Abstract
The physiology of N-methyl-d-aspartate (NMDA) receptors is fundamental to brain development and function. NMDA receptors are ionotropic glutamate receptors that function as heterotetramers composed mainly of GluN1 and GluN2 subunits. Activation of NMDA receptors requires binding of neurotransmitter agonists to a ligand-binding domain (LBD) and structural rearrangement of an amino-terminal domain (ATD). Recent crystal structures of GluN1–GluN2B NMDA receptors bound to agonists and an allosteric inhibitor, ifenprodil, represent the allosterically inhibited state. However, how the ATD and LBD move to activate the NMDA receptor ion channel remains unclear. Here we applied X-ray crystallography, single-particle electron cryomicroscopy and electrophysiology to rat NMDA receptors to show that, in the absence of ifenprodil, the bi-lobed structure of GluN2 ATD adopts an open conformation accompanied by rearrangement of the GluN1–GluN2 ATD heterodimeric interface, altering subunit orientation in the ATD and LBD and forming an active receptor conformation that gates the ion channel.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
Atomic coordinates and structure factor for the apo-GluN1b–GluN2B ATD is deposited in the Protein Data Bank under the accession code 5B3J; the cryo-EM coordinates are deposited under the accession codes 5FXG, 5FXH, 5FXI, 5FXJ and 5FXK. The cryo-EM maps are deposited in EMDB under accession codes EMD-3352, EMD-3353, EMD-3354, EMD-3355 and EMD-3356.
References
Traynelis, S. F. et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol. Rev. 62, 405–496 (2010)
Benveniste, M. & Mayer, M. L. Structure-activity analysis of binding kinetics for NMDA receptor competitive antagonists: the influence of conformational restriction. Br. J. Pharmacol. 104, 207–221 (1991)
Clements, J. D. & Westbrook, G. L. Activation kinetics reveal the number of glutamate and glycine binding sites on the N-methyl-d-aspartate receptor. Neuron 7, 605–613 (1991)
Johnson, J. W. & Ascher, P. Glycine potentiates the NMDA response in cultured mouse brain neurons. Nature 325, 529–531 (1987)
Nowak, L., Bregestovski, P., Ascher, P., Herbet, A. & Prochiantz, A. Magnesium gates glutamate-activated channels in mouse central neurones. Nature 307, 462–465 (1984)
Mayer, M. L., Westbrook, G. L. & Guthrie, P. B. Voltage-dependent block by Mg2+ of NMDA responses in spinal cord neurones. Nature 309, 261–263 (1984)
Mayer, M. L., MacDermott, A. B., Westbrook, G. L., Smith, S. J. & Barker, J. L. Agonist- and voltage-gated calcium entry in cultured mouse spinal cord neurons under voltage clamp measured using arsenazo III. J. Neurosci. 7, 3230–3244 (1987)
Granger, A. J. & Nicoll, R. A. Expression mechanisms underlying long-term potentiation: a postsynaptic view, 10 years on. Phil. Trans. R. Soc. Lond. B 369, 20130136 (2014)
Paoletti, P., Bellone, C. & Zhou, Q. NMDA receptor subunit diversity: impact on receptor properties, synaptic plasticity and disease. Nature Rev. Neurosci. 14, 383–400 (2013)
Armstrong, N., Sun, Y., Chen, G. Q. & Gouaux, E. Structure of a glutamate-receptor ligand-binding core in complex with kainate. Nature 395, 913–917 (1998)
Furukawa, H., Singh, S. K., Mancusso, R. & Gouaux, E. Subunit arrangement and function in NMDA receptors. Nature 438, 185–192 (2005)
Mayer, M. L. Crystal structures of the GluR5 and GluR6 ligand binding cores: molecular mechanisms underlying kainate receptor selectivity. Neuron 45, 539–552 (2005)
Hansen, K. B., Furukawa, H. & Traynelis, S. F. Control of assembly and function of glutamate receptors by the amino-terminal domain. Mol. Pharmacol. 78, 535–549 (2010)
Gielen, M., Siegler Retchless, B., Mony, L., Johnson, J. W. & Paoletti, P. Mechanism of differential control of NMDA receptor activity by NR2 subunits. Nature 459, 703–707 (2009)
Yuan, H., Hansen, K. B., Vance, K. M., Ogden, K. K. & Traynelis, S. F. Control of NMDA receptor function by the NR2 subunit amino-terminal domain. J. Neurosci. 29, 12045–12058 (2009)
