Abstract
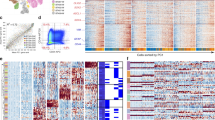
Although human tumours are shaped by the genetic evolution of cancer cells, evidence also suggests that they display hierarchies related to developmental pathways and epigenetic programs in which cancer stem cells (CSCs) can drive tumour growth and give rise to differentiated progeny1. Yet, unbiased evidence for CSCs in solid human malignancies remains elusive. Here we profile 4,347 single cells from six IDH1 or IDH2 mutant human oligodendrogliomas by RNA sequencing (RNA-seq) and reconstruct their developmental programs from genome-wide expression signatures. We infer that most cancer cells are differentiated along two specialized glial programs, whereas a rare subpopulation of cells is undifferentiated and associated with a neural stem cell expression program. Cells with expression signatures for proliferation are highly enriched in this rare subpopulation, consistent with a model in which CSCs are primarily responsible for fuelling the growth of oligodendroglioma in humans. Analysis of copy number variation (CNV) shows that distinct CNV sub-clones within tumours display similar cellular hierarchies, suggesting that the architecture of oligodendroglioma is primarily dictated by developmental programs. Subclonal point mutation analysis supports a similar model, although a full phylogenetic tree would be required to definitively determine the effect of genetic evolution on the inferred hierarchies. Our single-cell analyses provide insight into the cellular architecture of oligodendrogliomas at single-cell resolution and support the cancer stem cell model, with substantial implications for disease management.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Kreso, A. & Dick, J. E. Evolution of the cancer stem cell model. Cell Stem Cell 14, 275–291 (2014)
Lathia, J. D., Mack, S. C., Mulkearns-Hubert, E. E., Valentim, C. L. & Rich, J. N. Cancer stem cells in glioblastoma. Genes Dev . 29, 1203–1217 (2015)
Friedmann-Morvinski, D. et al. Dedifferentiation of neurons and astrocytes by oncogenes can induce gliomas in mice. Science 338, 1080–1084 (2012)
Louis, D. N., Ohgaki, H., Wiestler, O. D. & Cavenee, W. K. WHO Classification of Tumors of the Central Nervous System 4th edn (IARC, 2016)
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nature Protocols 9, 171–181 (2014)
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014)
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nature Neurosci . 17, 131–143 (2014)
Rousseau, A. et al. Expression of oligodendroglial and astrocytic lineage markers in diffuse gliomas: use of YKL-40, ApoE, ASCL1, and NKX2-2. J. Neuropathol. Exp. Neurol. 65, 1149–1156 (2006)
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014)
Feng, W. et al. The chromatin remodeler CHD7 regulates adult neurogenesis via activation of SoxC transcription factors. Cell Stem Cell 13, 62–72 (2013)
Ikushima, H. et al. Autocrine TGF-β signaling maintains tumorigenicity of glioma-initiating cells through Sry-related HMG-box factors. Cell Stem Cell 5, 504–514 (2009)
Suvà, M. L. et al. Reconstructing and reprogramming the tumor-propagating potential of glioblastoma stem-like cells. Cell 157, 580–594 (2014)
Rheinbay, E. et al. An aberrant transcription factor network essential for Wnt signaling and stem cell maintenance in glioblastoma. Cell Reports 3, 1567–1579 (2013)
Suvà, M. L., Riggi, N. & Bernstein, B. E. Epigenetic reprogramming in cancer. Science 339, 1567–1570 (2013)
Miller, J. A. et al. Transcriptional landscape of the prenatal human brain. Nature 508, 199–206 (2014)
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015)
Sugiarto, S. et al. Asymmetry-defective oligodendrocyte progenitors are glioma precursors. Cancer Cell 20, 328–340 (2011)
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016)
Shin, J. et al. Single-cell RNA-seq with waterfall reveals molecular cascades underlying adult neurogenesis. Cell Stem Cell 17, 360–372 (2015)
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015)
Kowalczyk, M. S. et al. Single-cell RNA-seq reveals changes in cell cycle and differentiation programs upon aging of hematopoietic stem cells. Genome Res . 25, 1860–1872 (2015)
The Cancer Genome Atlas Research Network Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372, 2481–2498 (2015)
