Abstract
It is widely accepted that complex interactions between cancer cells and their surrounding microenvironment contribute to disease development, chemo-resistance and disease relapse. In light of this observed interdependency, novel therapeutic interventions that target specific cancer stroma cell lineages and their interactions are being sought. Here we studied a mouse model of human T-cell acute lymphoblastic leukaemia (T-ALL) and used intravital microscopy to monitor the progression of disease within the bone marrow at both the tissue-wide and single-cell level over time, from bone marrow seeding to development/selection of chemo-resistance. We observed highly dynamic cellular interactions and promiscuous distribution of leukaemia cells that migrated across the bone marrow, without showing any preferential association with bone marrow sub-compartments. Unexpectedly, this behaviour was maintained throughout disease development, from the earliest bone marrow seeding to response and resistance to chemotherapy. Our results reveal that T-ALL cells do not depend on specific bone marrow microenvironments for propagation of disease, nor for the selection of chemo-resistant clones, suggesting that a stochastic mechanism underlies these processes. Yet, although T-ALL infiltration and progression are independent of the stroma, accumulated disease burden leads to rapid, selective remodelling of the endosteal space, resulting in a complete loss of mature osteoblastic cells while perivascular cells are maintained. This outcome leads to a shift in the balance of endogenous bone marrow stroma, towards a composition associated with less efficient haematopoietic stem cell function1. This novel, dynamic analysis of T-ALL interactions with the bone marrow microenvironment in vivo, supported by evidence from human T-ALL samples, highlights that future therapeutic interventions should target the migration and promiscuous interactions of cancer cells with the surrounding microenvironment, rather than specific bone marrow stroma, to combat the invasion by and survival of chemo-resistant T-ALL cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
Gene expression data have been deposited in ArrayExpress under accession number E-MTAB-4889.
References
Bowers, M. et al. Osteoblast ablation reduces normal long-term hematopoietic stem cell self-renewal but accelerates leukemia development. Blood 125, 2678–2688 (2015)
Colmone, A. et al. Leukemic cells create bone marrow niches that disrupt the behavior of normal hematopoietic progenitor cells. Science 322, 1861–1865 (2008)
Duan, C. W. et al. Leukemia propagating cells rebuild an evolving niche in response to therapy. Cancer Cell 25, 778–793 (2014)
Hanoun, M. et al. Acute myelogenous leukemia-induced sympathetic neuropathy promotes malignancy in an altered hematopoietic stem cell niche. Cell Stem Cell 15, 365–375 (2014)
Ishikawa, F. et al. Chemotherapy-resistant human AML stem cells home to and engraft within the bone-marrow endosteal region. Nature Biotechnol. 25, 1315–1321 (2007)
Saito, Y. et al. Induction of cell cycle entry eliminates human leukemia stem cells in a mouse model of AML. Nature Biotechnol. 28, 275–280 (2010)
Schepers, K. et al. Myeloproliferative neoplasia remodels the endosteal bone marrow niche into a self-reinforcing leukemic niche. Cell Stem Cell 13, 285–299 (2013)
Zhang, B. et al. Altered microenvironmental regulation of leukemic and normal stem cells in chronic myelogenous leukemia. Cancer Cell 21, 577–592 (2012)
Jin, L. et al. CXCR4 up-regulation by imatinib induces chronic myelogenous leukemia (CML) cell migration to bone marrow stroma and promotes survival of quiescent CML cells. Mol. Cancer Ther. 7, 48–58 (2008)
Weisberg, E. et al. Inhibition of CXCR4 in CML cells disrupts their interaction with the bone marrow microenvironment and sensitizes them to nilotinib. Leukemia 26, 985–990 (2012)
Sanda, T. et al. Interconnecting molecular pathways in the pathogenesis and drug sensitivity of T-cell acute lymphoblastic leukemia. Blood 115, 1735–1745 (2010)
Weng, A. P. et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306, 269–271 (2004)
Pui, C. H., Robison, L. L. & Look, A. T. Acute lymphoblastic leukaemia. Lancet 371, 1030–1043 (2008)
Lo Celso, C. et al. Live-animal tracking of individual haematopoietic stem/progenitor cells in their niche. Nature 457, 92–96 (2009)
Rashidi, N. M. et al. In vivo time-lapse imaging shows diverse niche engagement by quiescent and naturally activated hematopoietic stem cells. Blood 124, 79–83 (2014)