Zhu, S. & Paoletti, P. Allosteric modulators of NMDA receptors: multiple sites and mechanisms. Curr. Opin. Pharmacol. 20, 14–23 (2015)
Meyerson, J. R. et al. Structural mechanism of glutamate receptor activation and desensitization. Nature 514, 328–334 (2014)
Karakas, E., Regan, M. C. & Furukawa, H. Emerging structural insights into the function of ionotropic glutamate receptors. Trends Biochem. Sci. 40, 328–337 (2015)
Regan, M. C., Romero-Hernandez, A. & Furukawa, H. A structural biology perspective on NMDA receptor pharmacology and function. Curr. Opin. Struct. Biol. 33, 68–75 (2015)
Sobolevsky, A. I., Rosconi, M. P. & Gouaux, E. X-ray structure, symmetry and mechanism of an AMPA-subtype glutamate receptor. Nature 462, 745–756 (2009)
Kumar, J., Schuck, P. & Mayer, M. L. Structure and assembly mechanism for heteromeric kainate receptors. Neuron 71, 319–331 (2011)
Karakas, E. & Furukawa, H. Crystal structure of a heterotetrameric NMDA receptor ion channel. Science 344, 992–997 (2014)
Lee, C. H. et al. NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 511, 191–197 (2014)
Karakas, E., Simorowski, N. & Furukawa, H. Subunit arrangement and phenylethanolamine binding in GluN1/GluN2B NMDA receptors. Nature 475, 249–253 (2011)
Karakas, E., Simorowski, N. & Furukawa, H. Structure of the zinc-bound amino-terminal domain of the NMDA receptor NR2B subunit. EMBO J. 28, 3910–3920 (2009)
Banke, T. G. & Traynelis, S. F. Activation of NR1/NR2B NMDA receptors. Nature Neurosci. 6, 144–152 (2003)
Farina, A. N. et al. Separation of domain contacts is required for heterotetrameric assembly of functional NMDA receptors. J. Neurosci. 31, 3565–3579 (2011)
Jespersen, A., Tajima, N., Fernandez-Cuervo, G., Garnier-Amblard, E. C. & Furukawa, H. Structural insights into competitive antagonism in NMDA receptors. Neuron 81, 366–378 (2014)
Vance, K. M., Simorowski, N., Traynelis, S. F. & Furukawa, H. Ligand-specific deactivation time course of GluN1/GluN2D NMDA receptors. Nature Commun. 2, 294 (2011)
Sirrieh, R. E., MacLean, D. M. & Jayaraman, V. A. Conserved structural mechanism of NMDA receptor inhibition: A comparison of ifenprodil and zinc. J. Gen. Physiol. 146, 173–181 (2015)
Dürr, K. L. et al. Structure and Dynamics of AMPA Receptor GluA2 in Resting, Pre-Open, and Desensitized States. Cell 158, 778–792 (2014)
Kazi, R., Dai, J., Sweeney, C., Zhou, H. X. & Wollmuth, L. P. Mechanical coupling maintains the fidelity of NMDA receptor-mediated currents. Nature Neurosci. 17, 914–922 (2014)
Popescu, G. & Auerbach, A. The NMDA receptor gating machine: lessons from single channels. Neuroscientist 10, 192–198 (2004)
Popescu, G., Robert, A., Howe, J. R. & Auerbach, A. Reaction mechanism determines NMDA receptor response to repetitive stimulation. Nature 430, 790–793 (2004)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Grant, T. & Grigorieff, N. Automatic estimation and correction of anisotropic magnification distortion in electron microscopes. J. Struct. Biol. 192, 204–208 (2015)
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 A reconstruction of rotavirus VP6. eLife 4, e06980 (2015)
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015)
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Afonine, P. V., Headd, J. J., Terwilliger, T. C. & Adams, P. D. New tool: phenix.real_space_refine. Computational Crystallography Newsletter 4, 43–44 (2013)
Acknowledgements
We thank staff at the 23-ID beamlines at the Advanced Photon System in the Argonne National Laboratory. We are grateful to Z. Yu, C. Hong and R. Huang at the Janelia Research Center/HHMI EM facility for their support. This work was supported by the National Institutes of Health (MH085926 and GM105730), the Stanley Institute of Cognitive Genomics, Burroughs Wellcome Fund Collaborative Research Travel Grant, the Robertson Research Fund of Cold Spring Harbor Laboratory (all to H.F.), Japan Society for the Promotion of Science (to N.T.) and the Visiting Scientist program of the Janelia Research Center to allow H.F. to conduct cryo-EM work.
