Abstract
Umbilical cord blood-derived haematopoietic stem cells (HSCs) are essential for many life-saving regenerative therapies. However, despite their advantages for transplantation, their clinical use is restricted because HSCs in cord blood are found only in small numbers1. Small molecules that enhance haematopoietic stem and progenitor cell (HSPC) expansion in culture have been identified2,3, but in many cases their mechanisms of action or the nature of the pathways they impinge on are poorly understood. A greater understanding of the molecular circuitry that underpins the self-renewal of human HSCs will facilitate the development of targeted strategies that expand HSCs for regenerative therapies. Whereas transcription factor networks have been shown to influence the self-renewal and lineage decisions of human HSCs4,5, the post-transcriptional mechanisms that guide HSC fate have not been closely investigated. Here we show that overexpression of the RNA-binding protein Musashi-2 (MSI2) induces multiple pro-self-renewal phenotypes, including a 17-fold increase in short-term repopulating cells and a net 23-fold ex vivo expansion of long-term repopulating HSCs. By performing a global analysis of MSI2–RNA interactions, we show that MSI2 directly attenuates aryl hydrocarbon receptor (AHR) signalling through post-transcriptional downregulation of canonical AHR pathway components in cord blood HSPCs. Our study gives mechanistic insight into RNA networks controlled by RNA-binding proteins that underlie self-renewal and provides evidence that manipulating such networks ex vivo can enhance the regenerative potential of human HSCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Miller, P. H., Knapp, D. J. & Eaves, C. J. Heterogeneity in hematopoietic stem cell populations: implications for transplantation. Curr. Opin. Hematol. 20, 257–264 (2013)
Boitano, A. E. et al. Aryl hydrocarbon receptor antagonists promote the expansion of human hematopoietic stem cells. Science 329, 1345–1348 (2010)
Fares, I. et al. Pyrimidoindole derivatives are agonists of human hematopoietic stem cell self-renewal. Science 345, 1509–1512 (2014)
Novershtern, N. et al. Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144, 296–309 (2011)
Laurenti, E. et al. The transcriptional architecture of early human hematopoiesis identifies multilevel control of lymphoid commitment. Nature Immunol. 14, 756–763 (2013)
Hope, K. J. et al. An RNAi screen identifies Msi2 and Prox1 as having opposite roles in the regulation of hematopoietic stem cell activity. Cell Stem Cell 7, 101–113 (2010)
de Andrés-Aguayo, L. et al. Musashi 2 is a regulator of the HSC compartment identified by a retroviral insertion screen and knockout mice. Blood 118, 554–564 (2011)
Park, S. M. et al. Musashi-2 controls cell fate, lineage bias, and TGF-β signaling in HSCs. J. Exp. Med. 211, 71–87 (2014)
Ohyama, T. et al. Structure of Musashi1 in a complex with target RNA: the role of aromatic stacking interactions. Nucleic Acids Res. 40, 3218–3231 (2012)
Glimm, H. et al. Previously undetected human hematopoietic cell populations with short-term repopulating activity selectively engraft NOD/SCID-β2 microglobulin-null mice. J. Clin. Invest. 107, 199–206 (2001)
Cashman, J. D. & Eaves, C. J. Human growth factor-enhanced regeneration of transplantable human hematopoietic stem cells in nonobese diabetic/severe combined immunodeficient mice. Blood 93, 481–487 (1999)
Holyoake, T. L., Nicolini, F. E. & Eaves, C. J. Functional differences between transplantable human hematopoietic stem cells from fetal liver, cord blood, and adult marrow. Exp. Hematol. 27, 1418–1427 (1999)
Mimura, J. & Fujii-Kuriyama, Y. Functional role of AhR in the expression of toxic effects by TCDD. Biochim. Biophys. Acta 1619, 263–268 (2003)
Lo, R. & Matthews, J. High-resolution genome-wide mapping of AHR and ARNT binding sites by ChIP-seq. Toxicol. Sci. 130, 349–361 (2012)
Yeo, G. W. et al. An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Nature Struct. Mol. Biol . 16, 130–137 (2009)
Katz, Y. et al. Musashi proteins are post-transcriptional regulators of the epithelial-luminal cell state. Elife 3, e03915 (2014)
