Abstract
The retina extracts visual features for transmission to the brain. Different types of bipolar cell split the photoreceptor input into parallel channels and provide the excitatory drive for downstream visual circuits. Mouse bipolar cell types have been described at great anatomical and genetic detail, but a similarly deep understanding of their functional diversity is lacking. Here, by imaging light-driven glutamate release from more than 13,000 bipolar cell axon terminals in the intact retina, we show that bipolar cell functional diversity is generated by the interplay of dendritic excitatory inputs and axonal inhibitory inputs. The resulting centre and surround components of bipolar cell receptive fields interact to decorrelate bipolar cell output in the spatial and temporal domains. Our findings highlight the importance of inhibitory circuits in generating functionally diverse excitatory pathways and suggest that decorrelation of parallel visual pathways begins as early as the second synapse of the mouse visual system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Masland, R. H. The neuronal organization of the retina. Neuron 76, 266–280 (2012)
Baden, T. et al. The functional diversity of retinal ganglion cells in the mouse. Nature 529, 345–350 (2016)
Sanes, J. R. & Masland, R. H. The types of retinal ganglion cells: current status and implications for neuronal classification. Annu. Rev. Neurosci. 38, 221–246 (2015)
Euler, T., Haverkamp, S., Schubert, T. & Baden, T. Retinal bipolar cells: elementary building blocks of vision. Nat. Rev. Neurosci. 15, 507–519 (2014)
Helmstaedter, M. et al. Connectomic reconstruction of the inner plexiform layer in the mouse retina. Nature 500, 168–174 (2013)
Kim, J. S. et al. Space–time wiring specificity supports direction selectivity in the retina. Nature 509, 331–336 (2014)
Behrens, C., Schubert, T., Haverkamp, S., Euler, T. & Berens, P. Connectivity map of bipolar cells and photoreceptors in the mouse retina. eLife 5, e20041 (2016)
Wässle, H., Puller, C., Müller, F. & Haverkamp, S. Cone contacts, mosaics, and territories of bipolar cells in the mouse retina. J. Neurosci. 29, 106–117 (2009)
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016)
Greene, M. J., Kim, J. S. & Seung, H. S. Analogous convergence of sustained and transient inputs in parallel on and off pathways for retinal motion computation. Cell Reports 14, 1892–1900 (2016)
Awatramani, G. B. & Slaughter, M. M. Origin of transient and sustained responses in ganglion cells of the retina. J. Neurosci. 20, 7087–7095 (2000)
Breuninger, T., Puller, C., Haverkamp, S. & Euler, T. Chromatic bipolar cell pathways in the mouse retina. J. Neurosci. 31, 6504–6517 (2011)
Euler, T., Schneider, H. & Wässle, H. Glutamate responses of bipolar cells in a slice preparation of the rat retina. J. Neurosci. 16, 2934–2944 (1996)
DeVries, S. H. Bipolar cells use kainate and AMPA receptors to filter visual information into separate channels. Neuron 28, 847–856 (2000)
DeVries, S. H., Li, W. & Saszik, S. Parallel processing in two transmitter microenvironments at the cone photoreceptor synapse. Neuron 50, 735–748 (2006)
Lindstrom, S. H., Ryan, D. G., Shi, J. & DeVries, S. H. Kainate receptor subunit diversity underlying response diversity in retinal off bipolar cells. J. Physiol. (Lond.) 592, 1457–1477 (2014)
Puller, C., Ivanova, E., Euler, T., Haverkamp, S. & Schubert, T. OFF bipolar cells express distinct types of dendritic glutamate receptors in the mouse retina. Neuroscience 243, 136–148 (2013)