Scott, M. K., Akinduro, O. & Lo Celso, C. In vivo 4-dimensional tracking of hematopoietic stem and progenitor cells in adult mouse calvarial bone marrow. J. Vis. Exp. 91, e51683 (2014)
Kalajzic, I. et al. Use of type I collagen green fluorescent protein transgenes to identify subpopulations of cells at different stages of the osteoblast lineage. J. Bone Miner. Res. 17, 15–25 (2002)
Paic, F. et al. Identification of differentially expressed genes between osteoblasts and osteocytes. Bone 45, 682–692 (2009)
Méndez-Ferrer, S. et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 466, 829–834 (2010)
Inaba, H. & Pui, C. H. Glucocorticoid use in acute lymphoblastic leukaemia. Lancet Oncol. 11, 1096–1106 (2010)
Mullighan, C. G. et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 322, 1377–1380 (2008)
Lassailly, F., Foster, K., Lopez-Onieva, L., Currie, E. & Bonnet, D. Multimodal imaging reveals structural and functional heterogeneity in different bone marrow compartments: functional implications on hematopoietic stem cells. Blood 122, 1730–1740 (2013)
Visnjic, D. et al. Hematopoiesis is severely altered in mice with an induced osteoblast deficiency. Blood 103, 3258–3264 (2004)
Krevvata, M. et al. Inhibition of leukemia cell engraftment and disease progression in mice by osteoblasts. Blood 124, 2834–2846 (2014)
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007)
Rodda, S. J. & McMahon, A. P. Distinct roles for Hedgehog and canonical Wnt signaling in specification, differentiation and maintenance of osteoblast progenitors. Development 133, 3231–3244 (2006)
Hawkins, E. D. et al. Lethal giant larvae 1 tumour suppressor activity is not conserved in models of mammalian T and B cell leukaemia. PLoS ONE 9, e87376 (2014)
Aster, J. C. et al. Essential roles for ankyrin repeat and transactivation domains in induction of T-cell leukemia by notch1. Mol. Cell. Biol. 20, 7505–7515 (2000)
Real, P. J. et al. Gamma-secretase inhibitors reverse glucocorticoid resistance in T cell acute lymphoblastic leukemia. Nature Med. 15, 50–58 (2009)
Chiu, P. P., Jiang, H. & Dick, J. E. Leukemia-initiating cells in human T-lymphoblastic leukemia exhibit glucocorticoid resistance. Blood 116, 5268–5279 (2010)
Lo Celso, C., Lin, C. P. & Scadden, D. T. In vivo imaging of transplanted hematopoietic stem and progenitor cells in mouse calvarium bone marrow. Nature Protocols 6, 1–14 (2011)
Khorshed, R. A. et al. Automated identification and localization of hematopoietic stem cells in 3D intravital microscopy data. Stem Cell Reports 5, 139–153 (2015)
Preibisch, S., Saalfeld, S., Schindelin, J. & Tomancak, P. Software for bead-based registration of selective plane illumination microscopy data. Nature Methods 7, 418–419 (2010)
Carvalho, B. S. & Irizarry, R. A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 26, 2363–2367 (2010)
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015)
Hawkins, E. D. et al. Measuring lymphocyte proliferation, survival and differentiation using CFSE time-series data. Nature Protocols 2, 2057–2067 (2007)
Acknowledgements
E.D.H. was supported by a fellowship from the European Haematology Association and an RD Wright Career fellowship from the National Health and Medical Research Council of Australia (NHMRC) as well as a project grant from Bloodwise (12033). D.D. was supported by the GABBA PhD program (FCT fellowship SFRH/BD/52195/2013). L.E.P. was supported by a National Health and Medical Research Council of Australia Senior Research Fellowship. H.Q. is supported by Victorian Cancer Agency Early Career Seed Grant. D.B., D.P. and K.F. were funded by Cancer Research UK (CRUK) and the Francis Crick Institute. C.L.C. was funded by Bloodwise (12033), Human Frontier Science Program (RGP0051/2011), CRUK (C36195/A1183), Biotechnology and Biological Sciences Research Council (BB/I004033/1) Kay Kendall Leukaemia Fund (KKL460) and the European Research Council (337066). We are grateful to M. Spitaler, D. Keller and L. Carlin for support with imaging. We also thank S. Piperelis, E. Ibarguen, W. Steel and H. Goyal for logistical help; A. Ivan for support from the genomics facility; J. Srivastava and C. Simpson for support from the flow cytometry facility; M. Wall for her advice on the human cytogenetics; D. Ibanez for access to software; H. Fleming, V. Greco, R. E. Sinden, S. Mostowy and P. O’Donovan for critical feedback on the manuscript.
