Abstract
NLRs (nucleotide-binding domain and leucine-rich repeats) belong to a large family of cytoplasmic sensors that regulate an extraordinarily diverse range of biological functions. One of these functions is to contribute to immunity against infectious diseases, but dysregulation of their functional activity leads to the development of inflammatory and autoimmune diseases1. Cytoplasmic innate immune sensors, including NLRs, are central regulators of intestinal homeostasis2,3,4,5,6,7,8,9. NLRC3 (also known as CLR16.2 or NOD3) is a poorly characterized member of the NLR family and was identified in a genomic screen for genes encoding proteins bearing leucine-rich repeats (LRRs) and nucleotide-binding domains10,11. Expression of NLRC3 is drastically reduced in the tumour tissue of patients with colorectal cancer compared to healthy tissues12, highlighting an undefined potential function for this sensor in the development of cancer. Here we show that mice lacking NLRC3 are hyper-susceptible to colitis and colorectal tumorigenesis. The effect of NLRC3 is most dominant in enterocytes, in which it suppresses activation of the mTOR signalling pathways and inhibits cellular proliferation and stem-cell-derived organoid formation. NLRC3 associates with PI3Ks and blocks activation of the PI3K-dependent kinase AKT following binding of growth factor receptors or Toll-like receptor 4. These findings reveal a key role for NLRC3 as an inhibitor of the mTOR pathways, mediating protection against colorectal cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ting, J. P., Willingham, S. B. & Bergstralh, D. T. NLRs at the intersection of cell death and immunity. Nat. Rev. Immunol. 8, 372–379 (2008)
Zaki, M. H. et al. The NLRP3 inflammasome protects against loss of epithelial integrity and mortality during experimental colitis. Immunity 32, 379–391 (2010)
Zaki, M. H., Vogel, P., Body-Malapel, M., Lamkanfi, M. & Kanneganti, T. D. IL-18 production downstream of the Nlrp3 inflammasome confers protection against colorectal tumor formation. J. Immunol. 185, 4912–4920 (2010)
Allen, I. C. et al. The NLRP3 inflammasome functions as a negative regulator of tumorigenesis during colitis-associated cancer. J. Exp. Med. 207, 1045–1056 (2010)
Allen, I. C. et al. NLRP12 suppresses colon inflammation and tumorigenesis through the negative regulation of noncanonical NF-κB signaling. Immunity 36, 742–754 (2012)
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011)
Man, S. M. et al. Critical role for the DNA sensor AIM2 in stem cell proliferation and cancer. Cell 162, 45–58 (2015)
Wilson, J. E. et al. Inflammasome-independent role of AIM2 in suppressing colon tumorigenesis via DNA-PK and Akt. Nat. Med. 21, 906–913 (2015)
Zaki, M. H. et al. The NOD-like receptor NLRP12 attenuates colon inflammation and tumorigenesis. Cancer Cell 20, 649–660 (2011)
Harton, J. A., Linhoff, M. W., Zhang, J. & Ting, J. P. Cutting edge: CATERPILLER: a large family of mammalian genes containing CARD, pyrin, nucleotide-binding, and leucine-rich repeat domains. J. Immunol. 169, 4088–4093 (2002)
Conti, B. J. et al. CATERPILLER 16.2 (CLR16.2), a novel NBD/LRR family member that negatively regulates T cell function. J. Biol. Chem. 280, 18375–18385 (2005)
Liu, R. et al. Expression profile of innate immune receptors, NLRs and AIM2, in human colorectal cancer: correlation with cancer stages and inflammasome components. Oncotarget 6, 33456–33469 (2015)
Schneider, M. et al. The innate immune sensor NLRC3 attenuates Toll-like receptor signaling via modification of the signaling adaptor TRAF6 and transcription factor NF-κB. Nat. Immunol. 13, 823–831 (2012)