Tijet, N. et al. Aryl hydrocarbon receptor regulates distinct dioxin-dependent and dioxin-independent gene batteries. Mol. Pharmacol. 69, 140–153 (2006)
Doulatov, S. et al. PLZF is a regulator of homeostatic and cytokine-induced myeloid development. Genes Dev. 23, 2076–2087 (2009)
Majeti, R., Park, C. Y. & Weissman, I. L. Identification of a hierarchy of multipotent hematopoietic progenitors in human cord blood. Cell Stem Cell 1, 635–645 (2007)
Bagger, F. O. et al. HemaExplorer: a database of mRNA expression profiles in normal and malignant haematopoiesis. Nucleic Acids Res. 41, D1034–D1039 (2013)
Amendola, M., Venneri, M. A., Biffi, A., Vigna, E. & Naldini, L. Coordinate dual-gene transgenesis by lentiviral vectors carrying synthetic bidirectional promoters. Nature Biotechnol. 23, 108–116 (2005)
van Galen, P. et al. The unfolded protein response governs integrity of the haematopoietic stem-cell pool during stress. Nature 510, 268–272 (2014)
Lechman, E. R. et al. Attenuation of miR-126 activity expands HSC in vivo without exhaustion. Cell Stem Cell 11, 799–811 (2012)
Carow, C. E., Hangoc, G. & Broxmeyer, H. E. Human multipotential progenitor cells (CFU-GEMM) have extensive replating capacity for secondary CFU-GEMM: an effect enhanced by cord blood plasma. Blood 81, 942–949 (1993)
Milyavsky, M. et al. A distinctive DNA damage response in human hematopoietic stem cells reveals an apoptosis-independent role for p53 in self-renewal. Cell Stem Cell 7, 186–197 (2010)
Janky, R. et al. iRegulon: from a gene list to a gene regulatory network using large motif and track collections. PLOS Comput. Biol. 10, e1003731 (2014)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Kwon, A. T., Arenillas, D. J., Worsley Hunt, R. & Wasserman, W. W. oPOSSUM-3: advanced analysis of regulatory motif over-representation across genes or ChIP-Seq datasets. G3 2, 987–1002 (2012)
Colvin, G. A. et al. Murine marrow cellularity and the concept of stem cell competition: geographic and quantitative determinants in stem cell biology. Leukemia 18, 575–583 (2004)
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Darnell, R. CLIP (cross-linking and immunoprecipitation) identification of RNAs bound by a specific protein. Cold Spring Harb. Protoc. 2012, 1146–1160 (2012)
Lovci, M. T. et al. Rbfox proteins regulate alternative mRNA splicing through evolutionarily conserved RNA bridges. Nature Struct. Mol. Biol . 20, 1434–1442 (2013)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Dale, R. K., Pedersen, B. S. & Quinlan, A. R. Pybedtools: a flexible Python library for manipulating genomic datasets and annotations. Bioinformatics 27, 3423–3424 (2011)
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014)
Acknowledgements
We thank E. Lechman and P. Van Galen for experimental advice and for providing H1 and pSMALB vectors. The MA overexpression vector was a gift from L. Naldini. We also thank the SCC-RI core flow cytometry staff, the Obstetrics and Gynecology Unit at McMaster Children’s Hospital for cord blood, B. Doble and M. Bhatia for critical assessment of this work and all members of the Hope laboratory for experimental support and advice. This work was supported by an Ontario Institute for Cancer Research New Investigator Award (IA-033), an Ontario Institute for Cancer Research Cancer Stem Cell Program Team Grant (P.CSC.005) and a Canadian Institutes of Health Research (MOP-126030) grant to K.J.H. N.T.H was supported in part by a CIHR MD/PhD Studentship. M.S.B. was supported by an NSERC Alexander Graham Bell Doctoral Fellowship. S.R. is supported by a Canadian Blood Services Graduate Fellowship and Health Canada. The views expressed herein do not necessarily represent the view of the federal government of Canada. This work was partially supported by grants from the National Institute of Health (HG004659 and NS075449) and the California Institute of Regenerative Medicine (RB3-05219) to G.W.Y. G.P. was supported by a National Science Graduate Fellowship. G.W.Y. is an Alfred P. Sloan Research Fellow. We thank the UCSD Institute for Genomic Medicine’s Genomics Center for providing access to high-throughput sequencing facilities.