Author information
Authors and Affiliations
Contributions
E.D.H. and C.L.C. conceived the project. E.D.H., C.L.C., O.A. and M.K.S. designed and refined the intravital imaging systems. R.A.K. designed the image analysis platform; R.A.K., E.D.H. and D.D. performed all image analysis. S.R. designed the random data simulator. E.D.H., D.D. and N.R. performed imaging experiments. N.R. maintained all animal lines. M.N. and M.D.R. designed the microarray experiment and analysed the data. E.D.H., M.W. and R.W.J. performed experiments in Extended Data Fig. 1. D.P., K.F. and D.B. designed, performed and analysed osterix–GFP experiments. S.J.H., D.A.W. and J.G. provided human T-ALL samples. L.S. and L.E.P. performed histological analysis of human BM trephines. H.Q. funded human biopsy studies. D.P. and D.B. designed and D.P. performed the human T-ALL xenograft experiments and analysed data. E.D.H., D.D. and C.L.C. analysed data and wrote the manuscript. All authors contributed revisions to the manuscript. Note that O.A., R.A.K. and D.P. contributed equally to this work, and the same is for M.D.R., L.E.P. and D.B.
Corresponding authors
Additional information
Reviewer Information Nature thanks T. Look, C. Mullighan, J. van Rheenen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 T-ALL disease experimental model.
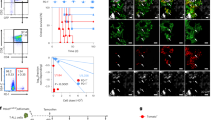
Fetal liver single-cell suspensions were isolated from embryonic day (E)14.5 wild-type (WT) embryos and transduced with DsRed alone or DsRed with NotchICNΔRamΔP then transplanted into primary lethally irradiated recipient mice. Recipient mice typically accumulated CD4+CD8+ cells in the peripheral blood from 4 weeks after transplant. Transformed leukaemic cells could be distinguished from non-malignant cells based on DsRed expression levels, where DsREDlo cells were transduced with the Notch construct yet were non-malignant as they had not yet acquired secondary mutations to drive leukaemogenesis, whereas DsRedhi cells were fully malignant. DsRedlo cells contained single-positive CD4+ and CD8+ T-cell populations, whereas DsRedhi cells had predominantly the leukaemic CD4+CD8+ phenotype. The accumulation of transformed leukaemic populations displayed large variation over time as shown in the ratio of transformed to non-malignant cells in peripheral blood of primary recipient mice at 6 and 9 weeks. Data shown are mean ± s.e.m. When DsRedhi cells dominated peripheral cell populations, mice were burdened with typical CD4+CD8+ T-ALL (now simply referred to as DsRed+). When primary recipient mice displayed enlarged lymph nodes and/or spleen, they were euthanized and DsRed+ cells were harvested, stored frozen and transplanted into secondary recipients. CXCR4 expression was measured in CD4+ T cells, CD8+ T cells and T-ALL by flow cytometry (T-ALL from four primary donors) and microarray gene expression analysis (triplicate biological replicates are shown for control T cells, and samples from nine individual secondary recipients injected with five independent primary T-ALL samples). To track disease progression, 10,000 primary T-ALL cells were transplanted into cohorts of sub-lethally irradiated secondary recipient mice. Secondary transplanted cells colonized the BM primarily, before spreading to peripheral organs and blood (n = 4 mice per time point; data shown are mean ± s.e.m.) and developed disease more rapidly and synchronously than primary recipients as injected cells were already transformed. Secondary recipients survived for up to 38 days (shown are survival data of mice injected with four independent primary T-ALL samples, n = 2 mice per primary sample represented by the following symbols: filled circle, open square, open triangle and asterisk). In selected cases, secondary T-ALL blasts were transplanted into tertiary recipients, which developed disease more rapidly but showed similar responses to chemotherapy. The primary samples used in this study were from primary recipients 151, 907, B2M2, B2M3, B2M10, B3M3 and B3M30. This nomenclature refers to the mouse numbering from three independent fetal liver transductions. First transduction: mouse 151 and 907; second transduction (B2): mouse 2, 3 and 10; third transduction (B3): mouse 3 and mouse 30.