Zhang, L. et al. NLRC3, a member of the NLR family of proteins, is a negative regulator of innate immune signaling induced by the DNA sensor STING. Immunity 40, 329–341 (2014)
Man, S. M. & Kanneganti, T. D. Converging roles of caspases in inflammasome activation, cell death and innate immunity. Nat. Rev. Immunol. 16, 7–21 (2016)
Shimobayashi, M. & Hall, M. N. Making new contacts: the mTOR network in metabolism and signalling crosstalk. Nat. Rev. Mol. Cell Biol. 15, 155–162 (2014)
Alessi, D. R. et al. Characterization of a 3-phosphoinositide-dependent protein kinase which phosphorylates and activates protein kinase Bα. Curr. Biol. 7, 261–269 (1997)
Stephens, L. et al. Protein kinase B kinases that mediate phosphatidylinositol 3,4,5-trisphosphate-dependent activation of protein kinase B. Science 279, 710–714 (1998)
Stokoe, D. et al. Dual role of phosphatidylinositol-3,4,5-trisphosphate in the activation of protein kinase B. Science 277, 567–570 (1997)
Sancak, Y. et al. Ragulator–Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141, 290–303 (2010)
Zoncu, R. et al. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H+-ATPase. Science 334, 678–683 (2011)
Monick, M. M. et al. Lipopolysaccharide activates Akt in human alveolar macrophages resulting in nuclear accumulation and transcriptional activity of β-catenin. J. Immunol. 166, 4713–4720 (2001)
Jones, B. W., Heldwein, K. A., Means, T. K., Saukkonen, J. J. & Fenton, M. J. Differential roles of Toll-like receptors in the elicitation of proinflammatory responses by macrophages. Ann. Rheum. Dis. 60 (Supp. 3), iii6–iii12 (2001)
Ojaniemi, M. et al. Phosphatidylinositol 3-kinase is involved in Toll-like receptor 4-mediated cytokine expression in mouse macrophages. Eur. J. Immunol. 33, 597–605 (2003)
Laudien, M. et al. Molecular signatures of a disturbed nasal barrier function in the primary tissue of Wegener’s granulomatosis. Mucosal Immunol. 4, 564–573 (2011)
Shiau, C. E., Monk, K. R., Joo, W. & Talbot, W. S. An anti-inflammatory NOD-like receptor is required for microglia development. Cell Reports 5, 1342–1352 (2013)
Pelletier, S., Gingras, S. & Green, D. R. Mouse genome engineering via CRISPR–Cas9 for study of immune function. Immunity 42, 18–27 (2015)
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014)
Zhu, Q. et al. Cutting edge: STING mediates protection against colorectal tumorigenesis by governing the magnitude of intestinal inflammation. J. Immunol. 193, 4779–4782 (2014)
Man, S. M. et al. The transcription factor IRF1 and guanylate-binding proteins target activation of the AIM2 inflammasome by Francisella infection. Nat. Immunol. 16, 467–475 (2015)
Boussemart, L. et al. eIF4F is a nexus of resistance to anti-BRAF and anti-MEK cancer therapies. Nature 513, 105–109 (2014)
Karki, R. et al. Concerted activation of the AIM2 and NLRP3 inflammasomes orchestrates host protection against Aspergillus infection. Cell Host Microbe 17, 357–368 (2015)
Acknowledgements
We thank the Transgenic Gene Knockout Shared Resource at St. Jude Children’s Research Hospital (SJCRH) for assistance with knockout mouse generation. Images were acquired at the SJCRH Cell & Tissue Imaging Center, which is supported by SJCRH and NCI grant P30 CA021765-35. Work from our laboratory is supported by the US National Institutes of Health (grants AI101935, AI124346, AR056296 and CA163507 to T.-D.K.), ALSAC (to T.-D.K.), and the ExC306 Inflammation at Interfaces, the DFG SFB 877 B9 and DFG SFB1182 C2 projects (to P.R.). S.M.M. is supported by the R. G. Menzies Early Career Fellowship from the National Health and Medical Research Council of Australia.