Puthussery, T. et al. Kainate receptors mediate synaptic input to transient and sustained OFF visual pathways in primate retina. J. Neurosci. 34, 7611–7621 (2014)
Masland, R. H. The tasks of amacrine cells. Vis. Neurosci. 29, 3–9 (2012)
Eggers, E. D. & Lukasiewicz, P. D. Multiple pathways of inhibition shape bipolar cell responses in the retina. Vis. Neurosci. 28, 95–108 (2011)
Grimes, W. N., Li, W., Chávez, A. E. & Diamond, J. S. BK channels modulate pre- and postsynaptic signaling at reciprocal synapses in retina. Nat. Neurosci. 12, 585–592 (2009)
Demb, J. B. & Singer, J. H. Intrinsic properties and functional circuitry of the AII amacrine cell. Vis. Neurosci. 29, 51–60 (2012)
Marvin, J. S. et al. An optimized fluorescent probe for visualizing glutamate neurotransmission. Nat. Methods 10, 162–170 (2013)
Borghuis, B. G., Marvin, J. S., Looger, L. L. & Demb, J. B. Two-photon imaging of nonlinear glutamate release dynamics at bipolar cell synapses in the mouse retina. J. Neurosci. 33, 10972–10985 (2013)
Chen, M., Lee, S., Park, S. J. H., Looger, L. L. & Zhou, Z. J. Receptive field properties of bipolar cell axon terminals in direction-selective sublaminas of the mouse retina. J. Neurophysiol. 112, 1950–1962 (2014)
Rosa, J. M., Ruehle, S., Ding, H. & Lagnado, L. Crossover inhibition generates sustained visual responses in the inner retina. Neuron 90, 308–319 (2016)
Dreosti, E., Esposti, F., Baden, T. & Lagnado, L. In vivo evidence that retinal bipolar cells generate spikes modulated by light. Nat. Neurosci. 14, 951–952 (2011)
Baden, T., Berens, P., Bethge, M. & Euler, T. Spikes in mammalian bipolar cells support temporal layering of the inner retina. Curr. Biol. 23, 48–52 (2013)
Burrone, J. & Lagnado, L. Synaptic depression and the kinetics of exocytosis in retinal bipolar cells. J. Neurosci. 20, 568–578 (2000)
Nikolaev, A., Leung, K.-M., Odermatt, B. & Lagnado, L. Synaptic mechanisms of adaptation and sensitization in the retina. Nat. Neurosci. 16, 934–941 (2013)
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013)
Werblin, F. S. & Dowling, J. E. Organization of the retina of the mudpuppy, Necturus maculosus. II. Intracellular recording. J. Neurophysiol. 32, 339–355 (1969)
Roska, B. & Werblin, F. Vertical interactions across ten parallel, stacked representations in the mammalian retina. Nature 410, 583–587 (2001)
Lukasiewicz, P. D. & Werblin, F. S. A novel GABA receptor modulates synaptic transmission from bipolar to ganglion and amacrine cells in the tiger salamander retina. J. Neurosci. 14, 1213–1223 (1994)
Euler, T. & Masland, R. H. Light-evoked responses of bipolar cells in a mammalian retina. J. Neurophysiol. 83, 1817–1829 (2000)
Euler, T. & Wässle, H. Different contributions of GABAA and GABAC receptors to rod and cone bipolar cells in a rat retinal slice preparation. J. Neurophysiol. 79, 1384–1395 (1998)
Vaney, D. I. The mosaic of amacrine cells in the mammalian retina. Prog. Retinal Res. 9, 49–100 (1990)
Eggers, E. D. & Lukasiewicz, P. D. Interneuron circuits tune inhibition in retinal bipolar cells. J. Neurophysiol. 103, 25–37 (2010)
Roska, B., Nemeth, E., Orzo, L. & Werblin, F. S. Three levels of lateral inhibition: A space–time study of the retina of the tiger salamander. J. Neurosci. 20, 1941–1951 (2000)
Ichinose, T. & Lukasiewicz, P. D. Inner and outer retinal pathways both contribute to surround inhibition of salamander ganglion cells. J. Physiol. (Lond.) 565, 517–535 (2005)
Buldyrev, I. & Taylor, W. R. Inhibitory mechanisms that generate centre and surround properties in ON and OFF brisk-sustained ganglion cells in the rabbit retina. J. Physiol. (Lond.) 591, 303–325 (2013)
