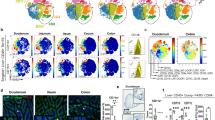
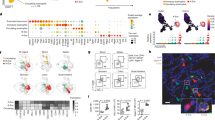
Abstract
Recognition and removal of apoptotic cells by professional phagocytes, including dendritic cells and macrophages, preserves immune self-tolerance and prevents chronic inflammation and autoimmune pathologies1,2. The diverse array of phagocytes that reside within different tissues, combined with the necessarily prompt nature of apoptotic cell clearance, makes it difficult to study this process in situ. The full spectrum of functions executed by tissue-resident phagocytes in response to homeostatic apoptosis, therefore, remains unclear. Here we show that mouse apoptotic intestinal epithelial cells (IECs), which undergo continuous renewal to maintain optimal barrier and absorptive functions3, are not merely extruded to maintain homeostatic cell numbers4, but are also sampled by a single subset of dendritic cells and two macrophage subsets within a well-characterized network of phagocytes in the small intestinal lamina propria5,6. Characterization of the transcriptome within each subset before and after in situ sampling of apoptotic IECs revealed gene expression signatures unique to each phagocyte, including macrophage-specific lipid metabolism and amino acid catabolism, and a dendritic-cell-specific program of regulatory CD4+ T-cell activation. A common ‘suppression of inflammation’ signature was noted, although the specific genes and pathways involved varied amongst dendritic cells and macrophages, reflecting specialized functions. Apoptotic IECs were trafficked to mesenteric lymph nodes exclusively by the dendritic cell subset and served as critical determinants for the induction of tolerogenic regulatory CD4+ T-cell differentiation. Several of the genes that were differentially expressed by phagocytes bearing apoptotic IECs overlapped with susceptibility genes for inflammatory bowel disease7. Collectively, these findings provide new insights into the consequences of apoptotic cell sampling, advance our understanding of how homeostasis is maintained within the mucosa and set the stage for development of novel therapeutics to alleviate chronic inflammatory diseases such as inflammatory bowel disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Poon, I. K., Lucas, C. D., Rossi, A. G. & Ravichandran, K. S. Apoptotic cell clearance: basic biology and therapeutic potential. Nat. Rev. Immunol. 14, 166–180 (2014)
Green, D. R., Ferguson, T., Zitvogel, L. & Kroemer, G. Immunogenic and tolerogenic cell death. Nat. Rev. Immunol. 9, 353–363 (2009)
Blander, J. M. Death in the intestinal epithelium—basic biology and implications for inflammatory bowel disease. FEBS J . 283, 2720–2730 (2016)
Eisenhoffer, G. T. et al. Crowding induces live cell extrusion to maintain homeostatic cell numbers in epithelia. Nature 484, 546–549 (2012)
Bekiaris, V., Persson, E. K. & Agace, W. W. Intestinal dendritic cells in the regulation of mucosal immunity. Immunol. Rev. 260, 86–101 (2014)
Gross, M., Salame, T. M. & Jung, S. Guardians of the gut – murine intestinal macrophages and dendritic cells. Front. Immunol. 6, 254 (2015)
Khor, B., Gardet, A. & Xavier, R. J. Genetics and pathogenesis of inflammatory bowel disease. Nature 474, 307–317 (2011)
Schlitzer, A. et al. IRF4 transcription factor-dependent CD11b+ dendritic cells in human and mouse control mucosal IL-17 cytokine responses. Immunity 38, 970–983 (2013)
Shrimpton, R. E. et al. CD205 (DEC-205): a recognition receptor for apoptotic and necrotic self. Mol. Immunol. 46, 1229–1239 (2009)
Kim, G. H., Dayam, R. M., Prashar, A., Terebiznik, M. & Botelho, R. J. PIKfyve inhibition interferes with phagosome and endosome maturation in macrophages. Traffic 15, 1143–1163 (2014)
Fang, W. F. et al. 5-Lipoxygenase activating protein (FLAP) dependent leukotriene biosynthesis inhibition (MK591) attenuates Lipid A endotoxin-induced inflammation. PLoS One 9, e102622 (2014)
Blander, J. M. A long-awaited merger of the pathways mediating host defence and programmed cell death. Nat. Rev. Immunol. 14, 601–618 (2014)