Author information
Authors and Affiliations
Contributions
S.R. designed and performed experiments, analysed data and wrote the manuscript. N.T.H. constructed CLIP–seq libraries. M.S.B. helped perform cord blood experiments. G.A.P. and G.W.Y advised on CLIP–seq library construction, performed CLIP–seq bioinformatic analyses and wrote the manuscript. B.T.W. performed RNA-seq analyses. V.V. and G.D.B performed RNA-seq bioinformatic analyses. K.J.H. conceived the project, supervised the study, analysed data, interpreted results and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
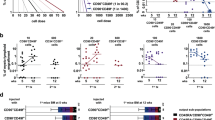
Extended Data Figure 1 MSI2 is highly expressed in human haematopoietic stem and progenitor cell populations.
a, Schematic of the human haematopoietic hierarchy showing key primitive cell populations and simplified surface marker expression. b, qRT–PCR analysis of MSI1 and MSI2 expression in Lin− cord blood (CB) cell populations (n = 3 independent Lin− CB samples). c, Gating strategy used to sort sub-fractions of Lin− CB HSPCs for MSI2 qRT–PCR expression analysis (n = 2 independent pooled Lin− CB samples). d, MSI2 expression across the human haematopoietic hierarchy. Each circle represents an independent gene expression data set curated by HemaExplorer. e, Intracellular flow cytometry analysis of MSI2 protein levels in Lin− CB. Histograms show background staining with secondary antibody (red) and positive staining with anti-MSI2 antibody plus secondary in Lin− CB (blue). MSI2 fluorescence intensity was divided into quartiles of negative (Q1), low (Q2), mid (Q3) and high (Q4) level expression. f, Plots show cell percentage within each quartile from e that are CD34+ CD38− (left) and CD34+ CD38+ (right) (n = 2 independent Lin− CB samples). All data presented as mean ± s.e.m. Unpaired t-test, *P < 0.05; ***P < 0.001.
Extended Data Figure 2 MSI2 overexpression enhances in vitro culture of primitive CB cells.
a, Top: schematic of bi-directional promoter lentivirus used to overexpress MSI2. Bottom: western blot and histogram showing intracellular flow validation of enforced MSI2 expression in 293FT cells (left) and Lin− CB (right), respectively. b, Representative images of secondary CFU made from replated control and MSI2-overexpressing (MSI2) CFU-GEMMs and types of colonies made. Scale bar, 200 μm. c, Fold change in Lin− CB transduced cell number after 7 days in culture following transduction (n = 5 experiments). d, Growth curve over 21 days of transduced Lin− CB cells (n = 4 experiments). e, Colony output of transduced Lin− CB from day 7 cultures (n = 8 cultures from 4 experiments). f, BrdU cell cycle analysis of transduced Lin− CB cells from day 10 cultures (n = 3 experiments). g, Ki67 cell cycle analysis of transduced Lin− CB cells from day 4 cultures (n = 4 experiments). h, Apoptotic and dead cells in day 7 cultures of transduced Lin− CB by Annexin V staining (n = 3 experiments). Western blot source data are available in Supplementary Fig. 1. All data presented as mean ± s.e.m. Unpaired t-test, **P < 0.01; ***P < 0.001.
