Abstract
In all domains of life, selenocysteine (Sec) is delivered to the ribosome by selenocysteine-specific tRNA (tRNASec) with the help of a specialized translation factor, SelB in bacteria. Sec-tRNASec recodes a UGA stop codon next to a downstream mRNA stem–loop. Here we present the structures of six intermediates on the pathway of UGA recoding in Escherichia coli by single-particle cryo-electron microscopy. The structures explain the specificity of Sec-tRNASec binding by SelB and show large-scale rearrangements of Sec-tRNASec. Upon initial binding of SelB–Sec-tRNASec to the ribosome and codon reading, the 30S subunit adopts an open conformation with Sec-tRNASec covering the sarcin–ricin loop (SRL) on the 50S subunit. Subsequent codon recognition results in a local closure of the decoding site, which moves Sec-tRNASec away from the SRL and triggers a global closure of the 30S subunit shoulder domain. As a consequence, SelB docks on the SRL, activating the GTPase of SelB. These results reveal how codon recognition triggers GTPase activation in translational GTPases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Böck, A. Biosynthesis of selenoproteins—an overview. Biofactors 11, 77–78 (2000)
Lescure, A., Fagegaltier, D., Carbon, P. & Krol, A. Protein factors mediating selenoprotein synthesis. Curr. Protein Pept. Sci. 3, 143–151 (2002)
Berchtold, H. et al. Crystal structure of active elongation factor Tu reveals major domain rearrangements. Nature 365, 126–132 (1993)
Paleskava, A., Konevega, A. L. & Rodnina, M. V. Thermodynamic and kinetic framework of selenocysteyl-tRNASec recognition by elongation factor SelB. J. Biol. Chem. 285, 3014–3020 (2010)
Forchhammer, K., Boesmiller, K. & Böck, A. The function of selenocysteine synthase and SELB in the synthesis and incorporation of selenocysteine. Biochimie 73, 1481–1486 (1991)
Itoh, Y., Sekine, S., Suetsugu, S. & Yokoyama, S. Tertiary structure of bacterial selenocysteine tRNA. Nucleic Acids Res. 41, 6729–6738 (2013)
Zinoni, F., Heider, J. & Böck, A. Features of the formate dehydrogenase mRNA necessary for decoding of the UGA codon as selenocysteine. Proc. Natl Acad. Sci. USA 87, 4660–4664 (1990)
Itoh, Y., Sekine, S. & Yokoyama, S. Crystal structure of the full-length bacterial selenocysteine-specific elongation factor SelB. Nucleic Acids Res. 43, 9028–9038 (2015)
Leibundgut, M., Frick, C., Thanbichler, M., Böck, A. & Ban, N. Selenocysteine tRNA-specific elongation factor SelB is a structural chimaera of elongation and initiation factors. EMBO J. 24, 11–22 (2005)
Kromayer, M., Wilting, R., Tormay, P. & Böck, A. Domain structure of the prokaryotic selenocysteine-specific elongation factor SelB. J. Mol. Biol. 262, 413–420 (1996)
Nissen, P., Thirup, S., Kjeldgaard, M. & Nyborg, J. The crystal structure of Cys–tRNACys–EF-Tu–GDPNP reveals general and specific features in the ternary complex and in tRNA. Structure 7, 143–156 (1999)
Hatfield, D. L., Carlson, B. A., Xu, X. M., Mix, H. & Gladyshev, V. N. Selenocysteine incorporation machinery and the role of selenoproteins in development and health. Prog. Nucleic Acid Res. Mol. Biol. 81, 97–142 (2006)
Metanis, N. & Hilvert, D. Natural and synthetic selenoproteins. Curr. Opin. Chem. Biol. 22, 27–34 (2014)
Bulteau, A. L. & Chavatte, L. Update on selenoprotein biosynthesis. Antioxid. Redox Signal. 23, 775–794 (2015)
Ringquist, S. et al. Recognition of the mRNA selenocysteine insertion sequence by the specialized translational elongation factor SELB. Genes Dev. 8, 376–385 (1994)
Moazed, D. & Noller, H. F. Interaction of tRNA with 23S rRNA in the ribosomal A, P, and E sites. Cell 57, 585–597 (1989)
Förster, C., Ott, G., Forchhammer, K. & Sprinzl, M. Interaction of a selenocysteine-incorporating tRNA with elongation factor Tu from E.coli. Nucleic Acids Res. 18, 487–491 (1990)
Rudinger, J., Hillenbrandt, R., Sprinzl, M. & Giegé, R. Antideterminants present in minihelix(Sec) hinder its recognition by prokaryotic elongation factor Tu. EMBO J. 15, 650–657 (1996)