Arbore, G. & Kemper, C. A novel “complement–metabolism–inflammasome axis” as a key regulator of immune cell effector function. Eur. J. Immunol. 46, 1563–1573 (2016)
Hu, H. & Sun, S. C. Ubiquitin signaling in immune responses. Cell Res . 26, 457–483 (2016)
Lee, M. S., Kim, B., Oh, G. T. & Kim, Y. J. OASL1 inhibits translation of the type I interferon-regulating transcription factor IRF7. Nat. Immunol. 14, 346–355 (2013)
Wakioka, T. et al. Spred is a Sprouty-related suppressor of Ras signalling. Nature 412, 647–651 (2001)
Guo, Z. et al. CD4+CD25+ regulatory T cells in the small intestinal lamina propria show an effector/memory phenotype. Int. Immunol. 20, 307–315 (2008)
Tran, D. Q. et al. GARP (LRRC32) is essential for the surface expression of latent TGFβ on platelets and activated FOXP3+ regulatory T cells. Proc. Natl Acad. Sci. USA 106, 13445–13450 (2009)
Josefowicz, S. Z., Lu, L. F. & Rudensky, A. Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012)
Maldonado, R. A. & von Andrian, U. H. How tolerogenic dendritic cells induce regulatory T cells. Adv. Immunol . 108, 111–165 (2010)
Kim, K. S. et al. Dietary antigens limit mucosal immunity by inducing regulatory T cells in the small intestine. Science 351, 858–863 (2016)
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012)
Moon, C. M. et al. Genetic variants in the IL12B gene are associated with inflammatory bowel diseases in the Korean population. J. Gastroenterol. Hepatol. 28, 1588–1594 (2013)
Guo, X. et al. Disruption of inducible 6-phosphofructo-2-kinase impairs the suppressive effect of PPARγ activation on diet-induced intestine inflammatory response. J. Nutr. Biochem. 24, 770–775 (2013)
Haberman, Y. et al. Pediatric Crohn disease patients exhibit specific ileal transcriptome and microbiome signature. J. Clin. Invest. 124, 3617–3633 (2014)
Bongers, G. et al. The cytomegalovirus-encoded chemokine receptor US28 promotes intestinal neoplasia in transgenic mice. J. Clin. Invest. 120, 3969–3978 (2010)
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem . 277, 33275–33283 (2002)
Jung, S. et al. In vivo depletion of CD11c+ dendritic cells abrogates priming of CD8+ T cells by exogenous cell-associated antigens. Immunity 17, 211–220 (2002)
Saito, M. et al. Diphtheria toxin receptor-mediated conditional and targeted cell ablation in transgenic mice. Nat. Biotechnol. 19, 746–750 (2001)
Bogunovic, M. et al. Origin of the lamina propria dendritic cell network. Immunity 31, 513–525 (2009)
Miller, J. C. et al. Deciphering the transcriptional network of the dendritic cell lineage. Nat. Immunol. 13, 888–899 (2012)
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res . 30, 207–210 (2002)
Bongers, G. et al. Interplay of host microbiota, genetic perturbations, and inflammation promotes local development of intestinal neoplasms in mice. J. Exp. Med. 211, 457–472 (2014)
Cummings, R. J. et al. Exposure to ionizing radiation induces the migration of cutaneous dendritic cells by a CCR7-dependent mechanism. J. Immunol. 189, 4247–4257 (2012)
Shi, C. & Pamer, E. G. Monocyte recruitment during infection and inflammation. Nat. Rev. Immunol. 11, 762–774 (2011)
Acknowledgements
We are grateful to S. V. Chittur and M. Kuentzel, SUNY at the Albany Center for Functional Genomics. We thank M. Bogunovic at Pennsylvania State University, S. Jung at the Weizmann Institute of Science, Blander Laboratory members, J. Ochando and C. Bare at the Icahn School of Medicine Flow Cytometry Core, and M. A. Blander and S. J. Blander for discussions, help, and support. This work was supported by institutional seed funds to J.M.B. J.M.B. and her laboratory were supported by NIH grants AI095245, AI123284, DK072201, the Burroughs Wellcome Fund, and the Leukemia and Lymphoma Society. R.J.C. was supported by NIH training grants 2T32A1007605-11 and 5T32DK007792-12. G.Ba. was supported by the Crohn’s and Colitis Foundation of America (CCFA) Research Fellowship Award. B.M.H: NIAID contract HHSN272201000054C and U19 AI117873. J.C.: R01 DK092235, U01 DK62429, U01 DK062422, philanthropic SUCCESS, Sanford J. Grossman Charitable Trust. S.A.L. and G.C.F.: NIH 5P01DK072201-09 and 5R01CA161373-04, CCFA 330239, and SUCCESS. G.Bo.: Jenna and Paul Segal grant.