Author information
Authors and Affiliations
Contributions
R.K., S.M.M. and T.-D.K. conceptualized the study; R.K., S.M.M., R.K.S.M. and S.K. designed the methodology; R.K., S.M.M., R.K.S.M., S.K., Q.Z., B.R.S., A.R.B., X.Q., S.P. and P.V. performed the experiments; R.K., S.M.M., R.K.S.M., S.K., Q.Z. and P.V. conducted the analysis; R.K., S.M.M. and T.-D.K. wrote the manuscript; P.R. and T.-D.K. provided resources; T.-D.K. provided overall supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 NLRC3 prevents colitis-associated colorectal tumorigenesis.
a, Timeline for azoxymethane (AOM) and DSS treatment. b, Relative expression levels of the gene encoding NLRC3 in tumour and non-tumour tissue in the colon of wild-type mice 80 days after azoxymethane injection. c, Targeting strategy used to generate Nlrc3−/− mice and PCR analysis for the gene encoding NLRC3 in wild-type C57BL/6 mice, Nlrc3+/– mice and Nlrc3−/− mice. The primers P1 (which binds a region between exon 1 and exon 2) and P2 (which binds a region between exon 3 and exon 4) were designed for ‘PCR1’ such that it generates a 4,804-bp PCR fragment for the wild-type allele and a 2,309-bp fragment for the knockout (KO) allele. However, PCR1 cannot differentiate heterozygote (HT) and knockout mice because the knockout 2,309-bp fragment outcompetes the wild-type 4,804-bp fragment. Therefore, we designed primers P3 and P4 for use in PCR2 to amplify a 940-bp fragment from exon 3 to confirm its presence in wild-type and HET mice and its absence in knockout mice. d, Histological scores of the colon tissue in wild-type and Nlrc3−/− mice 80 days after azoxymethane injection e, Percentages of mice with dysplasia 80 days after injection of azoxymethane. f, Percentages of mice with adenocarcinoma 80 days after injection of azoxymethane. g, Haematoxylin and eosin staining of colon crypts. Scale bar, 100 μm. Each symbol represents an individual mouse (b, d). ****P < 0.0001; NS, not statistically significant by one-way ANOVA (b) or two tailed t-test (d). Data represent two independent experiments (b, d–g) and are presented as mean ± s.e.m. (b, d).
Extended Data Figure 2 NLRC3 dampens intestinal inflammation.
a, Body-weight change of mice pooled from three independent experiments. b, Images of colon and colon length in wild-type mice and Nlrc3−/− mice at day 14. c, Histological scores at day 14. d, Levels of IL-18 and IL-1β in colon tissue at days 14 and 80. e, f, Levels of IL-6, TNF, GCSF, KC, MCP-1 and MIP-1α in colon tissue at days 14 and 80. g, Relative expression of genes encoding IL-6, TNF, GCSF and KC in colon tissue of untreated wild-type and Nlrc3−/− mice and in wild-type and Nlrc3−/− mice at day 14. h, Levels of IL-6, GCSF, KC and MIP-1α in sera of untreated wild-type and Nlrc3−/− mice and in wild-type and Nlrc3−/− mice at days 14 and 80. Each symbol represents an individual mouse (b–h). **P < 0.01; ****P < 0.0001; NS, not statistically significant by one-way ANOVA (a) or two tailed t-test (b–h). Data represent three independent experiments (a–h) and are presented as mean ± s.e.m. in a–h.
Extended Data Figure 3 NLRC3 governs colorectal tumorigenic susceptibility via inflammatory mediators and immune cells.
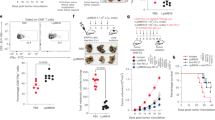
a, Relative expression of genes encoding IL-17a, levels of the IL-17 protein, and relative expression of genes encoding IL-22, IL-23p19, IFNβ and IFNγ in colon tissue of untreated wild-type and Nlrc3−/− mice and in wild-type and Nlrc3−/− mice at day 14. b, Immunoblot analysis of phosphorylated and total IκBα (P-IκBα and T-IκBα), ERK1 and ERK2 (P-ERK1/2 and T-ERK1/2), phosphorylated STAT3 (P-STAT3), and β-actin (loading control) in colon tissue of wild-type and Nlrc3−/− mice 14 days after injection of azoxymethane (top). The protein band intensity was normalized to the total protein counterpart and/or β-actin, and expressed relative to that of wild-type controls, set at 1 (bottom). c, Gating strategies used to generate data in d. d, Number of macrophages, CD11b+CD11c+ cells, neutrophils, B cells, CD4+ T cells, CD8+ T cells and natural killer cells per colon in wild-type and Nlrc3−/− mice at days 8 and 14. e, Splenocytes from wild-type and Nlrc3−/− mice were stimulated with CD3, CD28, and IL-2 and intracellular staining was performed for IFNγ and TNF. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; NS, not statistically significant by two-tailed t-test (a, b, d, e). Data pooled from two independent experiments (a) or represent one experiment representative of two independent experiments (b–e) and are presented as mean ± s.e.m. in a, b, d, e.