Yikilmaz, E., Chapman, S. J., Schrader, J. M. & Uhlenbeck, O. C. The interface between Escherichia coli elongation factor Tu and aminoacyl-tRNA. Biochemistry 53, 5710–5720 (2014)
Schrader, J. M., Chapman, S. J. & Uhlenbeck, O. C. Understanding the sequence specificity of tRNA binding to elongation factor Tu using tRNA mutagenesis. J. Mol. Biol. 386, 1255–1264 (2009)
Asahara, H. & Uhlenbeck, O. C. The tRNA specificity of Thermus thermophilus EF-Tu. Proc. Natl Acad. Sci. USA 99, 3499–3504 (2002)
Diaconu, M. et al. Structural basis for the function of the ribosomal L7/12 stalk in factor binding and GTPase activation. Cell 121, 991–1004 (2005)
Maracci, C. & Rodnina, M. V. Review: Translational GTPases. Biopolymers 105, 463–475 (2016)
Pape, T., Wintermeyer, W. & Rodnina, M. Induced fit in initial selection and proofreading of aminoacyl-tRNA on the ribosome. EMBO J. 18, 3800–3807 (1999)
Heider, J., Baron, C. & Böck, A. Coding from a distance: dissection of the mRNA determinants required for the incorporation of selenocysteine into protein. EMBO J. 11, 3759–3766 (1992)
Kotini, S. B., Peske, F. & Rodnina, M. V. Partitioning between recoding and termination at a stop codon-selenocysteine insertion sequence. Nucleic Acids Res. 43, 6426–6438 (2015)
Wohlgemuth, I., Pohl, C., Mittelstaet, J., Konevega, A. L. & Rodnina, M. V. Evolutionary optimization of speed and accuracy of decoding on the ribosome. Phil. Trans. R. Soc. Lond. B 366, 2979–2986 (2011)
Marshall, R. A., Aitken, C. E., Dorywalska, M. & Puglisi, J. D. Translation at the single-molecule level. Annu. Rev. Biochem. 77, 177–203 (2008)
Thompson, R. C., Dix, D. B. & Karim, A. M. The reaction of ribosomes with elongation factor Tu.GTP complexes. Aminoacyl-tRNA-independent reactions in the elongation cycle determine the accuracy of protein synthesis. J. Biol. Chem. 261, 4868–4874 (1986)
Ruusala, T., Ehrenberg, M. & Kurland, C. G. Is there proofreading during polypeptide synthesis? EMBO J. 1, 741–745 (1982)
Ogle, J. M., Murphy, F. V., Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002)
Ogle, J. M. et al. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 292, 897–902 (2001)
Dunkle, J. A. et al. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science 332, 981–984 (2011)
Fischer, N., Konevega, A. L., Wintermeyer, W., Rodnina, M. V. & Stark, H. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010)
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010)
Piepenburg, O. et al. Intact aminoacyl-tRNA is required to trigger GTP hydrolysis by elongation factor Tu on the ribosome. Biochemistry 39, 1734–1738 (2000)
Cochella, L. & Green, R. An active role for tRNA in decoding beyond codon:anticodon pairing. Science 308, 1178–1180 (2005)
Schmeing, T. M., Voorhees, R. M., Kelley, A. C. & Ramakrishnan, V. How mutations in tRNA distant from the anticodon affect the fidelity of decoding. Nat. Struct. Mol. Biol. 18, 432–436 (2011)
Moazed, D., Robertson, J. M. & Noller, H. F. Interaction of elongation factors EF-G and EF-Tu with a conserved loop in 23S RNA. Nature 334, 362–364 (1988)
Warner, G. J. et al. Inhibition of selenoprotein synthesis by selenocysteine tRNA[Ser]Sec lacking isopentenyladenosine. J. Biol. Chem. 275, 28110–28119 (2000)
Daviter, T., Wieden, H. J. & Rodnina, M. V. Essential role of histidine 84 in elongation factor Tu for the chemical step of GTP hydrolysis on the ribosome. J. Mol. Biol. 332, 689–699 (2003)
Maracci, C., Peske, F., Dannies, E., Pohl, C. & Rodnina, M. V. Ribosome-induced tuning of GTP hydrolysis by a translational GTPase. Proc. Natl Acad. Sci. USA 111, 14418–14423 (2014)
Cunha, C. E. et al. Dual use of GTP hydrolysis by elongation factor G on the ribosome. Translation (Austin) 1, e24315 (2013)
Adamczyk, A. J. & Warshel, A. Converting structural information into an allosteric-energy-based picture for elongation factor Tu activation by the ribosome. Proc. Natl Acad. Sci. USA 108, 9827–9832 (2011)