Author information
Authors and Affiliations
Contributions
R.J.C. and J.M.B: designed the study and wrote the manuscript. R.J.C. conducted most experiments; G.Ba. performed the initial set-up, protocol optimization and provided sorting expertise; L.M. performed initial VDTR characterizations; G.Bo. provided microarray and statistical analysis expertise; B.M.H. performed ImageStream acquisition and analyses; J.C. and K.G. performed IBD GWAS data comparison with differentially expressed phagocyte genes; G.C.F. and S.A.L.: VDTR strain derivation and data discussions. J.M.B. conceived the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 No gross anatomical changes in the small or large intestine of VDTR mice.
a, The primate diphtheria toxin receptor (DTR)–eGFP transgene driven by the mouse villin promoter (mVillin). b, qRT–PCR for relative expression of primate diphtheria toxin receptor (HBEGF (‘DTR’)) mRNA in both the small and large intestine of four different DTR–eGFP transgenic (VDTR) founder lines. Founder 32p was selected to propagate the VDTR line on the basis of specific transgene expression in the intestinal tract and not in other organs. c, d, Haematoxylin and eosin staining (H&E) of small intestine and large intestine paraffin sections from treated VDTR (c) and untreated C57BL/6 (B6) and VDTR (d) as indicated. e–i, Immunofluorescence of small intestine (e, f, h) and large intestine (g, i) cryo-sections from indicated mice after staining with phalloidin or pan-cytokeratin and DAPI as indicated. j, TUNEL (eGFP-quenched after paraffin embedding). p(A), polyadenylation. Scale bars, 100 μm (c, d, j), 50 μm (e–i).
Extended Data Figure 2 The intestinal microenvironment does not become inflamed following 2 ng g−1 diphtheria toxin administration.
a, Haematoxylin and eosin stain of paraffin-embedded sections from small intestine of VDTR mice at 4 h following PBS or diphtheria toxin administration. b, Flow cytometry of SILP cells from VDTR mice treated with PBS or diphtheria toxin. Pre-gated on live CD45+MHCII+ cells. Numbers above gates indicate percentage of positively stained cells. Quantification is shown in bottom panels. IM, inflammatory monocytes. We noted no tissue destruction or discernible inflammation (a), nor infiltration of Ly6Chi monocytes35 (b) at 4 h with either dose of diphtheria toxin, in contrast to infiltration of these cells at 16 h following 10 ng g−1 diphtheria toxin. n = 9 mice for PBS, n = 6 mice for diphtheria toxin at 1–4 h, and n = 3 for PBS and diphtheria toxin at 16 h. One-way ANOVA; **P < 0.01; NS, not significant. Flow cytometry gates are representative of at least three independent experiments. Data are mean ± s.e.m. c, d, In situ hybridization with a eubacterial probe on large intestine cryo-sections following PBS or diphtheria toxin administration to VDTR mice (c), or 3% dextran sodium sulphate (DSS) treatment of B6 mice for 5 days (d). c, Arrowheads indicate presence of eubacterial probe in SILP. There was no bacterial translocation to the intestinal lamina propria at 4-h post-treatment with either 2 ng g−1 or 10 ng g−1 diphtheria toxin, as evidenced by luminal confinement of the in-situ-hybridized eubacterial probe signal, and similar to that in PBS-treated controls. e, Immunofluorescence on small-intestine paraffin sections stained with antibodies to CC3 at 4 h following PBS injection. f, Quantification from e and Fig. 1c. Scale bars, 100 μm (a, e), 25 μm (c, d).