Author information
Authors and Affiliations
Contributions
The authors jointly contributed to project design. N.T. and H.F. performed X-ray crystallography and electrophysiology. N.S. purified and characterized antibodies critical for the x-ray crystallographic study. T.G., R.D.A., N.G. and H.F. were involved in structural analysis by cryo-EM. E.K. expressed and purified proteins for the cryo-EM analysis and conducted model building and refinement of the cryo-EM structures. N.T., E.K., T.G., N.G. and H.F. were involved in manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Domain organization and constructs.
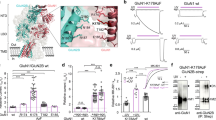
a, The construct design for GluN1b and GluN2B ATD used in this study. GluN1b from Xenopus laevis is combined with GluN2B from rat, as in the previous study on the ATD24. b, The construct design for the intact GluN1b/GluN2B NMDA receptors from rat. A similar construct was used in previous studies and shown to be fully functional22.
Extended Data Figure 2 Structure of the apo-GluN1b–GluN2B ATD.
a, Representative 2Fo − Fc electron density map contoured at 1.2σ showing continuous density throughout GluN1b, GluN2B and Fab. The quality of the electron density map is at a sufficient level to model amino acid side chains (see lower panel). b, Crystal packing of GluN1b–GluN2B ATD–Fab showing that the packing is mediated robustly by Fab molecules (green). The colour coding for the ATD is the same as in Fig. 1. c, Comparison of the apo-GluN1b–GluN2B ATD and ifenprodil–GluN1b–GluN2B ATD (grey) by stereo presentation. Colour coding for the apo-GluN1b–GluN2B ATD is the same as in Fig. 1. Here the two structures are superimposed at GluN2B R1.
Extended Data Figure 3 Validation of the crystal structure by disulfide cross-linking.
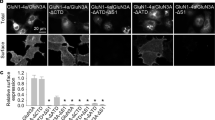
a, Crystal structure of the apo-GluN1b–GluN2B ATD showing locations of the mutated residues, GluN1b Phe113, GluN1b Gly331, GluN2B Ala107 and GluN2B Glu75 in spheres. b, Western blots using anti-GluN1 (left) and anti-GluN2B (right) antibodies on purified intact GluN1b/GluN2B NMDA receptor that lacks the CTD. Upper and lower panels are blots run in the absence and presence of β-mercaptoethanol (βME), respectively. Bands highlighted by arrow 1 are consistent with the molecular weight of GluN1–GluN2B heterodimers, whereas those highlighted by arrows 2 and 3 are consistent with the molecular weights of monomers of GluN1–4b and GluN2B.
Extended Data Figure 4 Conformational trap shows the apo-GluN1b–GluN2B ATD structure to represent ‘active’ form-II.