Extended Data Figure 2 Four-dimensional multi-position imaging of leukaemia cells in the BM space.
a, Representative maximum projection tile scan of a Col2.3–GFP recipient mouse (from Fig. 2) 12 days after transplantation of DsRed+ T-ALL. Red: T-ALL cells; green: GFP+ osteoblastic cells; blue: blood vessels; grey: bone collagen. b, Individual positions (framed in a) were selected and imaged at 3 min intervals for 3 h to measure cell migration and division. Shown here are a single time frame/position. For full time-lapse data see Supplementary Videos 2. c, Three-dimensional rendering of the three-dimensional tracks of individual leukaemia cells, measured using semi-automated tracking (red), overlaid to vasculature (blue). Spheres represent the beginning of each track. d, Data from c are shown with osteoblasts included in green. e, Tracks from c colour-coded based on time. Long stretches of the same colour correspond to faster movement, while rapid colour shifts represent slower movement. Data are representative of >30 time-lapse videos collected from eight secondary recipients (biological replicates) injected with three independent primary T-ALL samples.
Extended Data Figure 3 T-ALL expansion is not associated with Col2.3–GFP+ osteoblastic cells and nestin–GFP+ cells.
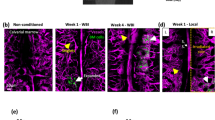
a, Representative tile scan of a Col2.3–GFP mouse 15 days after transplantation of T-ALL. Zooms P1–P4 illustrate that the expansion of disease is not associated with the presence or absence of GFP+ osteoblaststic cells. Red: T-ALL; green: osteoblastic cells; blue: blood vessels. Image is representative of five mice injected with two individual T-ALL primary samples. b, Representative tile scan of a nestin–GFP mouse 15 days after transplantation of T-ALL. Zooms P1–P3 illustrate that the expansion of disease is not associated with the presence or absence of nestin–GFP+ cells. Red: T-ALL; green: nestin+ cells; blue: blood vessels; grey: bone collagen second harmonic generation (SHG) signal. Image is representative of three individual mice (biological replicates).
Extended Data Figure 4 T-ALL expansion is not associated with BM areas containing nestin–GFP+, Col2.3–GFP+ cells or any combination of them.
Representative tile scan of a Col2.3–CFP/nestin–GFP double-transgenic mouse 15 days after transplantation of T-ALL. Zooms P1–P3 illustrate that the expansion of disease is not associated with the presence or absence of any combination of Col2.3–CFP+ or nestin–GFP+ cells. Yellow: T-ALL; green: nestin–GFP+ cells; blue: Col2.3–CFP+ cells; magenta: Cy5-labelled blood vessels; grey: bone collagen SHG signal. Image is representative of four mice (biological replicates).
Extended Data Figure 5 Dexamethasone-resistant T-ALL cells do not associate with nestin–GFP+ cells.
a, Representative nestin–GFP mouse transplanted with T-ALL cells and imaged 18 days post-transplant to confirm complete BM infiltration (left). Tile-scan imaging was repeated after 2 days of treatment with 15 mg kg−1 dexamethasone i.v. (right). b, Magnified view of representative positions, framed in a. c, We observed no preferential positioning of T-ALL surviving cells relative to nestin+ cells compared to simulated data. Red: T-ALL; green: nestin–GFP+ cells; blue: blood vessels. Data are representative of four individual mice injected with two T-ALL primary samples. Error bars: mean ± s.d. d, Chimaeric mice were generated by transplanting mTmG–tomato+ BM into Col2.3–GFP recipients. The high reconstitution efficiency of mTmG cells provided a more robust traceable marker of steady-state haematopoiesis for intravital imaging than MigR1–DsRed-transduced fetal liver cells (<40% reconstitution, data not shown). e, Representative tile scans of a mTmG/Col2.3–GFP chimaeric mouse performed before and after three doses of dexamethasone treatment showing that healthy BM is not affected by dexamethasone treatment or sub-lethal irradiation. Red: tomato+, healthy mTmG BM; green: GFP+ osteoblastic cells; blue: blood vessels. n = 2 mice. NS, not significant.