Lange, C. & Calegari, F. Cdks and cyclins link G1 length and differentiation of embryonic, neural and hematopoietic stem cells. Cell Cycle 9, 1893–1900 (2010)
Koyama-Nasu, R. et al. The critical role of cyclin D2 in cell cycle progression and tumorigenicity of glioblastoma stem cells. Oncogene 32, 3840–3845 (2013)
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nature Biotechnol . 30, 413–421 (2012)
Bettegowda, C. et al. Mutations in CIC and FUBP1 contribute to human oligodendroglioma. Science 333, 1453–1455 (2011)
Padul, V., Epari, S., Moiyadi, A., Shetty, P. & Shirsat, N. V. ETV/Pea3 family transcription factor-encoding genes are overexpressed in CIC-mutant oligodendrogliomas. Genes Chromosom. Cancer 54, 725–733 (2015)
Liu, C. et al. Mosaic analysis with double markers reveals tumor cell of origin in glioma. Cell 146, 209–221 (2011)
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nature Biotechnol . 33, 495–502 (2015)
Mohapatra, G. et al. Glioma test array for use with formalin-fixed, paraffin-embedded tissue: array comparative genomic hybridization correlates with loss of heterozygosity and fluorescence in situ hybridization. J. Mol. Diagn. 8, 268–276 (2006)
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011)
Shalek, A. K. et al. Single-cell RNA-seq reveals dynamic paracrine control of cellular variation. Nature 510, 363–369 (2014)
Whitfield, M. L. et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res . 20, 1297–1303 (2010)
Cibulskis, K. et al. ContEst: estimating cross-contamination of human samples in next-generation sequencing data. Bioinformatics 27, 2601–2602 (2011)
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nature Biotechnol . 31, 213–219 (2013)
Costello, M. et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res . 41, e67 (2013)
Ramos, A. H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–E2429 (2015)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Acknowledgements
We thank L. Gaffney for graphic support. This work was supported by grants from the National Brain Tumor Society (to M.L.S. and D.N.L.), the Smith Family Foundation (to M.L.S.), NIH-NCI SPORE on brain cancer Career Enhancement Project and Developmental Research Project (to M.L.S.), the Broad Institute Broadnext10 program (to M.L.S. and O.R.R.), the American Cancer Society (to M.L.S.) and start-up funds from the MGH department of Pathology. A.S.V. was supported by the NIH R25 fellowship (NS065743) and research grants from the American Brain Tumor Association and Neurosurgery Research and Education Foundation. I.T. was supported by a Human Frontier Science Program fellowship and a Rothschild fellowship. A.R. was supported by funds from the Howard Hughes Medicine Institute, the Klarman Cell Observatory, STARR cancer consortium, NCI grant 1U24CA180922, by the Koch Institute Support (core) grant P30-CA14051 from the National Cancer Institute, the Ludwig Center and the Broad Institute. A.R. is a scientific advisory board member for ThermoFisher Scientific and Syros Pharmaceuticals and a consultant for Driver Group. Flow cytometry and sorting services were supported by shared instrumentation grant 1S10RR023440-01A1. M.M. was supported by the California Institute of Regenerative Medicine (CIRM) grants RB4-06093 and RN3-06510 and the Virginia and D.K. Ludwig Fund for Cancer Research.
Author information
Authors and Affiliations
Contributions
I.T., A.S.V., A.R. and M.L.S. conceived the project, designed the study, and interpreted results. A.S.V., C.H., L.E.E. and C.N. collected single cells and generated single-cell sequencing data. I.T. performed computational analyses. J.M.Fra, K.Y. and G.G. provided support for genomic and genetic analyses. J.M.Fis, C.R. and K.J.L. designed and performed qPCR experiments. C.C.L. and R.M. provided flow cytometry expertise. C.M. and M.M. developed normal human cell cultures used in the study. N.D., N.R., M.N.R., M.L.O. and A.J.I. performed in situ hybridization and FISH experiments. A.P.P., A.A.P., D.G., B.I., J.N., R.S., M.G.F., B.V.N., D.P.C., W.T.C., R.L.M., M.P.F., O.R.R., T. R.G., B.E.B. and D.N.L. provided experimental and analytical support. I.T., A.R. and M.L.S. wrote the manuscript with feedback from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks P. Dirks, J. Rich and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Single-cell RNA-seq analysis of human oligodendroglioma samples.