Wallin, G., Kamerlin, S. C. & Aqvist, J. Energetics of activation of GTP hydrolysis on the ribosome. Nat. Commun. 4, 1733 (2013)
Tourigny, D. S., Fernández, I. S., Kelley, A. C. & Ramakrishnan, V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science 340, 1235490 (2013)
Rodnina, M. V. et al. Thiostrepton inhibits the turnover but not the GTPase of elongation factor G on the ribosome. Proc. Natl Acad. Sci. USA 96, 9586–9590 (1999)
Milon, P. et al. Transient kinetics, fluorescence, and FRET in studies of initiation of translation in bacteria. Methods Enzymol. 430, 1–30 (2007)
Rodnina, M. V. & Wintermeyer, W. GTP consumption of elongation factor Tu during translation of heteropolymeric mRNAs. Proc. Natl Acad. Sci. USA 92, 1945–1949 (1995)
Thanbichler, M. & Böck, A. Purification and characterization of hexahistidine-tagged elongation factor SelB. Protein Expr. Purif. 31, 265–270 (2003)
Paleskava, A., Konevega, A. L. & Rodnina, M. V. Thermodynamics of the GTP-GDP-operated conformational switch of selenocysteine-specific translation factor SelB. J. Biol. Chem. 287, 27906–27912 (2012)
Thanbichler, M. & Böck, A. Selenoprotein biosynthesis: purification and assay of components involved in selenocysteine biosynthesis and insertion in Escherichia coli. Methods Enzymol. 347, 3–16 (2002)
Kothe, U., Paleskava, A., Konevega, A. L. & Rodnina, M. V. Single-step purification of specific tRNAs by hydrophobic tagging. Anal. Biochem. 356, 148–150 (2006)
Fischer, N. et al. Towards understanding selenocysteine incorporation into bacterial proteins. Biol. Chem. 388, 1061–1067 (2007)
Konevega, A. L. et al. Purine bases at position 37 of tRNA stabilize codon-anticodon interaction in the ribosomal A site by stacking and Mg2+-dependent interactions. RNA 10, 90–101 (2004)
Rodnina, M. V., Fricke, R. & Wintermeyer, W. Transient conformational states of aminoacyl-tRNA during ribosome binding catalyzed by elongation factor Tu. Biochemistry 33, 12267–12275 (1994)
Gromadski, K. B. & Rodnina, M. V. Kinetic determinants of high-fidelity tRNA discrimination on the ribosome. Mol. Cell 13, 191–200 (2004)
Fischer, N. et al. Structure of the E. coli ribosome-EF-Tu complex at <3 Å resolution by Cs-corrected cryo-EM. Nature 520, 567–570 (2015)
Valle, M. et al. Cryo-EM reveals an active role for aminoacyl-tRNA in the accommodation process. EMBO J. 21, 3557–3567 (2002)
Stark, H. et al. Ribosome interactions of aminoacyl-tRNA and elongation factor Tu in the codon-recognition complex. Nat. Struct. Biol. 9, 849–854 (2002)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014)
Pettersen, E. F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Kleywegt, G. J. & Jones, T. A. Software for handling macromolecular envelopes. Acta Crystallogr. D 55, 941–944 (1999)
Brunger, A. T. Version 1.2 of the Crystallography and NMR system. Nat. Protocols 2, 2728–2733 (2007)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
DiMaio, F. et al. Improved molecular replacement by density- and energy-guided protein structure optimization. Nature 473, 540–543 (2011)
DiMaio, F., Tyka, M. D., Baker, M. L., Chiu, W. & Baker, D. Refinement of protein structures into low-resolution density maps using rosetta. J. Mol. Biol. 392, 181–190 (2009)
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005)
Selmer, M. & Su, X. D. Crystal structure of an mRNA-binding fragment of Moorella thermoacetica elongation factor SelB. EMBO J. 21, 4145–4153 (2002)
Soler, N., Fourmy, D. & Yoshizawa, S. Structural insight into a molecular switch in tandem winged-helix motifs from elongation factor SelB. J. Mol. Biol. 370, 728–741 (2007)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Keating, K. S. & Pyle, A. M. Semiautomated model building for RNA crystallography using a directed rotameric approach. Proc. Natl Acad. Sci. USA 107, 8177–8182 (2010)
Chou, F. C., Sripakdeevong, P., Dibrov, S. M., Hermann, T. & Das, R. Correcting pervasive errors in RNA crystallography through enumerative structure prediction. Nat. Methods 10, 74–76 (2013)