Liang, Z. & Freed, M. A. The ON pathway rectifies the OFF pathway of the mammalian retina. J. Neurosci. 30, 5533–5543 (2010)
Zaghloul, K. A., Boahen, K. & Demb, J. B. Different circuits for ON and OFF retinal ganglion cells cause different contrast sensitivities. J. Neurosci. 23, 2645–2654 (2003)
Atick, J. J. & Redlich, A. N. Towards a theory of early visual processing. Neural Comput. 320, 1–13 (1990)
Barlow, H. Possible principles underlying the transformations of sensory messages. Sensory Comm. 6, 57–58 (1961)
Pitkow, X. & Meister, M. Decorrelation and efficient coding by retinal ganglion cells. Nat. Neurosci. 15, 628–635 (2012)
Vinje, W. E. & Gallant, J. L. Sparse coding and decorrelation in primary visual cortex during natural vision. Science 1273, 1273–1276 (2007)
Giridhar, S., Doiron, B. & Urban, N. N. Timescale-dependent shaping of the correlation by olfactory bulb lateral inhibition. Proc. Natl Acad. Sci. USA 108, 5843–5848 (2011)
Denève, S. & Machens, C. K. Efficient codes and balanced networks. Nat. Neurosci. 19, 375–382 (2016)
Kemmler, R., Schultz, K., Dedek, K., Euler, T. & Schubert, T. Differential regulation of cone calcium signals by different horizontal cell feedback mechanisms in the mouse retina. J. Neurosci. 34, 11826–11843 (2014)
Schubert, T. et al. Development of presynaptic inhibition onto retinal bipolar cell axon terminals is subclass-specific. J. Neurophysiol. 100, 304–316 (2008)
Euler, T. et al. Eyecup scope—optical recordings of light stimulus-evoked fluorescence signals in the retina. Pflugers Arch. 457, 1393–1414 (2009)
Baden, T. et al. A tale of two retinal domains: near-optimal sampling of achromatic contrasts in natural scenes through asymmetric photoreceptor distribution. Neuron 80, 1206–1217 (2013)
Yatsenko, D. et al. DataJoint: managing big scientific data using MATLAB or Python. Preprint at http://biorxiv.org/content/early/2015/11/14/031658 (2015)
Dorostkar, M. M., Dreosti, E., Odermatt, B. & Lagnado, L. Computational processing of optical measurements of neuronal and synaptic activity in networks. J. Neurosci. Methods 188, 141–150 (2010)
Szczurek, E., Biecek, P., Tiuryn, J. & Vingron, M. Introducing knowledge into differential expression analysis. J. Comput. Biol. 17, 953–967 (2010)
Bishop, C. M. Pattern Recognition and Machine Learning (Springer, 2006)
Acknowledgements
We thank G. Eske for technical support, C. Behrens for help with EM data, J. Jüttner for help with virus injection; X. Pitkow and R. Taylor for discussions; and L. L. Looger, the Janelia Research Campus of the Howard Hughes Medical Institute and the Genetically-Encoded Neuronal Indicator and Effector (GENIE) Project for making the viral constructs (AAV9.hSyn.iGluSnFR.WPRE.SV40, AAV9.CAG.Flex.iGluSnFR.WPRE.SV40 and AAV9.Syn.Flex.GCaMP6f.WPRE.SV40) publically available. This work was supported by the Deutsche Forschungsgemeinschaft (DFG; EXC307 to T.E.; BA 5283/1-1 to T.B.; BE 5601/1-1 to P.B.), the German Federal Ministry of Education and Research (BMBF; FKZ 01GQ1002 to M.B. and T.E.; FKZ 01GQ1601 to P.B.), the BW-Stiftung (AZ 1.16101.09 to T.B.), the intramural fortüne program of the University of Tübingen (2125-0-0 to T.B.), the European Commission (H2020 ERC-StG 677687 ‘NeuroVisEco’ to T.B.), the National Institute of Neurological Disorders and Stroke (U01NS090562 to T.E.) and the National Eye Institute (1R01EY023766 to T.E.) of the National Institutes of Health. The content is solely the responsibility of the authors and does not necessarily represent the official views of the funders.