a, Experimental workflow. b, Clinical information of the main and validation patient cohorts analysed in this study. Asterisk indicates a borderline result of chromosome 1p loss based on clinical testing. c, ISH (top left) and FISH (all other panels) in a representative tumour (MGH36). All our cases retain ATRX protein expression by ISH (top left) and show loss of chromosomes arms 1p (bottom left) and 19q (top right) by FISH. In addition, tumour-specific CNVs identified by single-cell RNA-seq were confirmed by FISH (for example, loss of chromosome 4 in MGH36, bottom right panel). d, Distributions of the total number of sequenced paired-end reads per cell (grey) and of paired-end reads that were mapped to the transcriptome and used to quantify gene expression (black).
Extended Data Figure 2 Diversity of expression programs in oligodendroglioma.
a, Two populations of non-cancer cells identified in oligodendroglioma. Selected genes that are differentially expressed among the two populations of non-cancer cells that lack CNVs (Fig. 1b, top), including markers of microglia (top) and oligodendrocytes (bottom). b, Expression programs in microglia cells from three tumours. The heat map shows relative expression of genes (rows) across microglia cells (columns). Above the dashed line are microglia markers expressed in all microglia cells and below the line are the genes of a microglia activation program, which is variably expressed, and includes cytokines, chemokines, early response genes and other immune effectors. This latter gene set might reflect a microglia activation program that could either be a general microglia program or potentially specific to the context of oligodendroglioma. Microglia cells (n = 235) (columns) are rank ordered by their relative expression of the activation program. The tumour of origin of each cell is colour-coded as indicated in the top row. c, PC2 and PC3 are associated with intermediate values of PC1. PC1 scores are shown along with PC2 (top) and PC3 (bottom) scores for cells in each of the three tumours profiled at high depth. The red line indicates local weighted regression (LOWESS) with a span of 5%, which demonstrates that PC2 and PC3 values tend to be highest in intermediate values of PC1 and to decrease in either high PC1 (that is, oligodendrocyte-like cells) or low PC1 (that is, astrocyte-like cells). d, Consistency of PCA across tumours. Shown are the Pearson correlations in gene loadings (over all analysed genes) between the top three PCs in PCA of the three tumours profiled at high depth (y axis, as shown in Fig. 1) and the top four PCs in alternative PCA of either all six tumours (left), as well as of PCA of each individual tumour (right). PC1–3 are highly consistent between the three-tumour and six-tumour PCAs (R > 0.9); PC1 is highly consistent (R > 0.8) between the three-tumour analysis and all other analysis. e, PC1 (x axis) and PC2 plus PC3 (y axis) scores of malignant cells from each of the three tumours profiled at intermediate depth, showing consistent patterns with those shown in Fig. 1d. f, Distribution of differences in PC1 loadings between the original PCA and the shuffled PCA (see description in the Methods section for principal component analysis) for all genes (black), oligodendrocyte-like (OC-like) genes (blue) and astrocyte-like (AC-like) genes (green). This analysis demonstrates that oligodendrocyte-like and astrocyte-like gene sets are highly skewed in the original PCA and their loadings are not recapitulated by shuffled data reflecting the effect of complexity.
Extended Data Figure 3 The stemness program in oligodendroglioma.