Extended Data Figure 3 Ileum lamina propria CD11c+ phagocytes extend dendrites towards apoptotic intestinal epithelial cells.
a–h, Conventional (a–e) and confocal (f–h) whole-mount microscopy of excised ileum from B6 or VDTR mice following PBS or diphtheria toxin as indicated. b, c, e, eGFP not overlaid. Insets from f and h depicted in bottom and right panels, respectively, without eGFP overlay. L, lumen. Arrowhead in a points to a CC3+ IEC; in d, to a CD11c+ dendrite. Scale bars, 25 μm (a, e), 50 μm (b, d, h), 100 μm (c, f, g).
Extended Data Figure 4 Flow cytometry gating strategy for identifying CD11c+ and CD11c−/lo phagocytes in the SILP of VDTR mice.
a–e, g, Flow cytometric analyses of SILP cells from VDTR mice. Numbers indicate the percentage of gated populations. a, Gating strategy for total CD11c+ phagocytes. b, Identification of the eGFP+ gate on the basis of the small intestine cellular profile from B6 (eGFP−) mice. c, CD103 and CD11b expression on gated cells in b. d, Monocytes pre-gated on live CD45+CD11cloMHCIIhiCD11b+Ly6Chi cells. e, Granulocytes pre-gated on live CD45+CD11c−MHCII−CD11b+Ly6Chi cells. f, Percentage of cells and absolute numbers from b. g, Dendritic cells and macrophages pre-gated on live CD45+MHCIIhiCD11c+ cells, further identified as CD64− and CD64+, respectively. Gating on differential CD64 expression, followed by delineation of CD103 and CD11b expression, also distinguished two macrophage and three dendritic cell populations5. Data represent at least three independent experiments and in f, n = 3 B6 mice; n = 6 VDTR mice treated with 2 ng g−1 diphtheria toxin for 1–4 h; n = 9 VDTR mice treated with PBS; one-way ANOVA; *P < 0.05. Data are mean ± s.e.m. h, Schematic of SILP phagocytes that sample apoptotic IECs (green).
Extended Data Figure 5 eGFP+ and eGFP−CD11c+ phagocyte populations exhibit distinct transcriptional profiles.
a, FACS-sorted dendritic cell and macrophage populations from the SILP of VDTR mice at 4 h following diphtheria toxin administration using the gating strategy described in Fig. 2f, which included CD24 and CD64. b, Principal Component Analysis (PCA) of the 1,534 genes (ANOVA Q < 0.05; 4.8% of total) with most variable expression in dendritic cell and macrophage (Mϕ) subsets, with (eGFP+) or without (eGFP−) apoptotic IEC cargo. PCA-separated macrophages from dendritic cells (left panel) and eGFP+ from eGFP− (right panel). We confirmed the purity and identity of sorted phagocytes on the basis of relative expression of genes encoding the molecules used for FACS sorting, as well as macrophage and dendritic-cell-specific genes. c–g, Signal intensity of genes encoding MHCII (H2-Ab1) and CD11c (Itgax), shared by both dendritic cells and macrophages, the macrophage-specific Csf1 receptor (Csf1r), and the dendritic-cell-specific kinase Flt3 (Flt3) in CD11c+ phagocytes31 (c); molecules used for FACS sorting (d); macrophage markers (e); dendritic cell markers (f); and IEC markers from analyses of microarray experiments (g). It should be noted that IEC-specific genes were not increased in eGFP+ compared to eGFP− phagocytes indicating negligible contamination by IEC-specific transcripts. Data represent five independent experiments with four mice per experiment and three biological replicates. White and black bars indicate expression from eGFP− and eGFP+ phagocytes, respectively. h, Validation by qRT–PCR for IEC-specific genes including Muc2 (mucin-2) and Elf3 (E74-like factor 3), which were not expressed by phagocytes but readily detectable in sorted IECs. Conversely, phagocyte-specific transcripts (Aldh1a2, Siglec1 and Timd4)31 were expressed in sorted phagocytes but not in IECs. Data represent two independent experiments depicting relative expression (log2). i, qRT–PCR quantification of relative eGFP expression and the primate diphtheria toxin receptor (HBEGF or 'DTR') from VDTR IECs, spleen cells, and eGFP− and eGFP+ phagocytes. As expected, expression of the eGFP and DTR (HBEGF) transgenes were confined to IECs in VDTR mice and not present in sorted dendritic cell or macrophages. Data represent two independent experiments depicting fold change over C57BL/6J litter mate controls. j, qRT–PCR relative expression (log2) of inflammatory transcripts from