Abstract
Phosphorus is an important nutrient for crop productivity. More than 60% of the total phosphorus in cereal crops is finally allocated into the grains and is therefore removed at harvest. This removal accounts for 85% of the phosphorus fertilizers applied to the field each year1,2. However, because humans and non-ruminants such as poultry, swine and fish cannot digest phytate, the major form of phosphorus in the grains, the excreted phosphorus causes eutrophication of waterways. A reduction in phosphorus accumulation in the grain would contribute to sustainable and environmentally friendly agriculture. Here we describe a rice transporter, SULTR-like phosphorus distribution transporter (SPDT), that controls the allocation of phosphorus to the grain. SPDT is expressed in the xylem region of both enlarged- and diffuse-vascular bundles of the nodes, and encodes a plasma-membrane-localized transporter for phosphorus. Knockout of this gene in rice (Oryza sativa) altered the distribution of phosphorus, with decreased phosphorus in the grains but increased levels in the leaves. Total phosphorus and phytate in the brown de-husked rice were 20–30% lower in the knockout lines, whereas yield, seed germination and seedling vigour were not affected. These results indicate that SPDT functions in the rice node as a switch to allocate phosphorus preferentially to the grains. This finding provides a potential strategy to reduce the removal of phosphorus from the field and lower the risk of eutrophication of waterways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rose, T. J., Liu, L. & Wissuwa, M. Improving phosphorus efficiency in cereal crops: Is breeding for reduced grain phosphorus concentration part of the solution? Front. Plant Sci. 4, 444 (2013)
Lott, J. N., Bojarski, M., Kolasa, J., Batten, G. D. & Campbell, L. C. A review of the phosphorus content of dry cereal and legume crops of the world. Int.J. Agric. Resour. Gov. Ecol. 8, 351–370 (2009)
Batjes, N. H. A world data set of derived soil properties by FAO-UNESCO soil unit for global modeling. Soil Use Manage . 13, 9–16 (1997)
Withers, P. J. A., Sylvester-Bradley, R., Jones, D. L., Healey, J. R. & Talboys, P. J. Feed the crop not the soil: rethinking phosphorus management in the food chain. Environ. Sci. Technol. 48, 6523–6530 (2014)
Kochian, L. V., Hoekenga, O. A. & Pineros, M. A. How do crop plants tolerate acid soils? Mechanisms of aluminum tolerance and phosphorous efficiency. Annu. Rev. Plant Biol. 55, 459–493 (2004)
Syers, J. K., Johnston, A. E. & Curtin, D. Efficiency of soil and fertilizer phosphorus use. FAO Fertilizer and Plant Nutrition Bulletin 18, (2008)
Cordell, D., Drangert, J. O. & White, S. The story of phosphorus: global food security and food for thought. Glob. Environ. Change 19, 292–305 (2009)
Batten, G. D. Phosphorus fractions in the grain of diploid, tetraploid, and hexaploid wheat grown with contrasting phosphorus supplies. Cereal Chem. 63, 384–387 (1986)
Karlen, D. L., Flannery, R. A. & Sadler, E. J. Aerial accumulation and partitioning of nutrients by corn. Agron. J . 80, 232–242 (1988)
Raboy, V. Seeds for a better future: ‘low phytate’ grains help to overcome malnutrition and reduce pollution. Trends Plant Sci . 6, 458–462 (2001)
White, P. J. & Veneklaas, E. J. Nature and nurture: the importance of seed phosphorus content. Plant Soil 357, 1–8 (2012)
Pariasca-Tanaka, J. et al. Does reducing seed-P concentrations affect seedling vigor and grain yield of rice? Plant Soil 392, 253–266 (2015)
Wang, F., Rose, T., Jeong, K., Kretzschmar, T. & Wissuwa, M. The knowns and unknowns of phosphorus loading into grains, and implications for phosphorus efficiency in cropping systems. J. Exp. Bot. 67, 1221–1229 (2016)
Dobermann, A. & Fairhurst, T. Rice: Nutrient Disorders & Nutrient Management Vol. 1. Int. Rice Res. Inst. (2000)
Raboy, V. Approaches and challenges to engineering seed phytate and total phosphorus. Plant Sci. 177, 281–296 (2009)
Rose, T. J. & Wissuwa, M. Rethinking internal phosphorus utilization efficiency: a new approach is needed to improve PUE in grain crops. Adv. Agron . 116, 185–217 (2012)
Baker, A. et al. Replace, reuse, recycle: improving the sustainable use of phosphorus by plants. J. Exp. Bot. 66, 3523–3540 (2015)
Yamaji, N. & Ma, J. F. The node, a hub for mineral nutrient distribution in graminaceous plants. Trends Plant Sci . 19, 556–563 (2014)
Yamaji, N., Sakurai, G., Mitani-Ueno, N. & Ma, J. F. Orchestration of three transporters and distinct vascular structures in node for intervascular transfer of silicon in rice. Proc. Natl Acad. Sci. USA 112, 11401–11406 (2015)
Takahashi, H., Buchner, P., Yoshimoto, N., Hawkesford, M. J. & Shiu, S.-H. Evolutionary relationships and functional diversity of plant sulfate transporters. Front. Plant Sci. 2, 119 (2012)
Yamaji, N., Xia, J., Mitani-Ueno, N., Yokosho, K. & Feng Ma, J. Preferential delivery of zinc to developing tissues in rice is mediated by P-type heavy metal ATPase OsHMA2. Plant Physiol. 162, 927–939 (2013)
Nakagawa, T. et al. Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104, 34–41 (2007)
Kuwano, M. et al. Molecular breeding for transgenic rice with low-phytic-acid phenotype through manipulating myo-inositol 3-phosphate synthase gene. Mol. Breed. 18, 263–272 (2006)
Ma, J. F. & Takahashi, E. Effect of silicon on the growth and phosphorus uptake of rice. Plant Soil 126, 115–119 (1990)
Miyaji, T. et al. AtPHT4;4 is a chloroplast-localized ascorbate transporter in Arabidopsis . Nat. Commun. 6, 5928 (2015)
Ma, J. F. et al. A silicon transporter in rice. Nature 440, 688–691 (2006)
Acknowledgements
This work was supported by Grant-in-Aid for Specially Promoted Research (JSPS KAKENHI Grant Number 15H04469 to N.Y. and 16H06296 to J.F.M.).