a, Cell–cell correlation matrix based on all analysed genes across all malignant cells in MGH54 (n = 1,174). Cells are ordered by average linkage hierarchical clustering, and coloured boxes indicate distinct clusters. Clusters are marked based on the identity of differentially expressed genes as OC-like (blue), AC-like (yellow), cycling (pink) stem-like (purple) and intermediate cells that do not score highly for any of those expression programs (orange). b, Most differentially expressed genes. Shown is the average expression in each of the OC-like, AC-like, stem-like and intermediate cell clusters (columns) of differentially expressed genes (rows) defined by comparing cells from each of the OC-like, AC-like and stem-like clusters to cells from the remaining clusters with a t-test. Similar genes are highlighted as in Fig. 1 (OC-like: OMG, OLIG1, OLIG2, SOX8; AC-like: ALDOC, APOE, SOX9; Stem-like: SOX4, SOX11, CCND2, SOX2). Stem-like genes also include CTNNB1, USP22, and MSI1. c, Overlap with human GBM stemness program. We previously6 identified a GBM stemness program and determined the association of each gene with that program by the correlation between the expression of that gene and the average expression of the stemness program’s genes across individual cells (‘CSC gradient’) in each of five GBM tumours. Shown is the average correlation (x axis) of each analysed gene (green dots) across the five cases and the P values of those correlations as determined with a t-test (y axis). Genes identified in the oligodendroglioma stemness program (this work) are marked in black and are significantly enriched for the GBM stemness genes (1.5 × 10−4, hypergeometric test), defined as those with P < 0.05 and an average correlation above 0.1. d, Preferential expression of the oligodendroglioma stemness program in neurons but not in OPCs. Genes expressed in the oligodendroglioma single cells were divided into six bins (bars) based on their relative expression (log2-ratio) in stem-like cells with high PC2/3 and intermediate PC1 scores compared to all other cells. Each panel shows for each bin the average relative expression in each of three normal brain cell types (y axis) based on data from the Barres laboratory RNA-seq database9,18: mice oligodendrocyte progenitor cells (mOPC, top), mouse neurons (mNeurons, middle), and human neurons (hNeurons, bottom). Relative expression of each gene in each cell type was defined as the log2-ratio between the respective cell type divided by the average over AC, OC and neurons. Error bars denote standard error as defined by bootstrapping. Asterisks denote bins with significantly different relative expression (in the respective normal cell type) compared to all genes expressed in oligodendroglioma, based on P < 0.001 (by t-test) and average expression change of at least 30%. e, Correlation with mouse activated NSC program. Shown is the distribution of correlation values (x axis) of either all genes (grey) or genes from the oligodendroglioma stemness program (black) with the expression program of mice NSC activation states, as previously quantified by ‘pseudotime’, across single mouse NSCs19. The average correlation of the NSC activation program genes with oligodendroglioma stemness genes is significantly higher than with all other genes (P = 3 × 10−6; t-test). f, Correlation with human NPC program. Shown is the distribution of correlation values (x axis) of either all genes (grey) or genes from the oligodendroglioma stemness program (black) with an expression program of human NPCs identified by PCA (Extended Data Fig. 4). Each gene’s correlation to the average expression of the NPC program genes was calculated across single human NPCs. The average correlation with oligodendroglioma stemness genes is significantly higher than with all other genes (P = 2 × 10−35, t-test).
Extended Data Figure 4 Analysis of human NPCs.
a–d, Differentiation potential of human SVZ NPCs. Human SVZ NPCs isolated from 19-week-old fetuses form neurospheres in culture (a), and can be differentiated to neuronal (neurofilament, b), oligodendrocytic (OLIG2, c), or astrocytic (GFAP, d) lineages in vitro. Scale bars, 25 μm (a), 10 μm (b–d). We note that although OLIG2 can represent different cell types, it is expressed at a low level in the fetal NPCs before differentiation (an average log2(TPM + 1) of 0.82, compared to a threshold of 4 that we use to define expressed genes in our analysis, and with zero cells with expression above this threshold). Thus, the undifferentiated NPCs do not express OLIG2, and we interpret the expression of OLIG2 as a sign of oligodendroglial lineage differentiation. e, f, Single-cell RNA-seq analysis of NPCs. e, NPCs have an expression program similar to the oligodendroglioma stemness program. Heat map shows the expression of genes (rows) most positively (top) or negatively (bottom) correlated with PC1 of a PCA of RNA-seq profiles for 431 single NPCs, across NPC cells (columns) rank ordered by their PC1 scores. Selected genes are indicated, and a full list of correlated genes for PC1 and PC2 is given in Supplementary Table 2. f, NPC cell scores for PC1 (y axis) and PC2 (x axis). PC2 correlated genes are associated with the cell cycle. Cells with the highest PC1 scores tend to be non-cycling (low PC2 score), indicating that while the stemness program is coupled to the cell cycle in oligodendroglioma, it is decoupled from the cell cycle in NPCs.