Author information
Authors and Affiliations
Contributions
K.F., P.B., T.E. and T.B. designed the study; K.F. performed experiments and pre-processing; K.F. performed viral injections with help from T.S.; P.B. developed clustering with input from M.B.; K.F., P.B. and T.B. analysed the data with input from T.E.; K.F., P.B., T.E. and T.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. Diamond and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 ROI detection.
a, Mean correlation (±s.d. shading, n = 100 pixels) between noise-response traces of two individual pixels from scan field shown in Fig. 1c plotted against the distance of each pixel pair. Dotted line shows linear fit to the data above x = 10 μm and its extrapolation towards x = 0 μm. Exponential fit for distances <5 μm in orange. b, Zoomed-in version of a. The space constant obtained from the exponential fit (orange) was used to determine a scan field’s specific correlation minimum for ROI detection. Blue shading indicates the range of allowed ROI radii (0.375–2 μm). c, Scan field from Fig. 1c, with each pixel colour-coded by its correlation with the noise trace of pixel indicated in white. d, As c, with the red shading corresponding to pixels with a correlation coefficient greater than the correlation minimum from b, resulting in the red ROI in e. f, As a, averaged for n = 71 scan fields recorded at 48 × 12 μm. g, Histogram of ROI (black) and BC axon terminal (red) areas. The terminal area was determined from BC axonal arborizations labelled in GCaMP6-injected Pcp2 mice where individual axon terminals can be distinguished (see Extended Data Fig. 2a). Inset shows a histogram of the number of ROIs per BC axon terminal. h, Distribution of ROI numbers per scan field. i, Histogram illustrating sampling of ROIs against IPL depth. Data collected specifically for drug experiments (Methods) contributed to the ‘oversampling’ of the two ChAT bands.
Extended Data Figure 2 GCaMP6 signals in mouse BC axon terminals.
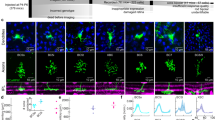
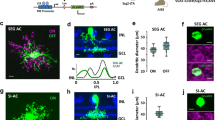
a–c, High-resolution scan of GCaMP6f-expressing BC axon terminal systems in the IPL of a Pcp2 mouse (a) and corresponding scan field (b) with automatically generated ROI mask overlaid (c, see Extended Data Fig. 1). Our ROI algorithm reliably detected individual axon terminals and preferably assigned two ROIs to a single terminal before merging two terminals into one ROI (compare with Extended Data Fig. 1g). d, Exemplary mean local-chirp responses of individual ROIs shown in c of the two different BCs shown in a. e, Scan field from c with ROI mask and spatial receptive fields (2 s.d. outlines of Gaussian fit) overlaid. f, Distribution of receptive field diameters estimated from On BC terminal calcium (GCaMP6) and glutamate release (iGluSnFR). Black dots correspond to mean receptive field diameters. Receptive field sizes estimated from calcium signals of single terminals closely fit those estimated from single iGluSnFR ROIs (56.1 ± 10 μm for GCaMP6 and 57.5 ± 10.6 μm for iGluSnFr; P > 0.05, n = 261 (GCaMP6) and n = 3,540, non-parametric non-paired Wilcoxon signed-rank test) and matched the anatomical dimensions of BC dendritic fields7,8. The findings shown here suggest that each ROI probably captured the light-driven glutamate signal of at most one individual BC axon terminal. g, h, High-resolution scan of GCaMP6-expressing RBC axon terminals (g) and corresponding scan field (h), with two individual terminals indicated. i, j, Mean responses of RBC terminals shown in h to full-field flashes (i, n = 20 trials) and local chirp stimulus (j, n = 5 trials).
Extended Data Figure 3 Alternative clustering including the glutamatergic monopolar interneuron (GluMI).
a, b, Heat maps of local chirp responses of C1, C2 and CGluMI (a) and their respective cluster means (b). Superimposed cluster means (b, bottom) illustrate response similarity between clusters. c, d, As in a and b, respectively, but for full-field responses.