Extended Data Figure 3 MSI2 overexpression does not affect STRC lineage output and extends STRC-mediated engraftment.
a, Schematic of STRC LDA experimental setup. b, Left: gating strategy to identify engrafted GFP+ CD45+ progenitor and myelo-lympho lineage-positive cell types or GFP+ CD45− erythroid cells and platelets. Right: summary of lineage output in the injected femur 3 weeks after transplantation (n = 4 mice for control and n = 18 mice for MSI2 overexpressing cells). MK, megakaryocyte; E, erythroid cells; P, platelets. c, Representative flow plots and summary of transduced STRC read out for engraftment with human CD45+ cells at 6.5 weeks post-transplant (n = 4 mice per condition). All data presented as mean ± s.e.m.
Extended Data Figure 4 MSI2 knockdown impairs secondary CFU replating potential and HSC engraftment capacity.
a, Left: schematic of MSI2- and control RFP-targeted shRNA lentiviruses. Right: confirmation of MSI2 protein knockdown (both isoforms that can be detected by western blot) in transduced NB4 cells. b, CFU production by shMSI2- and shControl-transduced Lin− CB (n = 8 cultures from 4 experiments). c, Secondary CFU output from shMSI2-transduced Lin− CB and images of representative secondary CFUs (scale bar, 200 μm; performed on n = 4 cultures from 2 experiments). d, Fold change in transduced cell number after 7 days in culture (n = 4 experiments). e, Growth curves of cultures initiated with transduced Lin− CB cells (n = 4 experiments). f, Experimental design to read out changes in HSC capacity with MSI2 knockdown. g, Left: representative flow analysis of transduced CD34+ CD38−-derived human chimaerism in NSG mouse bone marrow. Right: ratio of the percentage of GFP+ cells in the CD45+ population post-transplant to the initial pre-transplant GFP+ cell percentage. Dotted line indicates that the proportion of GFP+ cells is unchanged relative to input. One sample t-test, no change = 1; n = 6 mice receiving shControl and n = 8 mice receiving shMSI2-transduced cells pooled from two experiments. h, Representative flow plots and summary of multilineage engraftment with shControl and shMSI2 cells (gated on GFP+ cells). Western blot source data are shown in Supplementary Fig. 1. Data presented as mean ± s.e.m. Unpaired t-test, *P < 0.05; ***P < 0.001.
Extended Data Figure 5 MSI2 overexpression confers an HSC gene expression signature.
a, Genes that are upregulated (21 genes, logFC >0) or downregulated (156 genes, logFC <0) in MSI2-overexpressing (OE) cells relative to control cells with FDR < 0.05 were compared to expression data from MSI2 knockdown cells normalized to shControl expression data. Red circles represent 177 genes that were significantly differentially expressed in MSI2-overexpressing cells. Gray outlined circles represent random genes (equal number of grey circles and red circles). Only genes that were significantly up- or downregulated in MSI2-overexpressing cells showed anti-correlation with MSI2 knockdown cells. b, Genes that were differentially expressed between MSI2-overexpressing and control cells (FDR < 0.05) compared to DMAP populations. Numbers beside each bar indicate the percentage of time for which the observed value (set of up- or downregulated genes) was better represented in that population than random values (equal number of randomly selected genes based on 1,000 trials).
Extended Data Figure 6 MSI2 overexpression enhances HSC activity after ex vivo culture.