Author information
Authors and Affiliations
Contributions
N.Y. and Y.T. contributed equally to this work. N.Y. and J.F.M. designed research; N.Y. and Y.T. performed most experiments. Transport activity for Pi was determined by T.M. in vitro and by N.M.-U. in oocytes. K.T.Y. determined the phytic acid concentration; N.Y., Y.T. and J.F.M. analysed data; and N.Y. and J.F.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks C. Vance and M. Wissuwa and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Gene structure, Tos-17 insertion mutants and phylogeny of SPDT.
a, Structure of the SPDT gene. The gene consists of 10 exons and 9 introns. The exons are indicated as white boxes. Positions of the Tos-17 insertion are indicated as triangles. Primers (F1 and R2) for reverse transcription PCR (RT–PCR) are shown as arrows. b, Expression of SPDT mRNA in wild-type rice and three Tos-17 insertion mutants (spdt-1–spdt-3) examined by RT–PCR. c, Phylogenetic tree of sulfate transporter (Sultr) proteins in rice (Os) and Arabidopsis thaliana (At). Bootstrap values from 1,000 trials are indicated. The 0.1 scale shows substitution distance.
Extended Data Figure 2 Expression pattern of SPDT gene at different growth stages of rice.
a, b, Response of SPDT expression to different P (a) or S (b) supply at the vegetative growth stage. Seedlings (12 days old) were exposed to various external P or S concentrations for 1 week. The roots, shoot basal region (5 mm above the root-shoot junction), and shoots were sampled for RNA extraction. c, Expression pattern of SPDT at the reproductive growth stage. Samples of various organs were taken from rice grown in a paddy field at flowering and grain filling stages. The expression level was determined by quantitative RT–PCR. Expression relative to the shoot basal region in the control condition (a, b) or roots at the flowering stage (c) are shown. Histone H3 was used as the internal standard. Data are mean ± s.d. of biological replicates (n = 3).
Extended Data Figure 3 Subcellular localization of SPDT.
a–h, GFP:SPDT (a–d) or GFP control (e–h) together with DsRed were transiently introduced into onion epidermal cells by particle bombardment. Fluorescence signals from GFP (a, e), DsRed (b, f) and the merged images (c, g) are shown. Magnified image of boxed area in c and g are shown in d and h, respectively. Data are representative of three independent cells. Scale bars, 100 μm.
Extended Data Figure 4 Growth of wild-type rice and spdt mutants at the vegetative growth stage under low P and S conditions.
a–c, Wild-type rice and spdt mutants were grown in a nutrient solution containing 90 μM P and 460 μM S (a, control), 1 μM P and 460 μM S (b, low P), or 90 μM P and 10 μM S (c, low S) for 42 days. Plants were separated into roots, shoot basal region and individual leaves (2 to 9 from older to younger). Inset in b shows magnified image of leaf blade of leaf 7. For comparison, part pictures in a and b (WT and spdt-1) are the same as those in Fig. 3a, b. Pictures are representative of three biological replicates. d–f, Dry weight of each part sampled in a–c, respectively. Data are mean ± s.d. of biological replicates (n = 3). **P < 0.01, mutants compared with wild type (Tukey’s multiple comparison test).
Extended Data Figure 5 Accumulation, distribution and redistribution of P at the vegetative growth stage.
a, b, P content (a) and P concentration (b) in different organs. Wild-type rice and spdt mutants were grown hydroponically under a P-sufficient condition (90 μM) for 32 days. c, P re-distribution. Wild-type rice and three mutants were exposed to the nutrient solution free of P for 1 week. Samples of each organ were taken before and after P starvation treatment. The difference in P content (∆P) of each organ was calculated. d, Short-term (6 h) distribution analysis of P. Plants were treated with 1 μM radiolabelled 32P for 6 h. Distribution rate of newly absorbed P in different organ is shown. Data are mean ± s.d. of 3 (a–c) or 5 (d) biological replicates. *P < 0.05, **P < 0.01, mutants compared with wild type (Tukey’s test).