Afonine, P. V., Headd, J. J., Terwilliger, T. C. & Adams, P. D. New tool: phenix.real_space_refine. Computational Crystallography Newsletter 43–44 (2013)
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000)
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Bock, L. V. et al. Energy barriers and driving forces in tRNA translocation through the ribosome. Nat. Struct. Mol. Biol. 20, 1390–1396 (2013)
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013)
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010)
Berendsen, H. J. C., Grigera, J. R. & Straatsma, T. P. The missing term in effective pair potentials. J. Phys. Chem. 91, 6269–6271 (1987)
Joung, I. S. & Cheatham, T. E., III . Determination of alkali and halide monovalent ion parameters for use in explicitly solvated biomolecular simulations. J. Phys. Chem. B 112, 9020–9041 (2008)
Aduri, R. et al. AMBER Force Field Parameters for the Naturally Occurring Modified Nucleosides in RNA. J. Chem. Theory Comput. 3, 1464–1475 (2007)
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 25, 247–260 (2006)
Amadei, A., Linssen, A. B. & Berendsen, H. J. Essential dynamics of proteins. Proteins 17, 412–425 (1993)
Schrader, J. M. & Uhlenbeck, O. C. Is the sequence-specific binding of aminoacyl-tRNAs by EF-Tu universal among bacteria? Nucleic Acids Res. 39, 9746–9758 (2011)
Haruna, K., Alkazemi, M. H., Liu, Y., Söll, D. & Englert, M. Engineering the elongation factor Tu for efficient selenoprotein synthesis. Nucleic Acids Res. 42, 9976–9983 (2014)
Itoh, Y. et al. Decameric SelA•tRNA(Sec) ring structure reveals mechanism of bacterial selenocysteine formation. Science 340, 75–78 (2013)
Li, W. et al. Recognition of aminoacyl-tRNA: a common molecular mechanism revealed by cryo-EM. EMBO J. 27, 3322–3331 (2008)
Takyar, S., Hickerson, R. P. & Noller, H. F. mRNA helicase activity of the ribosome. Cell 120, 49–58 (2005)
Budkevich, T. V. et al. Regulation of the mammalian elongation cycle by subunit rolling: a eukaryotic-specific ribosome rearrangement. Cell 158, 121–131 (2014)
Frank, J. & Agrawal, R. K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000)
Acknowledgements
We thank F. Würriehausen, A. Bursy, O. Geintzer, S. Kappler, C. Kothe, T. Niese, T. Wiles, and M. Zimmermann for expert technical assistance and M. Lüttich and T. Koske for support in high-performance computation. The work was supported by the Deutsche Forschungsgemeinschaft Grant FOR 1805 (to H.S., M.V.R. and H.G.) and by the Sonderforschungsbereich 860 (to R.F.).
Author information
Authors and Affiliations
Contributions
N.F. conceived the project, performed cryo-EM experiments and data analysis and drafted the paper. P.N. built atomic models and performed pseudo-crystallographic refinement. L.V.B. performed and analyzed molecular dynamics simulations. C.M. performed the kinetic analysis. A.P. and A.L.K prepared ribosome complexes for cryo-EM. Z.W. and G.F.S. performed homology modelling. N.F., M.V.R. and H.S. finalized the paper with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks A. Amunts, M. Wahl and S. Yokoyama for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Computational sorting of ribosome particle images.
a, Hierarchical sorting scheme. Numbers refer to classification steps. SVC, supervised classification by projection matching; ‘bad’ 70S, the particles of low quality and/or those showing mixtures of different compositions and conformations; initial complex (hybrid) (IC (hybrid)), ratcheted state of the ribosome with one tRNA in the hybrid state. b, Mask used for focused classification in step 4. c, Cryo-EM reconstruction from 100,000 random unsorted particle images. Left, initial refinement stage at 10 Å resolution showing only scattered density (red) for SelB, Sec-tRNASec and SECIS. Right, final cryo-EM map at 3.7 Å resolution. d, Structural changes resolved by computational sorting as quantified by the r.m.s.d. and the changes in mass due to ligand binding and dissociation.