Extended Data Figure 4 The inhibitory effect of NLRC3 is more dominant in intestinal epithelial cells than in haematopoietic cells.
a, Left panel shows colon tumours in, from left to right, wild-type → wild-type (n = 10); Nlrc3−/− → wild-type (n = 9); wild-type → Nlrc3−/− (n = 8); Nlrc3−/− → Nlrc3−/− (n = 9) bone-marrow chimeric mice at day 80. Percentages of the tumour size of mice are shown on the right. b, Left panel shows colon tumours in littermate Nlrc3fl/fl (n = 8), LysMcreNlrc3fl/fl (n = 11), Vav1creNlrc3fl/fl (n = 9), Vil1creNlrc3fl/fl (n = 7) and Nlrc3−/− (n = 8) mice at day 80. The percentages of tumours of each size of each mouse type are shown on the right. c, Relative expression of genes encoding LGR5, WNT1, β-catenin (Ctnnb1), Axin2, TCF4, TCF7, and LEF1 in colon tissue of untreated wild-type and Nlrc3−/− mice or in wild-type and Nlrc3−/− mice at day 14. d, Immunohistochemical staining of β-catenin in colon tissue of wild-type and Nlrc3−/− mice. Scale bar, 20 μm. Each symbol represents one mouse (c). NS, not statistically significant by two-tailed t-test. Data represent two independent experiments and are presented as mean ± s.e.m. in c.
Extended Data Figure 5 Dysregulation of mTOR signalling precedes dysregulation of NF-kB signalling.
a, Immunoblot analysis of phosphorylated mTOR, S6, 4E-BP1, AKT and IκBα, and GAPDH (loading control) in colon tissue of wild-type and Nlrc3−/− mice at day 8 (left). The protein band intensity was normalized to GAPDH and expressed as a level relative to that of the wild-type controls, set at 1 (right). b, Levels of IL-18, IL-1β, IL-6, TNF, GCSF, KC, MCP-1 and MIP-1α in colon tissue at day 8. c, Levels of IL-18, IL-1β, IL-6, TNF, GCSF, KC, MCP-1 and MIP-1α in sera. Each symbol represents an individual mouse (b, c). *P < 0.05; **P < 0.01; NS, not statistically significant by two-tailed t-test (a–c). Data represent two independent experiments and are presented as mean ± s.e.m. in a–c. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 6 NLRC3 regulates mTOR activity.
a, Immunoblot analysis of phosphorylated AKT, total AKT, phosphorylated 4E-BP1 and GAPDH (loading control) in the colon tissue of wild-type, Nlrc3+/− and Nlrc3−/− mice at day 14. The protein-band intensity was normalized to the total protein counterpart and/or the loading control and expressed relative to that of wild-type controls, set at 1 (right). b, Immunoblot analysis of phosphorylated AKT, S6K, S6 and 4E-BP1, and GAPDH (loading control) in wild-type fibroblasts transfected with a control siRNA or Nlrc3 siRNA left untreated or treated with IGF-1 (top). Densitometry analysis as in a over 120 min (bottom). c, Relative expression of the gene encoding NLRC3 in wild-type fibroblasts transfected with a control siRNA or compared with wild-type fibroblasts transfected with an Nlrc3 siRNA. d, Immunoblotting of phosphorylated S6K, S6, 4E-BP1, AKT, and total AKT in primary fibroblasts either left untreated or treated with IGF-1 (top). Densitometry analysis as in a (bottom). e, Immunoblotting of phosphorylated AKT and mTOR, and β-actin (loading control) in primary fibroblasts either left untreated or treated with IGF-1 (left). Densitometry analysis as in b (right). *P < 0.05; **P < 0.01; ***P < 0.001 by one-way ANOVA (a, b, d, e). Data are from one experiment representative of two (a, c) or four independent experiments (b, d and e) and are presented as mean ± s.e.m. in a, b, d, e. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 7 NLRC3 regulates mTOR activity in fibroblasts.
a–d, Immunofluorescent stain of phosphorylated S6 (a), 4E-BP1 (b), AKT (c), and AKT (d) in primary fibroblasts either left untreated or treated with IGF-1 for 30 min is shown in the left panels. Quantification of the fluorescence intensity in each cell (n = 150 or more) is shown in the right panels. ADU, average density unit. Scale bar, 20 μm. Each symbol represents an individual cell. **** P < 0.0001; NS, not statistically significant by two-tailed t-test. Data represent one experiment representative of two independent experiments and are presented as mean ± s.e.m.