Extended Data Figure 6 Distribution of Mg, S, K and Ca in different organs at the vegetative growth stage.
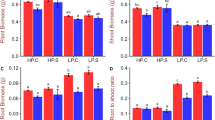
a–d, Wild-type rice and spdt mutants were grown hydroponically until the 8-leaf stage in half-strength Kimura B solution containing 90 μM Pi. Plants were separated into roots, shoot basal region and individual leaves (2 to 8 from older to younger) for the determination of Mg (a), S (b), K (c) and Ca (d) concentrations. Data are mean ± s.d. of biological replicates (n = 3). **P < 0.01, mutants compared with wild type (Tukey’s multiple comparison test).
Extended Data Figure 7 Comparison of growth and yield between wild-type rice and spdt mutants.
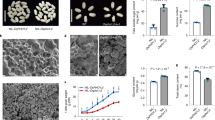
a–d, Wild-type rice and spdt mutants were grown in a paddy field until ripened. At harvest, plant height (a), stem length (b), tiller number (c) and panicle number (d) were recorded. Data are mean ± s.d. of 24 biological replicates (3 plots with 8 replicates each). *P < 0.05, **P < 0.01, mutants compared with wild type (Tukey’s multiple comparison test). e, f, Panicle (e) and brown rice (f) of wild-type rice and spdt mutants. Images are representative of 24 lines. Scale bars, 10 mm.
Extended Data Figure 8 Concentration and distribution of P at the reproductive growth stage.
Wild-type rice and spdt mutants were grown in a paddy field until ripened. a, P concentration in different organs of above-ground part. b, P content in different organs per tiller. c, Dry weight in different organs per tiller. d, Dry weight distribution ratio in different organs of above-ground part. e, Total P content of above-ground part per plant. f, Inorganic phosphate (Pi) concentration in brown rice. g, Stem-fed short-term distribution analysis. At the grain-filling stage, the plants were cut at the internode III. Solution containing 1 μM radiolabelled P was fed from the cut-end of the stem for 24 h. Distribution ratio of newly absorbed P in different organs above node I was calculated. Data are mean ± s.d. of 9 (3 plots with 3 replicates each; a–e), 3 (f) or 10 (g) biological replicates. *P < 0.05, **P < 0.01, compared with wild-type (Tukey’s test).
Extended Data Figure 9 Effect of SPDT knockout on seed germination and early growth.
a, Day-dependent germination rate. Thirty seeds each of wild-type rice and spdt mutants were used for the germination test, which was conducted in water at 30 °C in the dark. The water was changed every day. Data are mean ± s.d. of biological replicates (n = 3). b, c, Time-dependent growth of the shoot (b) and root (c) after germination. Twenty germinated seeds were grown in a 0.5 mM CaCl2 solution up to 5 days at 25 °C. Shoot and root lengths were measured with a ruler daily. Data are mean ± s.d. of biological replicates (n = 20). d, Phenotype of wild-type rice and spdt mutants grown in soil for 3 weeks.
Extended Data Figure 10 Element concentration of brown rice.
Wild-type rice and three spdt mutants were grown in a paddy field until ripened. Concentrations of elements in the brown rice were determined with a CHN analyser (C and N), MP-AES (S) or by ICP-MS (other elements) after digestion. Data are mean ± s.d. of 3 (C and N) or 9 (3 plots with 3 replicates each; other elements) biological replicates. *P < 0.05, **P < 0.01, compared with wild type (Tukey’s test).
Supplementary information
Supplementary Figure 1
The file contains the uncropped scans for Figure 2a and Extended Data Figure 1b. (PDF 1492 kb)
Supplementary Information
This file contains a Supplementary Discussion and additional references. (PDF 198 kb)
Source data
Rights and permissions
About this article
Cite this article
Yamaji, N., Takemoto, Y., Miyaji, T. et al. Reducing phosphorus accumulation in rice grains with an impaired transporter in the node. Nature 541, 92–95 (2017). https://doi.org/10.1038/nature20610
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20610
This article is cited by
-
OsCRLK2, a Receptor-Like Kinase Identified by QTL Analysis, is Involved in the Regulation of Rice Quality
Rice (2024)
-
A clade of receptor-like cytoplasmic kinases and 14-3-3 proteins coordinate inositol hexaphosphate accumulation
Nature Communications (2024)
-
Exploring the phosphorus–starch content balance mechanisms in maize grains using GWAS population and transcriptome data
Theoretical and Applied Genetics (2024)
-
Effects of Arbuscular Mycorrhizal Fungi on the Growth and Physiological Performance of Sophora davidii Seedling Under Low-Phosphorus Stress
Journal of Plant Growth Regulation (2024)
-
Phosphate environment and phosphate uptake studies: past and future
Journal of Plant Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.