Extended Data Figure 2 Cryo-EM map and model quality.
a, Fourier shell correlation (FSC) curves and model validation. FSC curves are shown for each state. Black, the FSC curve computed between the masked independent half-maps from cryo-EM refinement (half1 and half2); blue, the FSC curve between the final cryo-EM map (map) and the final model (model); red, the FSC curve between half-map 1 (half1) and the model obtained by refinement only against half-map 2 (model2). The vertical black dashed line indicates the resolution according to the 0.143 criterion (grey line) and the maximum resolution at which the full atomic models were refined. b, FSC curves (FSCwork) computed for each state between reflections from solvent-flattened cryo-EM maps and model maps generated from refined atomic coordinates. Differences to d largely result from solvent-flattening (Methods). c, CCwork curves as obtained by refinement for each state. For reliable resolution estimates CCwork is expected to be >0.2 in the highest resolution shell. d, Cryo-EM densities and models of the quarternary complex, SelB–GDPNP–Sec-tRNASec–SECIS-mRNA for each SelB-bound ribosome state. Densities are coloured as in Fig. 1b and rendered at ~2σ, except wh3, wh4 and SECIS of the GTPase-activated state, which are rendered at ~1.5σ. Top right: Sec-tRNASec density (purple) in the initial binding state at 5.3 Å resolution; the conformation of the invariant histidine 61 is not discernible (question mark). Middle right: Density for histidine 61 (blue) in the codon reading state indicating a partially flipped-in conformation. e, Densities for rRNA modifications as seen in the GTPase-activated state at 3.4 Å resolution. Densities are rendered at ~3σ. Arrows denote the characteristic distortion of the nucleobase of D2449 and methyl groups for the other modified nucleotides. f, g, Cryo-EM reconstructions of the individual states (as indicated) coloured according to local resolution. Left: Surface view; right: Cut-away view. Heat maps are adjusted to the respective resolutions ranges of the cryo-EM maps in f and g. The arrow denotes the substantially lower local resolution of SelB residues 485 to 614 in the GTPase-activated state; atomic models for such regions were correspondingly refined at lower resolution (Methods).
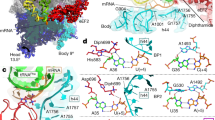
Extended Data Figure 3 Structural basis for the specificity of tRNASec recognition by SelB.
All superpositions are based on domains 1 and 2 of SelB and EF-Tu, respectively. a, Overview of SelB–GDPNP–Sec-tRNASec bound to the ribosome in the GTPase-activated state. Numbers refer to the close-ups in b, e and g, respectively, which depict specific SelB–Sec-tRNASec interactions. SelB d1–3, domains 1 to 3 of SelB; VA, variable arm. b, Interactions between SelB (domain 3, red, and the linker, residues 340–363 in khaki) and the backbone of the extra-long variable arm of tRNASec. These interactions do not form in the canonical EF-Tu–aa-tRNA complexes. c, The structure of tRNASec. In contrast to canonical tRNAs, tRNASec entails an additional base-pair in the acceptor stem (A5a–U67a, red), a unique sequence in the T-stem, the so-called antideterminant box (C7–G66, G49–U65 and C50–G64, orange), and a very long variable arm (yellow). d, Superposition of tRNASec (purple) and the canonical tRNACys (dark grey) in the complex with SelB–GDPNP and EF-Tu–GDPNP, respectively. The structure of the canonical EF-Tu–GDPNP–Cys-tRNACys is from PDB structure 1B23 (ref. 11). Note the distortion in the acceptor stem of tRNASec (residues G1-U6/G72-A67) as compared to tRNACys that compensates for the additional base pair. As a result of the distortion, the main factor-binding sites of both tRNAs, the CCA end and the T-stem, overlap and the bases are arranged in register (‘in’). In the T-stem, the bases are shifted by exactly one base pair (‘Δ1’) between the two tRNAs, which shifts the bases out of register (‘out’) in the acceptor stems of the two tRNAs. e, Specific interaction between the extended loop (blue) in SelB domain 3 and the antideterminant box in the T-stem of tRNASec. f, Sequence-specific interaction between the loop (blue) in EF-Tu domain 3 and the T-stem of tRNACys (PDB ID 1B23 ref. 11). Note the different structure of the hairpin-loop as compared to SelB, contributing to the particularly low affinity of EF-Tu for tRNASec (ref. 89) (see also k, middle panel). g, Charge distribution in the amino-acid-binding pockets of SelB (left) and EF-Tu (right). The colour bar on the right denotes the electrostatic potential in kBTe−1. Note the similar geometry of tRNA binding in the two complexes. In SelB, the pocket is positively charged, facilitating the interaction with the negatively charged selenol group of Sec-tRNASec and the discrimination against the precursors of Sec-tRNASec biosynthesis, deacylated