pakistan, consanguinity rates are notably high2. Here we sequence the protein-coding regions of 10,503 adult participants in the Pakistan Risk of Myocardial Infarction Study (PROMIS), designed to understand the determinants of cardiometabolic diseases in individuals from South Asia3. We identified individuals carrying homozygous predicted loss-of-function (pLoF) mutations, and performed phenotypic analysis involving more than 200 biochemical and disease traits. We enumerated 49,138 rare (<1% minor allele frequency) pLoF mutations. These pLoF mutations are estimated to knock out 1,317 genes, each in at least one participant. Homozygosity for pLoF mutations at PLA2G7 was associated with absent enzymatic activity of soluble lipoprotein-associated phospholipase A2; at CYP2F1, with higher plasma interleukin-8 concentrations; at TREH, with lower concentrations of apoB-containing lipoprotein subfractions; at either A3GALT2 or NRG4, with markedly reduced plasma insulin C-peptide concentrations; and at SLC9A3R1, with mediators of calcium and phosphate signalling. Heterozygous deficiency of APOC3 has been shown to protect against coronary heart disease4,5; we identified APOC3 homozygous pLoF carriers in our cohort. We recruited these human knockouts and challenged them with an oral fat load. Compared with family members lacking the mutation, individuals with APOC3 knocked out displayed marked blunting of the usual post-prandial rise in plasma triglycerides. Overall, these observations provide a roadmap for a ‘human knockout project’, a systematic effort to understand the phenotypic consequences of complete disruption of genes in humans.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Eisenberg, D., Marcotte, E. M., Xenarios, I. & Yeates, T. O. Protein function in the post-genomic era. Nature 405, 823–826 (2000)
Bittles, A. H., Mason, W. M., Greene, J. & Rao, N. A. Reproductive behavior and health in consanguineous marriages. Science 252, 789–794 (1991)
Saleheen, D. et al. The Pakistan Risk of Myocardial Infarction Study: a resource for the study of genetic, lifestyle and other determinants of myocardial infarction in South Asia. Eur. J. Epidemiol. 24, 329–338 (2009)
Crosby, J. et al. Loss-of-function mutations in APOC3, triglycerides, and coronary disease. N. Engl. J. Med. 371, 22–31 (2014)
Jørgensen, A. B., Frikke-Schmidt, R., Nordestgaard, B. G. & Tybjærg-Hansen, A. Loss-of-function mutations in APOC3 and risk of ischemic vascular disease. N. Engl. J. Med. 371, 32–41 (2014)
Narasimhan, V. M. et al. Health and population effects of rare gene knockouts in adult humans with related parents. Science 352, 474–477 (2016)
Sulem, P. et al. Identification of a large set of rare complete human knockouts. Nat. Genet. 47, 448–452 (2015)
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016)
Di Angelantonio, E. et al. Lipid-related markers and cardiovascular disease prediction. J. Am. Med. Assoc. 307, 2499–2506 (2012)
Gregson, J. M. et al. Genetic invalidation of Lp-Pla2 as a therapeutic target: large-scale study of five functional Lp-Pla2-lowering alleles. Eur. J. Prev. Cardiol. (2016)
Polfus, L. M., Gibbs, R. A. & Boerwinkle, E. Coronary heart disease and genetic variants with low phospholipase A2 activity. N. Engl. J. Med. 372, 295–296 (2015)
White, H. D. et al. Darapladib for preventing ischemic events in stable coronary heart disease. N. Engl. J. Med. 370, 1702–1711 (2014)
O’Donoghue, M. L. et al. Effect of darapladib on major coronary events after an acute coronary syndrome: the SOLID-TIMI 52 randomized clinical trial. J. Am. Med. Assoc. 312, 1006–1015 (2014)
Carr, B. A., Wan, J., Hines, R. N. & Yost, G. S. Characterization of the human lung CYP2F1 gene and identification of a novel lung-specific binding motif. J. Biol. Chem. 278, 15473–15483 (2003)
Standiford, T. J. et al. Interleukin-8 gene expression by a pulmonary epithelial cell line. A model for cytokine networks in the lung. J. Clin. Invest. 86, 1945–1953 (1990)
Murray, I. A., Coupland, K., Smith, J. A., Ansell, I. D. & Long, R. G. Intestinal trehalase activity in a UK population: establishing a normal range and the effect of disease. Br. J. Nutr. 83, 241–245 (2000)
Christiansen, D. et al. Humans lack iGb3 due to the absence of functional iGb3-synthase: implications for NKT cell development and transplantation. PLoS Biol. 6, e172 (2008)