Extended Data Figure 6 Multi-day time course of response to chemotherapy.
a, Representative maximum projections of tile scans of calvarium BM of one Col2.3–CFP and one nestin–GFP mouse at 18 days after T-ALL transplant (pre-treatment) and 1 and 2 days of dexamethasone treatment (15 mg kg−1). Red squares indicate some areas of cell loss from day 1 to day 2. b, Three-dimensional measurement of the position of surviving cells in mice imaged at both 1 and 2 days of dexamethasone treatment. n = 1,499 and 352 T-ALL cells measured to osteoblastic CFP+ cells at day 1 and day 2, respectively, and 363 and 496 T-ALL cells measured to nestin–GFP+ cells at day 1 and day 2, respectively. Data are representative from three individual mice (biological replicates) of each genotype. Error bars: mean ± s.d. NS, not significant.
Extended Data Figure 7 Development of resistance to dexamethasone and gene-expression-based clustering of leukaemia samples collected before and after dexamethasone treatment.
a, Representative maximum projection of tile scan of calvarium bone marrow of a mouse after 7 days of daily dexamethasone treatment (15 mg kg−1, i.v.). Red, DsRed+ T-ALL cells. Data are representative of four independent mice injected with four independent T-ALL primary samples. b, Mice with T-ALL were either kept untreated or treated with dexamethasone for 7 days, at which point they were culled and the expression of CXCR4 on T-ALL cells analysed by flow cytometry. There was no statistically significant difference in the mean fluorescence intensity (MFI) of CXCR4 between the two groups. Data are representative of six untreated and five treated mice, injected with three independent T-ALL primary samples. Error bars: mean ± s.d. c, Multi-dimensional scaling (MDS) plot of control CD4+ T cells, CD8+ T cells, CD4+8+ thymocytes, whole BM and T-ALL samples with no treatment (grey) or after treatment with dexamethasone (red) based on microarray transcriptomics data for the 1,000 most variable genes. The name of the primary T-ALL sample used to inject each mouse is indicated next to the dot marking its position relative to all other samples. This nomenclature refers to the mouse numbering from three independent fetal liver transductions. First transduction: mouse 907; second transduction (B2): mouse 2, B2M2, mouse 3, B2M3 and 10, B2M10; third transduction (B3): mouse 3, B3M3 and mouse 30, B3M30. Control samples are purified by flow cytometry from three biological replicate mice and each circle represents an individual sample. NS, not significant.
Extended Data Figure 8 Analysis of human T-ALL cells during response to chemotherapy in NSG xenotransplant recipients.
a, b, Human T-ALL samples were transplanted into NOD/SCID/γ (NSG) mice and 12 days post-transplant, daily dexamethasone treatment at 15 mg kg−1 was initiated. Fourteen days later, the response was measured by flow cytometry. c, For intravital imaging, human T-ALL cells were labelled by injection of 10 μg anti-human CD45-PE 15–30 min before imaging. d, Cells were imaged at 3 min intervals for >60 min and migration was measured by manual tracking either before or after dexamethasone treatment. Pre-dexamethasone: n = 82 cells from 2 independent mice, 14 days; dexamethasone: n = 100 from 3 independent mice. Shown are cells from patient JH, wild-type NOTCH. e, BM sections were prepared from untreated and treated NSG mice and stained for human CD45 (red) and Ki-67 (green). In addition, nuclei were visualized using DAPI (blue) and bone by SHG signal (grey). Zooms are of the areas framed by the white boxes on their left. f, Analysis of 2,338 (untreated) and 1,576 (14 days dexamethasone) human CD45+ cells in sections from three mice per condition reveals no change in the fraction of proliferating Ki-67+ cells after dexamethasone treatment. NS, not significant. **P < 0.01. Error bars: mean ± s.d. (b, d, f).