tRNASec and Ser-tRNASec, as well as against most canonical tRNAs. In EF-Tu, the amino-acid-binding pocket is negatively charged, thereby discriminating against Sec-tRNASec. h, Recognition of the selenol group in Sec-tRNASec by SelB. The positively charged, highly conserved Arg181 and Arg236 in SelB contact the negatively charged selenol group (Se−, orange), whereas the aromatic ring of Tyr42 stacks onto the selenol group. The importance of these residues was demonstrated by mutational analysis9,90. The universally conserved Asp180, which is also important for Sec-tRNASec binding9, forms a secondary binding shell stabilizing Arg236. i, Solvent-exposed amino-acid-binding site as found in the crystal structure of isolated Aquifex aeolicus (aq) SelB–GDPNP co-crystallized with l-cysteine as Sec mimic (PDB ID 4ZU9)8. The position of Cys deviates by 5 Å from the amino acid position in the present SelB–GDPNP–Sec-tRNASec structure (grey), resulting in a distinct interaction pattern and a more solvent-exposed binding site of the Cys. j, Superpositions of SelB–GDPNP–Sec-tRNASec and EF-Tu–GDPNP–Cys-tRNACys revealing differences in T-stem recognition by the loop in SelB domain 3 versus EF-Tu. Top: The loop structure in SelB is compatible with binding of canonical tRNACys. Other tRNAs with different T stem sequences (particularly at position 63, red arrow) may be unfavourable for the interaction with the conserved Ser327 in SelB. Middle: The loop structure in EF-Tu is incompatible with the conformation of the antideterminant box (orange) in tRNASec. Bottom: In isolated aqSelB (PDB ID 4ZU9)8, the loop adopts a similar conformation as in the SelB ternary complex, suggesting a conserved loop structure that facilitates recognition of tRNASec. k, Experimental density depicting the interaction of SelB with the acceptor and T stem region of tRNASec in the GTPase-activated state (rendered at 2σ) suggesting alternative conformations of Arg329; green mesh, density rendered at 1σ. l, Scheme depicting the functional importance of rearrangements of the tRNASec variable arm for translation elongation after Sec incorporation. (1) Upon release from SelB, tRNASec accommodates in the aminoacyl site in the classical state. The variable arm undergoes a large reorientation from a SelB-bound distorted conformation in the GTPase-activated state to its free ground state conformation in the classical state (left close-up); the latter ground state conformation is similar to the conformation of the variable arm found in complexes of tRNASec with its conversion enzymes, seryl-tRNA synthetase6 and SelA91. (2) After peptide bond formation (resulting in the hybrid state, not shown), peptidyl-tRNASec is translocated to the classical peptidyl site. (3) Binding of the next aa-tRNA (orange) is only compatible with the variable arm adopting the classical state conformation; the GTPase-activated state-conformation of the variable arm would sterically interfere with binding of the next aa-tRNA (right close-up). The long variable arm of canonical class 2 aa-tRNA is not distorted upon EF-Tu binding92 and, consequently, does not require any changes during translation elongation.
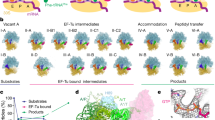
Extended Data Figure 4 The SelB–SECIS interactions on the ribosome and kinetic analysis of GTP hydrolysis.
a, Close-up of the SECIS–ribosome interactions in the GTPase-activated state. Ribosomal proteins S3, S4, and S5 contribute to the helicase activity of the ribosome93. Arg130 and Arg131 of S3 and Lys44 and Arg46 of S4 are important for mRNA unwinding93. Other potential helicase elements are Arg71 and Arg125 in S3, which interact with the base of the SECIS hairpin and may thereby facilitate unwinding of the mRNA secondary structure elements. The mode of SECIS recognition by SelB on the ribosome is similar to that in isolated SelB domain 4-SECIS complexes, including three essential bulged nucleotides in the SECIS (U21, G26 and U27; present data and refs 9, 25 and 72). b, Interaction of SelB domain 4 with the shoulder of the 30S subunit. Winged-helix motif 4 (wh4) of SelB forms salt bridges with protein S4, while wh2 and wh3 embrace helix 16 of 16S rRNA by interacting mainly with the rRNA backbone. c, Concentration dependence of the apparent rate of GTP hydrolysis by SelB measured with increasing concentrations of ribosomes programmed with mRNAs ‘UGA’ (black), ‘iSECIS’ (blue) and ‘UUC’ (red). d, mRNA construct ‘UGA’ used for structural and kinetic analyses containing the cognate UGA codon coding for Sec and the functional minimal fdhF-SECIS. The Sec codon and the essential bases of the SECIS are indicated by boxes; SD, Shine–Dalgarno sequence. e, f, mRNA constructs ‘iSECIS’ and ‘UUC’ used for kinetic analysis. Changes in comparison to the ‘UGA’ mRNA are indicated in blue (iSECIS) and red (UUC).