Extended Data Figure 5 Developmental hierarchy in oligodendroglioma.
a, Shown are plots as in Fig. 2d for each of the six tumours with cycling cells coloured as in Fig. 3. b, Lineage and stemness scores for three tumours with high-depth profiling, coloured based on sequencing batches, demonstrating the lack of considerable batch effects. c, For each of the three tumours profiled at high depth (horizontal panels) and for the two lineages (vertical panels), we calculated the significance of co-expression among sets of AC-related (top panels) or OC-related (bottom panels) genes within limited ranges of lineage scores (between the value of the x axis and that of the y axis). Significance was calculated by comparison of average co-expression to that of 100,000 control gene-sets with similar number of genes and distribution of average expression levels, and is indicated by colour. The significant co-expression patterns within limited ranges of lineage scores suggest that variability of lineage scores in these ranges cannot be driven by noise alone, and implies the existence of multiple states within each lineage, presumably reflecting intermediate differentiation states (see Supplementary Note 2). d, Characterization of tumour subpopulations by histopathology and tissue staining. Top/middle panels denote two predominant lineages of AC-like and OC-like cells. Shown are MGH53 with haematoxylin and eosin (H&E, top left), immunohistochemistry for OLIG2 (oligodendrocyte marker, top right) and GFAP (astrocyte marker, middle left), as well as in situ RNA hybridization for astrocytic markers ApoE (apolipoprotein E, astrocytic lineage, middle right), with patterns similar to GFAP immunohistochemistry. Bottom panels denote stem-like cells and association with cell cycle. In situ RNA hybridization for the stem/progenitor markers SOX4 and CCND2 (bottom left) and the proliferation marker Ki-67 (bottom right) in MGH36 identifies cells positive for both markers (arrows). Immunohistochemistry for GFAP (arrowhead, bottom right) and Ki-67 (arrow, bottom right) shows mutually exclusive expression patterns. e, Consistency of MGH60 hierarchy between the full-length SMART-Seq2 protocol used throughout this work (left panels), and an alternative protocol (right panels) in which only the 5′-ends of transcripts are analysed while incorporating random molecular tags (RMTs, also known us unique molecular identifiers, or UMIs) that decrease the biases of PCR amplification. Top panels: PC1 reflects an AC-like and OC-like distinction. Shown are heatmaps of the AC-like and OC-like specific genes (rows, as defined in Supplementary Table 1 and restricted to genes with average expression log2(TPM + 1) > 4 in each data set) with cells ordered by their PC1 score. Bottom, lineage (x axis) and stemness (y axis) scores (defined as in Fig. 2d).
Extended Data Figure 6 Cell-cycle analysis.
a, High expression of G1/S and G2/M gene sets in a subset of cycling cells. Shown are the average expression (top panels, lines) or the expression of all individual genes (bottom, heat maps) of the G1/S and G2/M gene sets, in all cells (n = 2,594) (left) or only among the putative cycling cells (n = 119) (right) from the three tumours profiled at high-depth ordered by cell cycle expression. Dashed lines (top right) separates the four inferred phases of cycling cells, corresponding to light blue, blue, green and red in Fig. 3a, respectively. b, Estimated fraction of cycling cells (y axis) in each of 3 tumours (x axis) based on single cell RNA-seq (left; different phases marked by colour code as in Fig. 3a) or Ki-67 immunohistochemistry (right). c, Variation in cycling cells between regions of the same tumour. Shown is Ki-67 immunohistochemistry in two regions in MGH36. Such regional variability in proliferation complicates direct comparisons as done in b. d, Cycling cells are enriched in stem-like and undifferentiated cells compared to differentiated cells. Shown is the percentage of cycling cells (y axis) in four bins based on stemness scores (top) or lineage scores (bottom). Black squares and error-bars correspond to the mean and standard deviation of the percentages in the three tumours profiled at high depth (MGH36, MGH53, MGH54), and red circles denote the percentages in individual tumours. Bins in left panel were defined as stemness scores below −1.5 (n = 711), between −1.5 and 0.5 (n = 1,100), between −0.5 and 0.5 (n = 939), and above 0.5 (n = 274), respectively. The first two bins are significantly depleted with cycling cells, while the last two bins are significantly enriched (P < 0.05, hypergeometric test). Bins in left panel were defined as AC score above 1 (n = 503), AC score between 0.5 and 1 (n = 1,013), AC and OC scores below 0.5 (n = 1,130), OC score between 0.5 and 1 (n = 855), and OC score above 1 (n = 597), respectively. The third bin is significantly enriched with cycling cells, while the four other bins are significantly depleted (P < 0.05, hypergeometric test). e, Correlation between the average expression of cell cycle (y axis) and that of stemness genes (x axis) across molecularly defined oligodendrogliomas (by IDH mutation, chromosome 1p and 19q co-deletion, and absence of P53 and ATRX mutations) profiled by TCGA (n = 69) with bulk RNA-seq. Average expression was defined by centring the log2-transformed RSEM gene quantifications. Also shown are the linear least-square regression and Pearson correlation coefficient. f, Specific enrichment of S/G2/M cells compared to G1 cells among stem-like or undifferentiated cells. Shown is the proportion (y axis) of each marked category of cells among the stem-like or undifferentiated subpopulations. Significant enrichments are marked (P < 0.01, hypergeometric test).