Extended Data Figure 9 Combined dexamethasone, vincristine and l-asparaginase treatment effectively reduces T-ALL burden.
a, Representative tile scan of a mouse calvarium fully infiltrated with T-ALL (pre-DVA) and after 2 days of combination therapy (dexamethasone, vincristine and l-asparaginase (DVA)). b, Zooms P1 and P2 illustrate effectiveness of DVA treatment and the small number of surviving T-ALL cells. Red: T-ALL; blue: blood vessels; grey: SHG bone collagen. Image is representative of four mice (biological replicates) injected with one individual T-ALL secondary sample.
Extended Data Figure 10 Analysis of the response of bone marrow structures to irradiation and dexamethasone treatment and of nestin–GFP+ cells to T-ALL.
a, b, Col2.3–GFP (a) or nestin–GFP (b) mice were treated with combinations of sublethal irradiation (administered >18 days before measurement) or dexamethasone treatment (administered for 2 days before measurement) as indicated. Then, using three-dimensional image analysis of tile scans, the total volume of GFP+ cells was quantified. Groups were analysed using analysis of variance (ANOVA) with Bonferroni correction for multiple groups. Error bars: mean ± s.d. c, Representative tile scan of nestin–GFP mouse transplanted with T-ALL 21 days earlier. At infiltration levels that eradicated osteoblasts, we still observed healthy nestin–GFP+ cells. d, Higher magnification of area P1 framed in A, with the signal from each channel split for clarity. e, Three-dimensional render at higher magnification of area P2 framed in A, showing healthy blood flow within the highly infiltrated BM space. Red: T-ALL; green: nestin–GFP+ cells; blue: blood vessels; grey: bone collagen SHG signal. n = 5 independent mice injected with two independent T-ALL primary samples. NS, not significant.
Supplementary information
Supplementary Table 1
This table shows differentially expressed genes in T-ALL cells harvested before and after dexamethasone treatment. 79 genes were statistically significantly differentially expressed before and after treatment. Provided are the Probe ID, gene title and GeneID, p values and adjusted p values. (XLSX 17 kb)
Supplementary Table 2
This table contains relevant T-ALL patient information for trephine histology and xenotransplantation experiments. * denotes the patient sample represented in Figure 4g; # the patient sample represented in Extended data figure 8. M: male; F: female; ND: not determined; Dic: dicentric; Del: deletion; Ins: insertion; rearr: rearrangement. (XLSX 35 kb)
Long-term single cell T-ALL time-lapse microscopy
Maximum projection of 14-hour time-lapse data collected at three-minute intervals (shown at 15 frames per second). Arrows in paused frames indicate a cell that is about to undergo mitosis. To maintain blood vessel signal, Cy5 dextran was injected I.V. at regular intervals throughout the imaging period. Red: T-ALL cells; green: GFP+ osteoblastic cells; blue: blood vessels. (MOV 9648 kb)
Heterogeneous behaviour of T-ALL cells in Col2.3-GFP reporter mice
Maximum projection time-lapse data (shown at 15 frames per second) of 5 areas from the same mouse presented in Figure 2b (positions 1-3) and extended data 2 (positions 1-5). Each position was filmed for three hours at three-minute intervals. Red: T-ALL cells; green: GFP+ osteoblastic cells; blue: blood vessels; grey: bone collagen. Arrows in paused frames point at cells that are about to undergo mitosis. (MOV 4679 kb)
Heterogeneous behaviour of T-ALL cells in Nestin-GFP reporter mice
Maximum projection time-lapse data (shown at 15 frames per second) of 2 areas from the same mouse representative of T-ALL cells undergoing mitosis both far away from and close to Nestin-GFP+ cells. Each position was filmed for three hours at three-minute intervals. Red: T-ALL cells; green: Nestin-GFP+ cells; blue: blood vessels. Arrows in paused frames point at cells that are about to undergo mitosis. (MOV 1883 kb)
Cell migration in T-ALL disease burdened mice
Maximum projection three-hour time-lapse data collected at three-minute intervals (shown at 15 frames per second) from the tilescan presented in Extended Data Figure 3a. Red: T-ALL cells; green: GFP+ osteoblastic cells; blue: blood vessels. (MOV 907 kb)