indicated eGFP− SILP macrophages over time. The unchanged expression of Il1b, Il6, and Tnf in eGFP−CD11b+ and CD103+CD11b+ macrophages, which served as sentinels of the microenvironment, provided further evidence for lack of inflammation induction by low-dose 2 ng g−1 diphtheria toxin. Note that the unique transcriptional profiles of eGFP+ phagocytes may belong to pre-existing subpopulations that become detectable only when marked by eGFP, owing to an inherently superior capacity for apoptotic cell sampling. However, we consider it more likely that the differential gene expression patterns between eGFP+ and eGFP− populations are a consequence of apoptotic IEC internalization. Data represent three independent analytical repeats and one biological replicate consisting of three mice per time point. Data are mean ± s.e.m.
Extended Data Figure 6 Further validation of the differential gene expression profiles following apoptotic IEC sampling.
a, Flow cytometry validation for MER and CD205 (Ly75) protein upregulation either in the steady state or 4 h after diphtheria toxin administration, reflecting transcript data shown in Fig. 3d. Flow cytometry representative of at least three independent experiments. b, g, Hierarchical clustering of differentially expressed ‘find me/eat me’ receptor (b) and ‘suppression of immune response’ (g) genes at 4 h after diphtheria toxin administration. Differences in expression did (g) and did not (b) meet statistical significance of at least 1.2-fold (ANOVA (Q < 0.05) and Tukey’s HSD post-hoc test (P < 0.05); −1.2> fold > 1.2) comparing eGFP+ and eGFP− phagocytes at 4 h following diphtheria toxin administration; expressed by row Z scale and relative expression (log2). In b, upregulated genes correlating with eGFP content included Cx3cr1 (the receptor for the ‘find me’ signal fractalkine) and TAM family receptor Axl by CD11b+ macrophages, as well as Tyro3 by CD103+ dendritic cells. Different members of the Stabilin family of scavenger receptors, Adgrb1, Stab1/2, required for apoptotic cell clearance1, as well as Timd4, which binds to exposed phosphatidylserine, were differentially expressed by macrophages and CD103+ dendritic cells. Data represent five independent experiments with four mice per experiment and three biological replicates. Many genes in b and g remained differentially regulated at 6 h after diphtheria toxin administration (see d and e), showing a stable profile not unique to the 4-h time point. c, qRT–PCR validation for Axl and Timd4 in SILP phagocytes at 4 h after diphtheria toxin administration. d, e, qRT–PCR quantification of relative gene expression in the indicated eGFP− and eGFP+ populations (log2). CD11b+ macrophages (d) and CD103+ dendritic cells (e) from the SILP at 6 h following diphtheria toxin administration. Expression of Ccr7, encoding the chemotactic receptor required for migration into mesenteric lymph nodes where priming of naive T cells occurs, was higher in SILP eGFP+CD103+ dendritic cells, but not macrophages, relative to their eGFP− counterparts (Extended Data Fig. 8c), and remained high in eGFP+CD103+ dendritic cells at 6 h after diphtheria toxin administration (e) compared to the downregulation by eGFP+ cells relative to eGFP−CD11b+ macrophages (d). Data represent two independent analytical repeats and one biological replicate consisting of three mice. f, Signal intensity of Cd300a, encoding an inhibitory receptor reported to bind apoptotic cell-exposed phosphatidylserine and suppress commensal-driven IFNβ production by large intestine lamina propria CD103−CD11b+CX3CR1+F4/80lo–int cells3, from microarray analyses conducted at 4 h after diphtheria toxin administration, showing similar levels in SILP eGFP+ and eGFP− macrophages and CD103+ dendritic cells indicated on the x axis. Data did not meet statistical significance of at least 1.2-fold (ANOVA (Q < 0.05) and Tukey’s HSD post-hoc test (P < 0.05); −1.2> fold > 1.2) and represent five independent experiments with 4 mice per experiment and three biological replicates. Data are mean ± s.e.m. White and black bar graphs indicate expression from eGFP− and eGFP+ phagocytes, respectively. h, i, Microarray validation by duplicate qRT–PCR for indicated genes represent three independent experiments.