Dahl, K., Buschard, K., Gram, D. X., d’Apice, A. J. & Hansen, A. K. Glucose intolerance in a xenotransplantation model: studies in alpha-gal knockout mice. APMIS 114, 805–811 (2006)
Casu, A. et al. Insulin secretion and glucose metabolism in alpha 1,3-galactosyltransferase knock-out pigs compared to wild-type pigs. Xenotransplantation 17, 131–139 (2010)
Schneider, M. R. & Wolf, E. The epidermal growth factor receptor ligands at a glance. J. Cell. Physiol. 218, 460–466 (2009)
Wang, G. X. et al. The brown fat-enriched secreted factor Nrg4 preserves metabolic homeostasis through attenuation of hepatic lipogenesis. Nat. Med. 20, 1436–1443 (2014)
Murtazina, R. et al. Tissue-specific regulation of sodium/proton exchanger isoform 3 activity in Na+/H+ exchanger regulatory factor 1 (NHERF1) null mice. cAMP inhibition is differentially dependent on NHERF1 and exchange protein directly activated by cAMP in ileum versus proximal tubule. J. Biol. Chem. 282, 25141–25151 (2007)
Karim, Z. et al. NHERF1 mutations and responsiveness of renal parathyroid hormone. N. Engl. J. Med. 359, 1128–1135 (2008)
Huff, M. W. & Hegele, R. A. Apolipoprotein C-III: going back to the future for a lipid drug target. Circ. Res. 112, 1405–1408 (2013)
Pollin, T. I. et al. A null mutation in human APOC3 confers a favorable plasma lipid profile and apparent cardioprotection. Science 322, 1702–1705 (2008)
Gaudet, D. et al. Antisense inhibition of apolipoprotein C-III in patients with hypertriglyceridemia. N. Engl. J. Med. 373, 438–447 (2015)
Gaudet, D. et al. Targeting APOC3 in the familial chylomicronemia syndrome. N. Engl. J. Med. 371, 2200–2206 (2014)
Graham, M. J. et al. Antisense oligonucleotide inhibition of apolipoprotein C-III reduces plasma triglycerides in rodents, nonhuman primates, and humans. Circ. Res. 112, 1479–1490 (2013)
Brown, S. D. & Moore, M. W. Towards an encyclopaedia of mammalian gene function: the International Mouse Phenotyping Consortium. Dis. Model. Mech. 5, 289–292 (2012)
Scott, E. M. et al. Characterization of Greater Middle Eastern genetic variation for enhanced disease gene discovery. Nat. Genet. 48, 1071–1076 (2016)
Kooner, J. S. et al. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat. Genet. 43, 984–989 (2011)
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007)
Tennessen, J. A. et al. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 337, 64–69 (2012)
Do, R. et al. Exome sequencing identifies rare LDLR and APOA5 alleles conferring risk for myocardial infarction. Nature 518, 102–106 (2015)
Fisher, S. et al. A scalable, fully automated process for construction of sequence-ready human exome targeted capture libraries. Genome Biol. 12, R1 (2011)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011)
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics http://dx.doi.org/10.1002/0471250953.bi1110s43 (2013)
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26, 2069–2070 (2010)
Karczewski, K. J. Loftee (Loss-of-Function Transcript Effect Estimator), https://github.com/konradjk/loftee (2015)
Jun, G. et al. Detecting and estimating contamination of human DNA samples in sequencing and array-based genotype data. Am. J. Hum. Genet. 91, 839–848 (2012)
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010)
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS One 5, e15004 (2010)
Hunter-Zinck, H. et al. Population genetic structure of the people of Qatar. Am. J. Hum. Genet. 87, 17–25 (2010)
Lander, E. S. & Botstein, D. Homozygosity mapping: a way to map human recessive traits with the DNA of inbred children. Science 236, 1567–1570 (1987)
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014)
Wright, S. Coefficients of Inbreeding and Relationship. Am. Nat. 56, 330–338 (1922)
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014)
Samocha, K. E. et al. A framework for the interpretation of de novo mutation in human disease. Nat. Genet. 46, 944–950 (2014)
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015)
Eppig, J. T., Blake, J. A., Bult, C. J., Kadin, J. A. & Richardson, J. E. The Mouse Genome Database (MGD): facilitating mouse as a model for human biology and disease. Nucleic Acids Res. 43, D726–D736 (2015)
Georgi, B., Voight, B. F. & Bućan, M. From mouse to human: evolutionary genomics analysis of human orthologs of essential genes. PLoS Genet. 9, e1003484 (2013)