Extended Data Figure 8 NLRC3 prevents colorectal cancer in an ApcMin/+ model of tumorigenesis.
a, Images of colon tumours (left), tumour number and colon length (middle), and size (right) of 120-day-old littermate ApcMin/+ and ApcMin/+Nlrc3−/− mice. b, Percentage of mice with dysplasia (left), total histology scores (middle), and histology scores of different parts of colon and different parameters (right) of mice in a. c, Haematoxylin and eosin (H&E, top), Ki67 (middle) and phosphorylated S6 (P-S6, bottom) staining of colon tumours. Scale bar, 200 μm. d, Quantification of the number (top) and size (bottom) of organoids derived from colonic stem cells of ApcMin/+ and ApcMin/+Nlrc3−/− mice left untreated or treated with LY294002 or rapamycin. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; NS, not statistically significant by two-tailed t-test (a, b, d). Data represent two independent experiments and are presented as mean ± s.e.m. in a, b, d.
Extended Data Figure 9 NLRC3 disrupts the assembly of the PI3K heterodimeric complex.
a, b, Immunoprecipitation and comparative analysis of the PI3K signalling complex levels between wild-type and Nlrc3−/− primary mouse embryonic fibroblasts (MEFs; a) and BMDMs (b). c, Schematic representation of the generation of deletion mutants of NLRC3. d, Loading inputs for e–g. e, Immunoprecipitation of wild-type-NLRC3 and its deletion mutants. e, f, Immunoblotting analysis of the interaction between NLRC3 and its mutants with the PI3K–p110 (f) and PI3K–p85 (g) subunit. Data represent two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 10 NLRC3 negatively regulates TLR4-induced activation of the PI3K–AKT–mTOR pathway.
a, Immunoblot analysis of phosphorylated AKT (at Thr308 and Ser473), total AKT and β-actin (loading control) in wild-type and Nlrc3−/− bone-marrow-derived macrophages (BMDMs) left untreated or treated with LPS (top). The protein-band intensity was normalized to β-actin, and expressed relative to that of wild-type controls, set at 1 (bottom). b, Immunoblot analysis of phosphorylated mTOR, phosphorylated 4E-BP1, and β-actin (loading control) in wild-type and Nlrc3−/− bone-marrow-derived macrophages (BMDMs) left untreated or treated with LPS (top). Densitometry analysis as in a (bottom). c, A model of the role of NLRC3 in the negative regulation of the PI3K–AKT–mTOR pathway. * P < 0.05; ** P < 0.01; *** P < 0.001 by two-tailed t-test (a, b). Data are from one experiment representative of four independent experiments and are presented as mean ± s.e.m. in a and b. For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Information
This file contains the uncropped gel source data. (PDF 5141 kb)
Supplementary Data
This file contains a list of primer sequences. (XLSX 35 kb)
Source data
Rights and permissions
About this article
Cite this article
Karki, R., Man, S., Malireddi, R. et al. NLRC3 is an inhibitory sensor of PI3K–mTOR pathways in cancer. Nature 540, 583–587 (2016). https://doi.org/10.1038/nature20597
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20597
This article is cited by
-
Nlrc3 signaling is indispensable for hematopoietic stem cell emergence via Notch signaling in vertebrates
Nature Communications (2024)
-
Chemoproteomic mapping of the glycolytic targetome in cancer cells
Nature Chemical Biology (2023)
-
The NLR gene family: from discovery to present day
Nature Reviews Immunology (2023)
-
Molecular mechanisms in colitis-associated colorectal cancer
Oncogenesis (2023)
-
Impact of intracellular innate immune receptors on immunometabolism
Cellular & Molecular Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.