a, Experimental procedure for measuring changes in HSC engraftment capacity and frequency with ex vivo culture. b, Representative flow plots of CD45+ GFP+ reconstitution from mice receiving the highest cell dose transplanted per time point. c, Multilineage engraftment of mice injected with D3 cultures. d, Proportion of the human CD45+ graft containing GFP+ cells from mice receiving the two highest doses of D3 primary grafts relative to pre-transplant levels of GFP+ cells before expansion (n = 8 mice for each dose). e, Proportion of the human CD45+ graft containing GFP+ cells from mice receiving the two highest doses of D10 primary grafts relative to pre-transplant levels of GFP+ cells after expansion (n = 8 mice for each dose, one-sample t-test, no change = 1). f, Multilineage engraftment of mice injected with D10 cultures. g, GFP mean fluoresence intensity (MFI) in D10 primary cell-engrafted mice. Data are from mice transplanted with the highest three doses; n = 11 control and 13 MSI2-overexpressing cell-engrafted mice. h, CD34 expression in GFPhigh (top 60%) relative to GFPlow (bottom 40%) gated cells (set per mouse) from engrafted recipients in e. i, Number of transduced phenotyped HSCs after 7 days of culture from HSC expansion experiment described in a. Symbols represent individual mice and shaded symbols represent mice grafted with MSI2-overexpressing cells. All data presented as mean ± s.e.m. Unpaired t-test, *P < 0.05.
Extended Data Figure 7 Predicted AHR targets and genes downregulated by SR1 or MSI2 overexpression are upregulated by MSI2 knockdown.
a, Predicted AHR targets were identified with the iRegulon tool and compared with MSI2 knockdown normalized to shControl-upregulated gene signature by GSEA. b, log fold-change of MSI2-overexpression and knockdown shared leading edge AHR target genes from GSEA. c, GSEA comparing gene sets downregulated by SR1 high and low dose with the MSI2 knockdown upregulated gene signature. d, Heatmap and log fold-change of shared leading edge genes identified by GSEA from MSI2 overexpression, MSI2 knockdown and SR1 at varying doses. e, The percentage of downregulated genes in UM171-treated, SR1-treated and MSI2-overexpressing cells containing at least one AHR-binding site within 1,500 bp upstream or downstream of the transcription start site. Dotted line indicates the background percentage of genes with AHR-binding sites. P values were generated relative to background with Fisher’s exact test.
Extended Data Figure 8 AHR antagonism with SR1 has redundant effects with MSI2 overexpression, and AHR activation with MSI2 overexpression results in a loss of HSPCs.
a, Representative flow plots and summary of CD34 expression in MSI2-overexpressing and control transduced CD34+ CB cells grown for 10 days in medium containing SR1 or DMSO vehicle (n = 3 experiments). b, Fold change in CD34 expression from results shown in a. c, Fold increase in CYP1B1 and AHRR transcript levels after FICZ treatment in transduced cultures (n = 3 experiments). d, Transduced CD34+ CB cells grown for 3 days in medium supplemented with FICZ and the corresponding change in CD34 expression. Each coloured pair (DMSO and FICZ) represents a matched CB sample (n = 3 experiments). e, Differences in culture CD34 levels from d. All data presented as mean ± s.e.m. Unpaired t-test, *P < 0.05.
Extended Data Figure 9 MSI2 preferentially binds mature mRNA within the 3′UTR.
a, Validation of the capacity of the anti-MSI2 antibody to immunoprecipitate MSI2 compared to IgG control pulldowns (heavy chain, HC; light chain, LC). b, Autoradiogram showing anti-MSI2 immunoprecipitated, MNase digested and radiolabelled RNA isolated for CLIP library construction and sequencing (red box). Low levels of MNase show a smearing pattern extending upwards from the modal weight of MSI2. c, Scatter plot of total number of uniquely mapped CLIP–seq reads for each gene, comparing both replicates. d, Heatmap indicating the number of different classes of Gencode-annotated genes that contain at least one predicted MSI-binding site. e, Consensus motifs within MSI2 clusters in the different genic regions. P values for the most statistically significant enriched motif are presented for all overlapping clusters between replicates. f, Cumulative distribution function of mean conservation score (Phastcons) of MSI2 clusters, compared to a shuffled background control, computed for all overlapping clusters and the top 40% of overlapping clusters. P values were obtained by a Kolgomorov–Smirnov two-tailed test comparing the distributions from actual and shuffled locations. g, Number of clusters within 200 bases of the annotated stop codon in known mRNA transcripts for all overlapping clusters between replicates and the top 40% of overlapping clusters. h, Cumulative distribution function of mean conservation score (Phastcons) of MSI2 clusters, compared to a shuffled background control, computed for overlapping clusters between the replicates and the top 40% of overlapping clusters found in different genic regions. Similarity between the 3′UTR conservation for the top 40% and the shuffled background control is probably due to MSI2 sites being small and not needing structural contexts for conservation. P values were obtained by a Kolgomorov–Smirnov two-tailed test comparing the distributions from actual and shuffled locations. i, Genome browser views displaying CLIP–seq mapped reads from replicate 1 (blue), predicted clusters (purple), exact matches for the GUAG sequence (black) and mammal conservation scores (PhyloP) in the 3′UTRs for a previously predicted Msi1 target.