Extended Data Figure 4 Clustering.
a, Exemplary distributions of prior probabilities for cluster allocation (right) taken from mean stratification profiles (left) of scan fields recorded at two different IPL depths (A: 1.7; B: 0). b, Temporal features extracted from glutamate traces in response to local (n = 6 features, top) and full-field (n = 12 features, middle) chirp and full-field flashes (n = 6 features, bottom). c, Heat maps of all recorded glutamate responses (n = 14 clusters plus ROIs discarded based on signal-to-noise (S/N) ratio) to the four visual stimuli (compare with Fig. 1); n = 11,101 ROIs from 29 retinas. Each line corresponds to the responses of individual ROIs with activity colour-coded. Block height represents the number of included ROIs per cluster. Within one cluster, ROIs are sorted on the basis of the quality of their receptive fields and local chirp response (Methods). Overlaid stratification profiles (left) illustrate overlap for some BC types. d, e, Cluster separation was determined for every cluster pair using the sensitivity index d′. Dotted lines in d illustrate transition between On and Off clusters and dotted line in e at d′ = 2, which corresponds to about 15% false positive/false negative rates. f, Separation of exemplary cluster pairs with a low (d′ = 2.0, top) and an average (d′ = 4.5, bottom) sensitivity index. g, Distribution of field entropies (left, Methods). Two exemplary scan fields (right) with ROIs colour-coded by cluster allocation illustrate low (1) and high (2) field entropy, respectively.
Extended Data Figure 5 Anatomical verification of clustering approach.
a, High-resolution scan of the axon terminal system of a filled BC (middle), with the iGluSnFR staining overlaid in green (top) and the corresponding scan field (bottom). b, Colour-coded ROI mask (cyan), with ROIs assigned to the labelled axon terminal system in red (top; Methods). Cluster allocations of all ROIs, and ROIs assigned to the labelled cell that passed the quality criterion, are shown in the middle and bottom panel, respectively. c, Reconstructed BC and its smoothed stratification profile (black), with the profile of the BC underlying the assigned functional cluster (CBC2) overlaid (red). d, Receptive field outlines of ROIs with QiRF > 0.4 allocated to one reconstructed cell. e, Mean local (top) and full-field chirp (bottom) responses of all ROIs assigned to the labelled cell (black), with s.d. shading (grey) and cluster mean of the assigned functional cluster (C2, red). f–j, As in a–e; k–n, as in b–e. o, Box plots illustrate the fraction of ROIs from one cell that were assigned to the same cluster for all reconstructed BCs (n = 8 cells, n = 3 mice). For the plot in the bottom, functionally very similar clusters (C3a and C3b; C5o, C5i and C5t) were merged. Owing to lower signal-to-noise ratios and a slower sampling rate (15.625 Hz vs. 31.25 Hz) for these single-cell data compared to all other data, the fraction of ‘correctly’ assigned ROIs is likely to be underestimated.
Extended Data Figure 6 Functional organization of the IPL.