Extended Data Figure 7 CCND2 is associated with both cycling and non-cycling stem/progenitor cells.
a, CCND2, but not CCND1 or CCND3, is upregulated in non-cycling stem-like oligodendroglioma cells. Shown are the average expression levels (y axis, log-scale) of three cyclin D genes (x axis) in non-cycling cells classified as OC-like cells (light blue), undifferentiated cells (grey) and stem-like cells (purple). CCND2 is approximately fourfold higher in stem-like non-cycling cells than in OC-like and undifferentiated cells (P < 0.001 by permutation test). Conversely, CCND1 and CCND3 are expressed at comparable levels in stem-like and OC-like cells. b, Upregulation of cyclin D genes in cycling cells compared to non-cycling cells. As in a but for up regulation (log2-ratio) in cycling cells vs. non-cycling cells. CCND2 levels further increase in cycling undifferentiated and stem-like cells but not in OC-like cells, whereas CCND1 and CCND3 levels increase in OC-like cycling cells more than in undifferentiated and stem-like cycling cells. c, Distinct expression patterns of cyclin D genes in human brain development. Shown are the expression patterns of three cyclin D genes (rows) in human brain samples at different points in pre- and post-natal development, sorted by age (columns) from the Allen Brain Atlas15. CCND2 is associated with prenatal samples, whereas CCND1 and CCND3 are expressed mostly in childhood and adult samples. d, CCND2 is upregulated in activated versus quiescent NSCs19, both among cycling and non-cycling cells. Activated NSCs were partitioned into non-cycling cells (black) and cycling cells in the G1/S (green) or G2/M (red) phases (Methods). Expression difference (y axis) for each of three genes (x axis) was quantified for each of these subsets as the log2-ratio of the average expression in the respective subset versus the quiescent NSCs, and was significant for each of the three subsets (P < 0.05 by permutation test). Although CCND2 (left) is induced in both cycling and non-cycling activated NSCs, two canonical cell cycle genes (PCNA, middle; and AURKB, right) are not induced in non-cycling genes but were induced preferentially in G1/S and G2/M cells, respectively.
Extended Data Figure 8 Distribution of cellular states in distinct genetic clones of MGH36 and MGH97.
a, Stemness (y axis) and lineage (x axis) score plots for MGH36 (top) and MGH97 (bottom), each separated into clone 1 (left) and clone 2 (right) as determined by CNV analysis (Fig. 1a, b). Cycling cells are coloured as in Fig. 3, with G1/S cells in blue, S/G2 cells in green, and G2/M cells in red. b, Colour-coded density of cells across the cellular hierarchy as shown in Fig. 2e, for the two clones (left: clone 1, right: clone 2) in each of the two tumours (top: MGH36, bottom: MGH97). c, The fraction of cells assigned to the different tumour compartments (y axis, Methods) based on either single-cell RNA-seq (blue) or RNA in situ hybridization (orange). Circles denote individual tumours; squares denote average of all tumours; error bars denote standard deviation across tumours, showing general agreement between scRNA-Seq and IHC estimates.
Extended Data Figure 9 Subclonal mutations tend to span the cellular hierarchy.