Extended Data Figure 7 Inflammatory cytokine and co-stimulatory genes in CD11c+ phagocytes are unchanged and decreased at 4 h and 6 h, respectively, following sampling of apoptotic IECs.
Signal intensity of genes encoding inflammatory cytokines (a), and co-stimulatory molecules (b), from microarray analyses conducted 4 h after diphtheria toxin administration on the SILP phagocyte populations indicated on the x axes where CD103CD11b and CD11b denote CD103+CD11b+ and CD11b+ macrophages, and CD103 denotes CD103+ dendritic cells. White and black bar graphs indicate expression from eGFP− and eGFP+ phagocytes, respectively. No statistically significant changes were observed as genes did not meet statistical significance of at least 1.2 fold (ANOVA (Q < 0.05) and Tukey’s HSD post-hoc test (P < 0.05); −1.2> fold > 1.2). Data represent five independent experiments with four mice per experiment and three biological replicates. c, d, qRT–PCR quantification of Il1b and Il6 transcripts expressed by eGFP+ and eGFP−CD11b+ macrophages (c) and CD103+ dendritic cells (d) from the SILP at 6 h after diphtheria toxin administration. Data represent three independent analytical repeats and one biological replicate consisting of three mice.
Extended Data Figure 8 Resident phagocytes in the mesenteric lymph node do not carry eGFP+ apoptotic IEC cargo.
a, Duplicate qRT–PCR quantification of Aldh1a2 transcript levels. b, Flow cytometry validation for CD274 protein expression at steady state or at 4 h after diphtheria toxin administration reflecting transcript data shown in Fig. 4a. Data represent at least three independent experiments. c, l, Signal intensity of Ccr7 (c) and Tgfb1 (l) from microarray analyses. Expression of Ccr7 was higher in SILP eGFP+CD103+ dendritic cells but not macrophages, relative to their eGFP− counterparts. Although Ccr7 transcripts showed a higher expression trend in CD103+ dendritic cells, relative expression of Ccr7 and Tgfb1 did not meet statistical significance of at least 1.2-fold (ANOVA (Q < 0.05) and Tukey’s HSD post-hoc test (P < 0.05); −1.2 > fold > 1.2). Data represent five independent experiments with four mice per experiment and three biological replicates. d–g, Flow cytometric analyses of MLN cells from naive, untreated B6 and VDTR mice. d, e, ‘Migratory’ cells were pre-gated on live CD45+MHCIIhiCD11c+ cells. Quantification in right (d) and bottom (e) panels. f, ‘Resident’ cells were pre-gated on live CD45+MHCII+CD11c+ cells and these cells were eGFP−. n = 3 mice per experiment, n = 6 mice per group; unpaired two-tailed t-test with Welch’s correction; ***P < 0.001. e, g, eGFP− and eGFP+ cells from d were further sorted on the basis of CD64, CD24, CD103 and CD11b expression. e, CD24+CD103+ dendritic cells from B6 control mice did not contain the eGFP label. g, The minor populations (~2%) of CD64+CD103+CD11b+ and CD64+CD11b+ macrophages within the migratory gate5 were also eGFP−. Data in e, g, represent three independent experiments. h–k, Flow cytometry analyses of migratory eGFP− and eGFP+ phagocytes in the MLN over time following a single injection of diphtheria toxin. Increasing the number of apoptotic IECs with diphtheria toxin treatment did not lead to a statistically significant increase in the frequency or absolute number of the indicated macrophage populations (h and i, respectively) or eGFP+CD103+ dendritic cells (j and k, respectively) over time within migratory MLN cells. This was expected given the non-inflammatory nature of the low dose diphtheria toxin and the small number (~0.5–2 × 104) of total CD103+ dendritic cell within the SILP, of which an even smaller fraction (~10%) are eGFP+ (Fig. 2d). Data represent four independent experiments with 4 mice per time point. Data mean ± s.e.m. m, qRT–PCR for Tgfb1 from indicated cell populations. Data represent three independent analytical repeats and one biological replicate consisting of 3 mice. n, Flow cytometry for surface CD40 expression on migratory eGFP+ and eGFP− MLN CD103+ dendritic cells from untreated VDTR mice representative of four independent experiments.