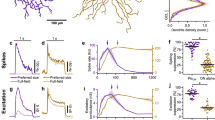
a, Glutamate responses to the local chirp step stimulus (first 8 s) of single ROIs assigned to C2 and C9. Shown are responses from three trials and a histogram of response amplitudes across each cluster’s 100 best-responding ROIs. On responses in Off BC cluster C2 are highlighted (arrows). b, Percentage of ROIs with at least one opposite polarity event in response to the local chirp step response for On and Off BC clusters. Dotted line illustrates mean incidence of spontaneous events (<1%). c, Time to peak of On events observed in the Off layer (grey) and On responses in On layer (red). Events were estimated from responses to single trials of the local chirp stimulus. Owing to the variability in timing, On events are not evident in traces averaged across the population of ROIs for each cluster (Fig. 2d). d, As in a, showing ‘spiking’ (C3b, CX) and non-spiking (C1, CR) responses in clusters of either polarity. e, f, Mean spectra (n = 5 trials) of local chirp step responses for two Off (C3b and C1) and two On (CX and CR) ROIs shown in d, with HFi estimated from the relative power of low (0.5–1 Hz) and high (2–16 Hz) frequencies (Methods). g, Response measures estimated from local chirp responses for all ROIs plotted against IPL depth (see Fig. 2e–h). h, Spatial receptive fields of individual ROIs assigned to C3a and C6. i, Mean response transience of BC clusters is not correlated with mean response delay (r = 0.01, P > 0.05, n = 14, linear correlation). j, Mean HFi and mean response delay are not correlated across BC clusters (r = 0.3, P > 0.05, n = 14, linear correlation). k, Mean chirp responses of two Off (C2 and C3b) clusters, with linear correlation coefficient (ρ) of whole trace or contrast ramp indicated. l, Cluster means of local (left) and full-field (right) chirp responses embedded in 2D feature space based on first and second principal components (PC). m, Response measures estimated from full-field chirp responses for all ROIs plotted against IPL depth (compare with Fig. 2e–h).
Extended Data Figure 7 GABA and glycinergic inhibition differentially shapes BC responses.
a, Mean responses (n = 5 trials) of individual ROIs to alternating local and full-field flashes under control conditions and with GABA (top) or glycine (bottom) receptor block. b, Change in mean peak amplitudes (±s.d. shading) of n = 15 ROIs originating from two scan fields during wash-in of GABA and glycine receptor blockers. c, Drug-induced changes in peak response amplitude across different BC clusters upon blocking GABA (top; P < 0.001, n = 9 clusters from 5 scan fields and 4 mice, non-parametric paired Wilcoxon signed-rank test) and glycine receptors (bottom; P < 0.001, n = 6 clusters from 5 scan fields and 4 mice). Mean ± s.d. in black. d, e, Local (grey) and full-field (black) chirp responses for control and drug conditions (d, GABA receptor block; e, glycine receptor block), with linear correlation coefficient (ρ) between each pair indicated. f, Local (grey) and surround (black) chirp responses for an exemplary On (C7, top) and Off (C3a, bottom) BC cluster. g, h, Spatial receptive fields with 2 s.d. outline of Gaussian fit shown in red (left) and quantification of changes in receptive field diameter across different BC clusters (right) upon blocking GABA (g; P < 0.05, n = 6 cluster from 3 scan fields and 2 mice, non-parametric paired Wilcoxon signed-rank test) and glycine receptors (h; P > 0.05, n = 5 cluster from 3 scan fields and 2 mice).
Extended Data Figure 8 Glycine-mediated crossover inhibition from the On pathway rectifies Off BCs.
a, Local chirp traces of an exemplary Off BC cluster (C3a) during control condition (left) and glycine receptor block (right). Magnified traces in the bottom illustrate an increase in tonic release upon drug application, with respective tonic release indices (TRi, Methods) indicated. Dotted line corresponds to baseline. Schematic (top right) illustrates effect of drug on inner retinal network. b, Quantification of drug-induced changes in tonic release across different Off BC clusters (P < 0.001, n = 5 clusters from 5 scan fields and 4 mice, non-parametric paired Wilcoxon signed-rank test). c, d, Local chirp responses of an exemplary On (c, C6) and Off (d, C2) BC cluster for control condition (left) and upon blocking the On BC pathway using l-AP4 (right). e, Tonic release indices of different Off BC clusters for control and l-AP4 conditions (P < 0.001, n = 5 clusters from 3 scan fields and 2 mice, non-parametric paired Wilcoxon signed-rank test). f, Average local chirp frequency and contrast responses of an exemplary Off (C1, top) and On (CX, bottom) BC cluster illustrate rectification of Off BC responses. g, Comparison of tonic release indices of On and Off BC clusters under control condition and Off BCs upon blocking crossover inhibition with l-AP4 or strychnine (*P < 0.05, **P < 0.01, non-parametric Kruskal–Wallis test).