Each panel shows lineage (x axis) and stemness (y axis) scores of cells in which we ascertained by single cell RNA-seq a mutant (red), a wild-type (blue) or none (black) of the alleles. Included are mutations for which at least three cells were identified as mutants and that were identified by WES as subclonal (fraction <60%). The gene names, tumour name, ABSOLUTE-derived fraction of mutant cells (E, expected fraction) and the fraction of cells detected as mutant by RNA-seq (O, observed) are also indicated within each panel. We note that identification of a wild-type allele (blue) does not imply a wild-type cell because mutations may be heterozygous, and thus cells could contain both alleles while only one may be detected by single-cell RNA-seq. The observed fraction of mutations (O) is much lower than expected (E) due to limited coverage of the single-cell RNA-seq data, as well as due to heterozygosity. The vast majority of mutations (20 of 22) are distributed across the hierarchy and span multiple compartments. Two remaining mutations (H2AFV and EIF2AK2) appear more restricted to the ‘undifferentiated’ region (intermediate lineage and stemness scores), which could reflect our limited detection rate of mutant cells and/or a bias of the mutation to a particular region. To test the significance of potential biases in the distribution of mutations we calculated, for each mutation, a Euclidean distance among all pairs of mutant cells (based on their lineage and stemness scores), and compared the average pairwise distances among mutant cells to that among randomly selected subsets of the same number of cells. None of the mutations were significant with a false discovery rate (FDR) of 0.1, although this could reflect our limited statistical power and we cannot exclude a potential bias. The apparent bias of mutant cells to the OC lineage over the AC lineage (that is, positive versus negative lineage scores) reflects the lower frequencies of AC-like cells compared to OC-like cells in MGH53 and MGH54 (MGH53: 17% AC versus 39% OC; MGH54: 23% AC vs. 45% OC); this bias is also observed for the detection of wild-type alleles (blue) demonstrating that there is no bias against mutation detection in the AC lineage.
Extended Data Figure 10 Loss-of-heterozygosity event in MGH54 reveals two clones that span the cellular hierarchy.
a, Chromosome 18 loss of heterozygosity (LOH) in MGH54. Allelic fraction analysis of MGH54 SNPs from WES shows an imbalance (red and blue dots) in the frequency of alternative alleles in chromosome 1p, 19q, as well as chromosome 18, despite the normal copy number at this chromosome (Fig. 1a). This is consistent with an LOH event in which presumably one copy of chromosome 18 was deleted, and the other copy amplified. The weaker imbalance compared to chromosomes 1p and 19q further suggests that this is a subclonal event. b, Each of two clones defined by chromosome 18 LOH status spans the full hierarchy. Shown are the lineage (x axis) and stemness (y axis) scores for each cell from MGH54 (n = 1,174) classified as pre-LOH (red), post-LOH (blue) and unresolved (black) based on RNA-seq reads that map to SNPs in the minor (that is, deleted) chromosome. Both the pre- and post-LOH clones span the different tumour subpopulations. Pre-LOH cells were defined as all cells with reads that map to minor alleles in chromosome 18; post-LOH cells were defined as all cells with reads that map to at least five different major alleles, but no reads that map to minor alleles in chromosome 18; all other cells were defined as unresolved.
Supplementary information
Supplementary Data
This file contains Supplementary Table 1. (XLSX 20 kb)
Supplementary Data
This file contains Supplementary Table 2. (XLSX 24 kb)
Supplementary Data
This file contains Supplementary Table 3. (XLSX 13 kb)
Supplementary Data
This file contains Supplementary Table 4. (XLSX 43 kb)
Supplementary Data
This file contains Supplementary Table 5. (XLSX 12 kb)
Supplementary Information
This file contains Supplementary Notes. (PDF 118 kb)
Rights and permissions
About this article
Cite this article
Tirosh, I., Venteicher, A., Hebert, C. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016). https://doi.org/10.1038/nature20123
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20123
This article is cited by
-
Development and validation of a disulfidptosis and disulfide metabolism-related risk index for predicting prognosis in lung adenocarcinoma
Cancer Cell International (2024)
-
EMP3 as a prognostic biomarker correlates with EMT in GBM
BMC Cancer (2024)
-
Deubiquitinase Mysm1 regulates neural stem cell proliferation and differentiation by controlling Id4 expression
Cell Death & Disease (2024)
-
Threonine fuels glioblastoma through YRDC-mediated codon-biased translational reprogramming
Nature Cancer (2024)
-
Deciphering the tumor immune microenvironment of imatinib-resistance in advanced gastrointestinal stromal tumors at single-cell resolution
Cell Death & Disease (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.