Extended Data Figure 5 Local and global conformational changes of the 30S subunit.
a, Rearrangements of A1913 in H69 of 23S rRNA. A1913 stabilizes A1492 and A1493 and guides Sec-tRNASec to the mRNA codon by interaction with isopentenyl-A37 (i6A37) of tRNASec. The respective conformation (syn or anti) is denoted; dashed lines indicate potential interactions with distances ≤4 Å. b, Close-up of the distance RASL between the UGA (N1 of G) and anticodon of tRNASec (N3 of C35). Note the potential interaction between the third codon position and the tRNA anticodon in the codon reading state. c, Global conformational changes of the 30S subunit upon accommodation (GTPase-activated to classical) and hybrid state formation (classical to hybrid). tRNA accommodation in the aminoacyl site does not cause major 30S subunit rearrangements, in contrast to the eukaryotic system, where tRNA accommodation correlates with a large-scale conformational change94. Subsequent tRNA hybrid state formation is coupled to the well-known rotational movement of the 30S subunit95. The typical rotational changes all over the 30S subunit are clearly distinct from the changes observed upon domain opening and closure, which are mainly found at the shoulder region (Fig. 3a). The heat map quantifies the movements of 16S rRNA backbone phosphate atoms as obtained by superposition on 23S rRNA of the initial complex state (Fig. 3a). d, Deviations of the 16S rRNA backbone phosphates in the initial binding, codon reading, or GTPase-activated states from the initial complex superimposed on 23S rRNA (top) or 16S rRNA (bottom). Negative values correspond to the 30S domain opening, positive values to domain closure. Landmarks of the 30S subunit are indicated. e, f, Experimental densities of the decoding centre region rendered at 2–2.5σ, if not indicated otherwise. For better visibility, densities for tRNASec were omitted for the GTPase-activated state. e, Anti-conformation of G530 and stacking interaction with C518 of 16S rRNA as seen in the different states. Bottom left: Note the density of tRNASec (purple) suggesting a partial interaction of U34 with the mRNA codon in the codon reading state. f, Different conformational states of A1492, A1493 (red) of 16S rRNA and A1913 (slate blue) of 23S rRNA. Top left: In the initial complex the reduced densities for the three nucleotides indicate a dynamic equilibrium between two states: (1) A1492, A1493 flipped out (‘out’, red) with A1913 in anti-configuration (‘anti’, slate blue) and (2) A1492, A1493 flipped in (‘in’, dark red) with A1913 in syn-configuration (‘syn’, dark slate blue). Top middle: The dynamic nucleotides in the initial complex are discernible at lower threshold (red mesh, density at ~1σ). Top right and bottom left: Density in the initial binding and codon reading states is compatible with the bases of A1492 and A1493 flipped-in and A1913 in the anti-configuration. Bottom right: In the GTPase-activated state, A1492 and A1493 are flipped-out and A1913 remains in the anti-conformation. g, Density for the codon–anticodon interaction in the GTPase-activated state rendered at ~3σ.
Extended Data Figure 6 Molecular dynamics simulations of the free ternary complex.