Extended Data Figure 9 Naive CD4+ T cells co-cultured with eGFP+ or eGFP−CD103 dendritic cells from MLN do not differentiate into T-bet+, GATA-3+, or RORγt+ cells.
a, b, d, Flow cytometric analyses of VDTR MLN. a, b, Gating strategy for total CD45+CD3+CD4+ T cells. b, T cells pre-gated as in a from non-treated and 3 days post-diphtheria-toxin-treated VDTR mice. Peripherally induced Treg were identified as FOXP3+Helios− cells. c, Flow cytometric analyses of splenic CD4+ T cells cultured ex vivo at 1 × 105 after enrichment of naive CD4+ T cells, for 5 days with FACS-sorted 1 × 104 eGFP+ or eGFP−CD103+ MLN dendritic cells. Data represent two independent experiments. d, MLN CD4+ T cells from non-treated and day 3 post-diphtheria-toxin-treated VDTR mice. T-bet, GATA-3 and RORγt, characteristic of TH1, TH2 and TH17 cells, respectively, were not present in ex vivo co-cultures (c) or in the MLN of diphtheria-toxin-treated mice (d). At least three independent experiments were performed for flow cytometry studies. Numbers in plots indicate the percentage of positively stained cells within each gate.
Extended Data Figure 10 Inflammation does not alter the population of phagocytes that traffics apoptotic IEC cargo to MLN, but does alter its gene expression.
B6 and VDTR mice for 5 days received drinking water supplemented with 3% DSS (yellow highlight in b, d). On day 5, DSS treatment was stopped and replaced with fresh drinking water. a, c, Paraffin haematoxylin and eosin sections from the large intestine (a), and small intestine (c) of indicated mice. Scale bars, 100 μm. b, Change in weight shown on the y axis monitored daily as shown on the x axis. DSS treatment of VDTR mice (green line) was accompanied by weight loss similar to that observed in B6 mice (black line), with minimal damage to the terminal ileum, as expected. n = 4 mice per group. d, f, qRT–PCR for indicated transcripts expressed by migratory eGFP+CD103+ MLN dendritic cells from untreated and day 5 DSS-treated VDTR mice (d), and eGFP− and eGFP+CD103+ MLN dendritic cell isolated from day 5 DSS treated VDTR mice (f). #, not detected. Data represent three independent analytical repeats and one biological replicate consisting of two mice. e, Flow cytometry analyses of MLN migratory cells from day-5 DSS-treated and non-DSS-treated mice. Data represent two independent experiments consisting of two mice. Numbers near gates indicate the percentage of positively stained cells.
Supplementary information
Supplementary Information
This file contains a glossary of gene names in alphabetical order. (PDF 104 kb)
Rights and permissions
About this article
Cite this article
Cummings, R., Barbet, G., Bongers, G. et al. Different tissue phagocytes sample apoptotic cells to direct distinct homeostasis programs. Nature 539, 565–569 (2016). https://doi.org/10.1038/nature20138
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20138
This article is cited by
-
Regional differences in the ultrastructure of mucosal macrophages in the rat large intestine
Cell and Tissue Research (2024)
-
Interferon-Driven Immune Dysregulation in Common Variable Immunodeficiency–Associated Villous Atrophy and Norovirus Infection
Journal of Clinical Immunology (2023)
-
αvβ8 integrin-expression by BATF3-dependent dendritic cells facilitates early IgA responses to Rotavirus
Mucosal Immunology (2021)
-
Cell death in the gut epithelium and implications for chronic inflammation
Nature Reviews Gastroenterology & Hepatology (2020)
-
Efferocytosis in health and disease
Nature Reviews Immunology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.