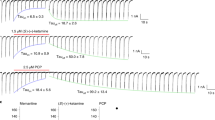
a, Location of engineered cysteines in the crystal structure of the apo-GluN1b–GluN2B ATD. The cysteine mutant pairs, GluN1–4b(Ala175Cys)/GluN2B(Gln180Cys) (green spheres) and GluN1–4b(Lys178Cys)/GluN2B(Asn184Cys) (blue spheres) are co-expressed in Xenopus oocytes and cross-linked by bifunctional MTS with different linker lengths (M2M, M4M and M8M). b, Application of 200 μM M4M in the presence or absence of 100 μM agonists (glycine (gly)/glutamate (glut)) potentiates the macroscopic current measured at the holding potential of −60 mV by two-electrode voltage clamp. No potentiation was observed when M4M was applied in the presence of ifenprodil (ifen). Shown here are the representative recording profiles for the GluN1–4b(Lys178Cys)/GluN2B(Asn184Cys) pair.
Extended Data Figure 5 Effect of bi-MTAs on cysteine mutants.
a, b, M4M specifically traps the active conformation at the engineered cysteines. Representative electrophysiological traces for the mutant pairs, GluN1–4b(Ala175Cys)/GluN2B(Gln180Cys) (green spheres) and GluN1–4b(Lys178Cys)/GluN2B(Asn184Cys) as well as mutant and wild-type pairs. The experiments were conducted by two-electrode voltage clamp as in Fig. 2. The potentiation by M4M (represented by IMTS/Io) is only observed when both GluN1 and GluN2B cysteine mutants are co-expressed. No potentiation was observed when the cysteine mutant of one subunit is combined with the wild type of the other, indicating that the effect of M4M modification is specific and validating the relevance of the experiments. c, d, Bar graphs presenting the degree of potentiation from the recordings in a and b. Error bars represent ±s.d. for data obtained from five different oocytes per mutant combination. e–g, M2M but not M8M potentiates the mutant GluN1b–GluN2B NMDA receptor. The same experiment as above or in Fig. 2 was conducted using M2M or M8M on GluN1–4b(Ala175Cys)/GluN2B(Gln180Cys) (e) GluN1–4b(Lys178Cys)/GluN2B(Asn184Cys) (f), and wild-type GluN1–4b/GluN2B (g). Shown are representative electrophysiological recordings used to estimate the degree of bi-MTS potentiation presented in Fig. 2c.
Extended Data Figure 6 Cryo-EM analysis on GluN1b/GluN2B NMDA receptors.
a, Representative motion-corrected image collected at 22,500 magnification. b, Two-dimensional class averages. c, d, Fourier shell correlation curves for unmasked data (c) and model versus electron microscopy map (d). Class X and Y are similar to ‘non-active 2’ and were not further analysed. e, Orientation plots for each class, plotting the distribution of Euler angles assigned to all particles contributing to that class with an occupancy of at least 80%. For each class, the number of particles which have that class as their highest occupancy value is also shown.
Extended Data Figure 7 Representative cryo-EM density, model fit and structural comparison of the ATD in inhibited and active conformations of cryo-EM structures.
a–e, Here, the cryo-EM maps for non-active 2 and active 3D classes are shown along with the refined models. Densities are shown at the ATD and LBD (a–d) for both of the 3D classes and at the TMD (e) for non-active 2. f, g, Superimposition of R1 lobes of GluN1b (f) and GluN2B (g) illustrates the relative ‘opening’ between R1 and R2 lobes in the inhibited and active forms of intact NMDA receptors. The extent of GluN2B ATD opening is similar to that observed between the crystal structures of the ifenprodil–GluN1b–GluN2B ATD and the apo-GluN1b–GluN2B ATD as in Fig. 1. GluN1b and GluN2B ATDs are shown in grey and yellow for the inhibited and active states, respectively. h, Comparison of the GluN1b–GluN2B ATD heterodimers between ifenprodil inhibited and active cryo-EM structures. Superimposition of the GluN2B R1 lobes reveals an ~12° rotation of the GluN1b ATD relative to the GluN2B ATD in the similar manner to the crystal structure of the apo-GluN1b–GluN2B ATD as in Fig. 1. The black rods indicate the axis of rotation between the two cryo-EM structures. The distance of the R2 lobes in the GluN1b–GluN2B heterodimers is measured between Cα atoms of GluN1b(Lys178) and GluN2B(Asn184) (green spheres).