a, tRNASec dynamics obtained by molecular dynamics simulations of the free SelB–GTP–Sec-tRNASec ternary complex. The free energy landscape for the conformational dynamics of the tRNASec body, excluding the variable arm, is shown as a heat map. The free energy was estimated from separate simulations starting from the structures of the ternary complex bound to the ribosome in initial binding, codon reading, and GTPase-activated states (upper and lower left panels), as well as from all simulations combined (lower right panel). The results of different simulations have a large overlap, especially in regions of low free energy. The crosses denote the tRNA conformations in the ribosome-bound cryo-EM states and the free energy minimum of the free ternary complex found in the molecular dynamics simulations. b, Weak coupling between the conformational dynamics of tRNASec excluding the variable arm (no-v-arm) and of the variable arm (v-arm). Each panel shows the correlation coefficients for a pair of conformational modes (red points, with standard deviation) for the simulations starting from each state (initial binding, codon reading, and GTPase-activated), as well as for all simulations combined. The grey bars denote the standard deviation of the correlation coefficient expected from random drawing of projections. All pairs of modes show a small but significant correlation, indicating a weak coupling. c, Coupling between tRNA dynamics (no-v-arm modes) and SelB motions (SelB domain 1, D1, relative to domain 2, D2). Only a weak overall correlation is seen between no-v-arm mode 1 and D1 mode 2. The three ribosome-bound conformations do not change much with regards to no-v-arm mode 1 (Fig. 3b), which suggests that there is no direct coupling between tRNA dynamics and SelB motions on the ribosome.
Extended Data Figure 7 Role of L11-rRNA arm, protein S12 and the dynamics of SelB domains.
a, Role of L11–rRNA arm in guiding Sec-tRNASec towards H89. Residues of protein L11, H43 and H89 of 23S rRNA are shown in mauve; dashed lines mark interactions with the D loop (DL) and T loop (TL) of tRNASec (purple); arrows denote changes with respect to the preceding state. b, Protein S12 as a pivot for tRNASec movement. S12 His76 interacts with the backbone of tRNASec (residue U69) guiding the tRNA from its position in the codon reading state (grey) to the GTPase-activated state (coloured). c, Inter-subunit rearrangement in SelB upon transition from initial binding to codon reading. Shown is the small movement observed in addition to the rotation of SelB relative to the 30S shoulder as depicted in Fig. 4f, upper panel. d, SelB residues facilitating SRL docking. Left and middle: Spontaneous fluctuations within SelB that are required—in addition to the rotation upon 30S domain closure (shown in Fig. 4, lower panel)—for transition from codon reading (grey) to the GTPase-activated state (red). Middle: 30S domain closure and tRNASec repositioning re-align SelB and in particular its long polar residues towards the SRL. Right: SelB–SRL interactions resulting from docking in the GTPase-activated state. e, Motions of SelB domain 1 (D1) relative to domain 2 (D2) in the molecular dynamics simulations of the free ternary complex. SelB domain 1 can rapidly and spontaneously sample arrangements required for the transitions from initial binding to codon reading and GTPase-activated states. f, Minimum distance between the imidazole ring of SelB His61 and γ-phosphate/oxygens of GTP (RGTP) and the minimum distance between the Van der Waals radii of Val9 and Met36 (RGate), residues adjacent to His61 in SelB and analogous to the presumed ‘gate’ in EF-Tu3. In the free ternary complex, the distance between GTP and His61 of SelB is larger than in the GTPase-activated state on the ribosome, although Val9 and Met36 can freely move apart. Distances for the crystal structure of SelB–GDPNP are labelled 4ZU9 (ref. 8). g, Metastable SelB conformation in the GTPase-activated state. SelB in the GTPase-activated state was taken as initial structure for molecular dynamics simulations. During the first 50 ns of the simulations, the positions of heavy atoms are restrained to allow equilibration of the solvent (top panel). As soon as the restraints are released (50–70 ns), the distance between His61 and GTP increases (middle panel), and residues Val9 and Met36 resume fluctuations between open and closed conformations. h, Comparison of the active sites in SelB and EF-Tu. Top: GTPase centre of SelB in the GTPase-activated state. Bottom: Activated GTPase centre of EF-Tu on the ribosome (PDB ID 4V5L)35.
Rights and permissions
About this article
Cite this article
Fischer, N., Neumann, P., Bock, L. et al. The pathway to GTPase activation of elongation factor SelB on the ribosome. Nature 540, 80–85 (2016). https://doi.org/10.1038/nature20560
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20560
This article is cited by
-
A large expert-curated cryo-EM image dataset for machine learning protein particle picking
Scientific Data (2023)
-
Modulation of translational decoding by m6A modification of mRNA
Nature Communications (2023)
-
Repurposing tRNAs for nonsense suppression
Nature Communications (2021)
-
New Directions for Understanding the Codon Redefinition Required for Selenocysteine Incorporation
Biological Trace Element Research (2019)
-
Visualization of translation termination intermediates trapped by the Apidaecin 137 peptide during RF3-mediated recycling of RF1
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.