Fuchs, M. et al. The p400 complex is an essential E1A transformation target. Cell 106, 297–307 (2001)
Fazzio, T. G., Huff, J. T. & Panning, B. An RNAi screen of chromatin proteins identifies Tip60-p400 as a regulator of embryonic stem cell identity. Cell 134, 162–174 (2008)
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006)
Sambrook, J. & Russell, D. W. Purification of nucleic acids by extraction with phenol:chloroform. CSH Protoc. http://dx.doi.org/10.1101/pdb.prot4455 (2006)
Mosteller, R. D. Simplified calculation of body-surface area. N. Engl. J. Med. 317, 1098 (1987)
Maraki, M. et al. Validity of abbreviated oral fat tolerance tests for assessing postprandial lipemia. Clin. Nutr. 30, 852–857 (2011)
Acknowledgements
D.S. is supported by grants from the National Institutes of Health, the Fogarty International, the Wellcome Trust, the British Heart Foundation, and Pfizer. P.N. is supported by the John S. LaDue Memorial Fellowship in Cardiology from Harvard Medical School. H.-H.W. is supported by a grant from the Samsung Medical Center, Korea (SMO116163). S.K. is supported by the Ofer and Shelly Nemirovsky MGH Research Scholar Award and by grants from the National Institutes of Health (R01HL107816), the Donovan Family Foundation, and Fondation Leducq. Exome sequencing was supported by a grant from the NHGRI (5U54HG003067-11) to S.G. and E.S.L. D.G.M. is supported by a grant from the National Institutes of Health (R01GM104371). J.D. holds a British Heart Foundation Chair, European Research Council Senior Investigator Award, and NIHR Senior Investigator Award. The Cardiovascular Epidemiology Unit at the University of Cambridge, which supported the field work and genotyping of PROMIS, is funded by the UK Medical Research Council, British Heart Foundation, and NIHR Cambridge Biomedical Research Centre. In recognition for PROMIS fieldwork and support, we also acknowledge contributions made by the following: M. Z. Ozair, U. Ahmed, A. Hakeem, H. Khalid, K. Shahid, F. Shuja, A. Kazmi, M. Qadir Hameed, N. Khan, S. Khan, A. Ali, M. Ali, S. Ahmed, M. W. Khan, M. R. Khan, A. Ghafoor, M. Alam, R. Ahmed, M. I. Javed, A. Ghaffar, T. B. Mirza, M. Shahid, J. Furqan, M. I. Abbasi, T. Abbas, R. Zulfiqar, M. Wajid, I. Ali, M. Ikhlaq, D. Sheikh, M. Imran, M. Walker, N. Sarwar, S. Venorman, R. Young, A. Butterworth, H. Lombardi, B. Kaur and N. Sheikh. Fieldwork in the PROMIS study has been supported through funds available to investigators at the Center for Non-Communicable Diseases, Pakistan and the University of Cambridge, UK.
Author information
Authors and Affiliations
Contributions
Sample recruitment and phenotyping was performed by D.S., P.F., J.D., A.R., M.Z., M.S., A.I., S.A., F.Ma., M.I., S.A., K.T., N.H.M., K.S.Z., N.Q., M.I., S.Z.R., F.Me., K.M., N.A., and R.M.K. D.S., P.F., J.D., and W.Z. performed array-based genotyping and runs-of-homozygosity analyses. Exome sequencing was coordinated by D.S., N.G., S.G., E.S.L., D.J.R., and S.K. P.N., W.Z., H.H.W., and R.D. performed exome-sequencing quality control and association analyses. P.N., I.M.A., K.J.K., A.H.O., B.W., and D.G.M. performed variant annotation. D.S., S.K., and D.J.R. performed confirmatory genotyping and lipoprotein biomarker assays. D.S. and A.R. conducted recall-based studies for the APOC3 knockouts. P.N. and M.J.D. performed bioinformatics simulations. P.N. and K.E.S. performed constraint score analyses. D.S., P.N., and S.K. designed the study and wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 pLoF mutations are typically seen in very few individuals.
The site-frequency spectrum of synonymous, missense, and high-confidence pLoF mutations is represented. Points represent the proportion of variants within a 1 × 10−4 minor allele frequency bin for each variant category. Lines represent the cumulative proportions of variants categories. The bottom inset highlights that most pLoF variants are often seen in no more than one or two individuals. The top inset highlights that virtually all pLoF mutations are very rare.
Extended Data Figure 2 Intersection of homozygous pLoF genes between PROMIS and other cohorts.
We compared the counts and overlap of unique homozygous pLoF genes in PROMIS with other exome sequenced cohorts.
Extended Data Figure 3 QQ-plot of recessive model pLoF association analysis across phenotypes.