Extended Data Figure 9 Extraction of BC centre–surround receptive fields.
a, Centre–surround maps obtained from the ring noise were averaged across ROIs assigned to one cluster (1). To isolate the surround component, the innermost two rings, which contained the majority of the centre component, were clipped (2) and the surround map estimated by SVD (3) was extrapolated across the centre by fitting a Gaussian (4). Next, the extrapolated surround map was subtracted from the average map and the centre component was extracted using SVD (6). The resultant centre–surround map was then subtracted from the average map to estimate the residual variance (7, Methods). Dotted lines at t = 0. b, Centre–surround maps of three ROIs assigned to CX from three independent experiments (top; bottom, left). Overlaid centre and surround time kernels (bottom, right) of ROIs (grey) and cluster mean (black) illustrate temporal precision and reproducibility. c, d, Heat maps of all centre (C) and surround (S) time kernels and cluster means ± s.d. for Off (c) and On (d) BC clusters.
Extended Data Figure 10 Centre–surround receptive fields of BC clusters.
a, Normalized time (top) and space (bottom) kernels of all On and Off BC clusters. Space kernels represent extrapolated Gaussian fits of centre and surround activation across rings shown in Fig. 5c, with circles corresponding to the original data points (Methods). b, c, Effect of GABA (b) and glycine (c) receptor block on centre–surround receptive fields of two exemplary BC clusters. Centre–surround maps correspond to averages of n > 5 ROIs of one scan field. d, Time to peak of centre time kernels were correlated with time to peak of surround time kernels (cluster mean ± s.e.m.; r = 0.91, P < 0.001, n = 14, linear correlation) and centre time kernel peaks preceded surround time kernel peaks, consistent with at least two additional synapses in the inhibitory pathway. Black line corresponds to linear fit and for the dashed line the slope is equal to 1. e, Half-maximal widths (cluster mean ± s.e.m.) of centre and surround time kernels were correlated (r = 0.83, P < 0.001, n = 14, linear correlation) and centre time kernel widths were consistently broader than surround time kernel widths. f, Half-maximal width of surround space kernels did not correlate with mean cluster IPL depth (r = 0.06, P > 0.05, n = 14, linear correlation). g, Time to peak of centre time kernels correlated with response delay estimated from local chirp step responses (mean ± s.e.m.; r = 0.82, P < 0.001, n = 14, linear correlation), indicating that the centre kernels adequately reflect BC responses to local stimuli. h, Predicted CSRs of BC clusters for different stimulus diameters (compare with Fig. 5d). i, j, Normalized temporal kernels predicted for local (100 μm diameter), full-field (500 μm diameter) and surround-only (500 and 100 μm outer and inner diameter, respectively) stimulation for Off (i) and On (j) BC clusters (compare with Fig. 5e).
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion and additional references. (PDF 296 kb)
Glutamate responses in the IPL
Background-subtracted and colour-coded glutamate signals recorded in the On ChAT band (64x16 pixels @31.25 Hz), where yellow corresponds to higher activity. Responses to the noise stimulus (exemplary 30 s, single trial) and local chirp (mean across n=5 trials) are shown. Scan field corresponds to the one shown in (Fig. 1c). Video in real-time. (MP4 29084 kb)
Rights and permissions
About this article
Cite this article
Franke, K., Berens, P., Schubert, T. et al. Inhibition decorrelates visual feature representations in the inner retina. Nature 542, 439–444 (2017). https://doi.org/10.1038/nature21394
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21394
This article is cited by
-
Ancient origin of the rod bipolar cell pathway in the vertebrate retina
Nature Ecology & Evolution (2024)
-
A presynaptic source drives differing levels of surround suppression in two mouse retinal ganglion cell types
Nature Communications (2024)
-
Distributed feature representations of natural stimuli across parallel retinal pathways
Nature Communications (2024)
-
Ancestral photoreceptor diversity as the basis of visual behaviour
Nature Ecology & Evolution (2024)
-
Heterogeneity of synaptic connectivity in the fly visual system
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.