T-ALL cell migration and mitosis in dexamethasone treated Col2.3-GFP+ mice
Maximum projection of three-hour time-lapse data collected at three-minute intervals (shown at 15 frames per second) from mice after 2 days of treatment with 15mg/kg dexamethasone as presented in figure 3h. Red: T-ALL cells; green: GFP+ osteoblastic cells; blue: blood vessels. Arrows in paused frames point at cells about to undergo mitosis. (MOV 1034 kb)
T-ALL cell migration, mitosis and death in dexamethasone treated Nestin-GFP+ mice
Maximum projection of time-lapse data collected at three-minute intervals (shown at 15 frames per second) from Nestin-GFP+ mice after 2 days of treatment with 15mg/kg dexamethasone as presented in Extended Data figure 5a. Red: T-ALL cells; green: Nestin-GFP+ cells; blue: blood vessels. In the first part, arrows in paused frames point at cells about to undergo mitosis. In the second part, arrows in paused frames point at cells about to undergo cell death. (MOV 1793 kb)
Human T-ALL cell migration in dexamethasone treated NOD/SCID/γ mice
Maximum projection of time-lapse data collected at three-minute intervals (shown at 9 frames per second) from NOD/SCID/γ mice before, and after 14 days of treatment with 15mg/kg dexamethasone treatment. Red: human T-ALL cells labelled by I.V. injection of anti-CD45 PE antibody; grey = bone collagen SHG. (MOV 901 kb)
T-ALL cell migration and mitosis in vincristine treated nestin-GFP+ mice
Maximum projection of three-hour time-lapse data collected at three-minute intervals (shown at 15 frames per second) from mice after 2 days of treatment with 0.15mg/kg vincristine as presented in figure 3j. Red: T-ALL cells; green: nestin-GFP+ cells; blue: blood vessels. Arrow in paused frame point at a cell about to undergo mitosis. Bone and blood vessel signals of the first frame were maintained throughout the video for clarity purposes. (MOV 872 kb)
T-ALL cell migration and mitosis in DVA treated mice
Maximum projection of three-hour time-lapse data collected at three-minute intervals (shown at 15 frames per second) from mice after 2 days of treatment with DVA combination therapy (dexamethasone 15 mg/kg, vincristine 0.15mg/kg, L-asparaginase 1000 IU/kg) as presented in figure 3i. Red: T-ALL cells; blue: blood vessels. Arrow in paused frame point at a cell about to undergo mitosis. Bone and blood vessel signals of the first frame were maintained throughout the video for clarity purposes. (MOV 127 kb)
T-ALL remodels the endosteal niche
Maximum projection of nine-hour time-lapse data collected at three minute intervals (shown at 15 frames per second) from data presented in Figure 4a (top panel) of >8 weeks post reconstitution with whole mTmG bone marrow 19 days after sublethal irradiation (first part) and in Figure 4a (lower panel) of a Col2.3-GFP mouse 19 days post transplantation with T-ALL (second part) or. Red: mTmG tomato+ haematopoietic cells (first part) / T-ALL cells (second part); green: GFP+ osteoblastic cells. Dotted lines highlight examples of two osteoblastic cells that change position during the course of the time lapse. Other examples are visible in the video. (MOV 5181 kb)
Rights and permissions
About this article
Cite this article
Hawkins, E., Duarte, D., Akinduro, O. et al. T-cell acute leukaemia exhibits dynamic interactions with bone marrow microenvironments. Nature 538, 518–522 (2016). https://doi.org/10.1038/nature19801
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19801
This article is cited by
-
Intravital imaging to study cancer progression and metastasis
Nature Reviews Cancer (2023)
-
Quantification of bone marrow interstitial pH and calcium concentration by intravital ratiometric imaging
Nature Communications (2022)
-
Multiphoton intravital microscopy of rodents
Nature Reviews Methods Primers (2022)
-
Mammary tumour cells remodel the bone marrow vascular microenvironment to support metastasis
Nature Communications (2021)
-
Adipocytes disrupt the translational programme of acute lymphoblastic leukaemia to favour tumour survival and persistence
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.