Extended Data Figure 10 MSI2 overexpression represses CYP1B1 and HSP90 3′UTR Renilla Luciferase reporter activity.
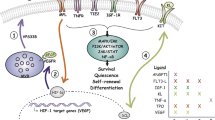
a, CLIP–seq reads (replicate 1 in blue and replicate 2 in green) and clusters (purple) mapped to the 3′UTR of HSP90. Matches to the GUAG motif are shown in black. Mammal PhyloP score listed in last track. b, c, Representative data of mean per cell fluoresence for HSP90 and CYP1B1 protein in transduced CD34+ CB. Protein level in cells during in vitro culture was analysed 3 days (D3) and 7 days (D7) after transduction and sorting for GFP. Corresponding secondary-alone antibody staining is shown for each experiment. Each circle represents a cell, and more than 200 cells were analysed per condition. d, e, Levels of renilla luciferase activity in NIH-3T3 cells co-transfected with control or MSI2 overexpression vectors and the CYP1B1 or HSP90 wild-type or TCC mutant 3′UTR luciferase reporter (dotted line indicates no change in renilla activity; n = 4 CYP1B1 and n = 3 HSP90 experiments). f, Flow plots of co-transduced CD34+ CB cells with MSI2 (GFP) and CYP1B1 (BFP) lentivirus. g, GFP+ BFP+ CFU-GEMMs generated from f were replated into secondary CFU assays and the total number of colonies formed was counted. A total of 24 CFU-GEMMs from MSI2-BFP and MSI2-CYP1B1 were replated (n = 2 experiments). Data presented as mean ± s.e.m. Unpaired t-test, ***P < 0.001, **P < 0.01. h, A model for AHR pathway attenuation by MSI2 post-transcriptional processing. MSI2 mediates the post-transcriptional downregulation of HSP90 at the outset of culture and continuously represses the prominent AHR pathway effector CYP1B1 to facilitate HSPC expansion. The resulting MSI2-mediated repression of AHR signalling enforces a self-renewal program and allows HSPC expansion ex vivo.
Supplementary information
Supplementary Figure
This file contains the uncropped western blots. (PDF 551 kb)
Supplementary Data
This file contains Supplementary Tables 1-7. (XLSX 6193 kb)
Rights and permissions
About this article
Cite this article
Rentas, S., Holzapfel, N., Belew, M. et al. Musashi-2 attenuates AHR signalling to expand human haematopoietic stem cells. Nature 532, 508–511 (2016). https://doi.org/10.1038/nature17665
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17665
This article is cited by
-
Loss of RNA-binding protein CELF2 promotes acute leukemia development via FAT10-mTORC1
Oncogene (2024)
-
Tumor cell-released kynurenine biases MEP differentiation into megakaryocytes in individuals with cancer by activating AhR–RUNX1
Nature Immunology (2023)
-
Impaired expression of BCAT1 relates to muscle atrophy of mouse model of sarcopenia
BMC Musculoskeletal Disorders (2022)
-
Expansion of Human Megakaryocyte-Lineage Progeny via Aryl Hydrocarbon Receptor Antagonism with CH223191
Stem Cell Reviews and Reports (2022)
-
Decoding Human Hematopoietic Stem Cell Self-Renewal
Current Stem Cell Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.