Extended Data Figure 8 Structural comparison of the GluN1–GluN2B LBD in non-active and active conformations to the GluA2 LBD in pre-open state.
a–d, The crystal structure of GluA2 AMPA receptor in the pre-open state (PDB ID, 4U1W; shown in green) aligned with the structures of GluN1–GluN2B in the non-active 2 (blue) (a, b) and active conformation (yellow) (c, d) by superimposing the LBDs of GluN2B onto GluA2. e–h, The equivalent superimposition with the cryo-EM structure of GluA2 AMPA receptor in the pre-open state (PDB ID, 4UQ6; shown in green). The overlaid structures are viewed through the LBD heterodimer interface (a, c or e, g) and the dimer of heterodimer interface (b, d or f, h). Here, the GluN2B LBD of the GluN1b–GluN2B NMDA receptor is superimposed onto the LBD of the GluA2 AMPA receptor and the shift of the GluN1 LBD is measured with respect to the other GluA2 LBD. The homodimeric arrangement of GluA2 AMPA receptor in the pre-open state is similar to the heterodimeric arrangements of GluN1b–GluN2B NMDA receptors in both non-active 2 and active states (a, c or e, g). However, when the dimer of homodimer arrangement of GluA2 AMPA receptor is compared to the dimer of heterodimers arrangement of the GluN1b–GluN2B NMDA receptor, a greater difference is observed for the active NMDA receptor (d, h) than for the non-active 2 NMDA receptor (b, f). Here, the non-active 2 NMDA receptor as in Fig. 3 is subjected to superimposition. The non-active 1 and non-active 2 NMDA receptors have similar subunit arrangements in the LBD. The numbers in each panel represent degrees of rotations and translations.
Supplementary information
Conformational transition of GluN1b-GluN2B NMDA receptors during activation
This video shows a morph of the GluN1b-GluN2B NMDA receptor structure from ‘non-active state 2’ to ‘active state’ as viewed from the top (left) and side (right). The linkers between the LBD and the TMD except the LBD-M3 linkers are omitted for clarity. The GluN1 (α), GluN1 (β), GluN2B (α) and GluN2B (β) subunits are colored in red, orange, blue and cyan, respectively. The first ordered residues on the linker between the M3 helices on TMD and the LBD in the active state structure (GluN1b Arg 684 and GluN2B Glu 658) are shown as spheres. Note that the TMD of GluN1b-GluN2B NMDA receptor in the ‘active’ state was not well resolved, and that the TMD shown in this video is that of GluN1b-GluN2B NMDA receptor in ‘non-active state 2.’ (MOV 648 kb)
Conformational transition of GluN1b-GluN2B NMDA receptors during ifenprodil inhibition
This video shows a morph of the GluN1b-GluN2B NMDA receptor structure from the ‘non-active state 2’ to the ‘inhibited’ state by ifenprodil (PDB ID: 4PE5) as viewed from the top (left) and side (right). The GluN1 (α), GluN1 (β), GluN2B (α) and GluN2B (β) subunits are colored in red, orange, blue and cyan, respectively. TMD of the GluN1b-GluN2B NMDA receptor in the ‘inhibited’ state was replaced with the TMD of the GluN1b-GluN2B NMDA receptor in the ‘non-active 2’ state. Ifenprodil is shown as green spheres. (MOV 617 kb)
Rights and permissions
About this article
Cite this article
Tajima, N., Karakas, E., Grant, T. et al. Activation of NMDA receptors and the mechanism of inhibition by ifenprodil. Nature 534, 63–68 (2016). https://doi.org/10.1038/nature17679
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17679
This article is cited by
-
Therapeutic potential of N-methyl-D-aspartate receptor modulators in psychiatry
Neuropsychopharmacology (2024)
-
Downstream Allosteric Modulation of NMDA Receptors by 3-Benzazepine Derivatives
Molecular Neurobiology (2023)
-
Structural insights into binding of therapeutic channel blockers in NMDA receptors
Nature Structural & Molecular Biology (2022)
-
Development and characterization of functional antibodies targeting NMDA receptors
Nature Communications (2022)
-
Complex functional phenotypes of NMDA receptor disease variants
Molecular Psychiatry (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.