Analyses to determine whether homozygous pLoF carrier status was associated with traits was performed where there were at least two homozygous pLoF carriers phenotyped per trait. The observed versus the expected results from 15,263 associations are displayed here demonstrating an excess of associations beyond a Bonferroni threshold.
Extended Data Figure 4 Carriers of pLoF alleles in CYP2F1 have increased IL-8 concentrations.
Participants who had pLoF mutations in the CYP2F1 gene had higher concentrations of IL-8, whereas heterozygotes had a more modest effect when compared to the rest of the cohort of non-carriers. IL-8 concentration is natural log transformed. Bars represent 1.5× interquartile range beyond the 25th and 75th percentiles.
Extended Data Figure 5 Carriers of pLoF alleles in TREH have decreased concentrations of several lipoprotein subfractions.
Participants who had pLoF mutations in the TREH gene had lower concentrations of several lipoprotein subfractions. Bars represent 1.5× interquartile range beyond the 25th and 75th percentiles.
Extended Data Figure 6 Nondiabetic homozygous pLoF carriers for A3GALT2 have diminished insulin C-peptide concentrations.
Among nondiabetics, those who were homozygous pLoF for A3GALT2 had substantially lower fasting insulin C-peptide concentrations. This observation was not evident in nondiabetic heterozygous pLoF A3GALT2 participants. Insulin C-peptide is natural log transformed. Bars represent 1.5× interquartile range beyond the 25th and 75th percentiles.
Extended Data Figure 7 Example of a second polymorphism in-phase which rescues a putative protein-truncating mutation.
Short-reads that align to genomic positions 65,339,112 to 65,339,132 on chromosome 1 are displayed for one individual with a putative homozygous pLoF genotype in this region. The SNP at position 65,339,122 from G to T is annotated as a nonsense mutation in the JAK1 gene. However, all three homozygotes of this mutation carried a tandem SNP in the same codon (A to G at 65,339,124) thus resulting in a glutamine and effectively rescuing the protein-truncating mutation.
Extended Data Figure 8 Anticipated number of genes knocked out with increasing sample sizes by minimum knockout count.
We simulate the number of genes expected to be knocked out by minimum knockout count per gene at increasing sample sizes. We perform this simulation with and without the observed inbreeding.
Extended Data Figure 9 PROMIS participants have an excess burden of runs of homozygosity compared with other populations.
Consanguinity leads to regions of genomic segments that are identical by descent and can be observed as runs of homozygosity. Using genome-wide array data in 17,744 PROMIS participants and reference samples from the International HapMap3, the burden of runs of homozygosity (minimum 1.5 Mb) per individual was derived and population-specific distributions are displayed, with outliers removed. This highlights the higher median runs of homozygosity burden in PROMIS and the higher proportion of individuals with very high burdens.
Extended Data Figure 10 Down-sampling of synonymous and high confidence pLoF variants to validate simulation.
a, b, We ran simulations to estimate the number of unique, completely knocked out genes at increasing sample sizes. Before applying our model, we first applied this approach to a range of sample sizes below 7,078 for variants that were not under constraint, synonymous variants (a), and for high-confidence null variants (b). At the observed sample size, we did not observe significant selection. We expect that at increasing sample sizes, there may be a subset of genes that will not be tolerated in a homozygous pLoF state. In fact, our estimates are slightly more conservative when comparing outbred simulations with a recent description of >100,000 Icelanders using a more liberal definition for pLoF mutations.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-9, the full legend for Supplementary Table 1 (supplied as a separate spreadsheet) and Supplementary References. (PDF 621 kb)
Supplementary Table
This file contains Supplementary Table 1 – see the Supplementary Information document for the full description. (XLSX 241 kb)
Rights and permissions
About this article
Cite this article
Saleheen, D., Natarajan, P., Armean, I. et al. Human knockouts and phenotypic analysis in a cohort with a high rate of consanguinity. Nature 544, 235–239 (2017). https://doi.org/10.1038/nature22034
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22034
This article is cited by
-
Comparative study on genomic and epigenomic profiles of retinoblastoma or tuberous sclerosis complex via nanopore sequencing and a joint screening framework
Cancer Gene Therapy (2024)
-
A novel mouse model of familial combined hyperlipidemia and atherosclerosis
Acta Pharmacologica Sinica (2024)
-
The Role of Genetics in Advancing Cardiometabolic Drug Development
Current Atherosclerosis Reports (2024)
-
Burden of Mendelian disorders in a large Middle Eastern biobank
Genome Medicine (2024)
-
Exploring the complex spectrum of dominance and recessiveness in genetic cardiomyopathies
Nature Cardiovascular Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.