Abstract
Cytolytic proteins and peptide toxins are classical virulence factors of several bacterial pathogens which disrupt epithelial barrier function, damage cells and activate or modulate host immune responses. Such toxins have not been identified previously in human pathogenic fungi. Here we identify the first, to our knowledge, fungal cytolytic peptide toxin in the opportunistic pathogen Candida albicans. This secreted toxin directly damages epithelial membranes, triggers a danger response signalling pathway and activates epithelial immunity. Membrane permeabilization is enhanced by a positive charge at the carboxy terminus of the peptide, which triggers an inward current concomitant with calcium influx. C. albicans strains lacking this toxin do not activate or damage epithelial cells and are avirulent in animal models of mucosal infection. We propose the name ‘Candidalysin’ for this cytolytic peptide toxin; a newly identified, critical molecular determinant of epithelial damage and host recognition of the clinically important fungus, C. albicans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
01 April 2016
A label in Fig. 3b of the PDF file initially published online was corrupted and was replaced.
07 April 2016
A missing citation to the Extended Data Tables was corrected in the HTML on 7 April 2016.
References
Brown, G. D. et al. Hidden killers: human fungal infections. Sci. Transl Med. 4, 165rv113 (2012)
Jacobsen, I. D. et al. Candida albicans dimorphism as a therapeutic target. Expert Rev. Anti Infect. Ther. 10, 85–93 (2012)
Moyes, D. L. et al. A biphasic innate immune MAPK response discriminates between the yeast and hyphal forms of Candida albicans in epithelial cells. Cell Host Microbe 8, 225–235 (2010)
Moyes, D. L. et al. Candida albicans yeast and hyphae are discriminated by MAPK signaling in vaginal epithelial cells. PLoS ONE 6, e26580 (2011)
Murciano, C. et al. Candida albicans cell wall glycosylation may be indirectly required for activation of epithelial cell proinflammatory responses. Infect. Immun. 79, 4902–4911 (2011)
Moyes, D. L. et al. Activation of MAPK/c-Fos induced responses in oral epithelial cells is specific to Candida albicans and Candida dubliniensis hyphae. Med. Microbiol. Immunol. (Berl.) 201, 93–101 (2012)
Murciano, C. et al. Evaluation of the role of Candida albicans agglutinin-like sequence (Als) proteins in human oral epithelial cell interactions. PLoS ONE 7, e33362 (2012)
Moyes, D. L. & Naglik, J. R. Mucosal immunity and Candida albicans infection. Clin. Dev. Immunol. 2011, 346307 (2011)
Naglik, J. R. & Moyes, D. Epithelial cell innate response to Candida albicans. Adv. Dent. Res. 23, 50–55 (2011)
Naglik, J. R., Moyes, D. L., Wachtler, B. & Hube, B. Candida albicans interactions with epithelial cells and mucosal immunity. Microbes Infect. 13, 963–976 (2011)
Hebecker, B., Naglik, J. R., Hube, B. & Jacobsen, I. D. Pathogenicity mechanisms and host response during oral Candida albicans infections. Expert Rev. Anti Infect. Ther. 12, 867–879 (2014)
Naglik, J. R. Candida immunity. New J. Science 2014, 390241 (2014)
Naglik, J. R., Richardson, J. P. & Moyes, D. L. Candida albicans Pathogenicity and epithelial immunity. PLoS Pathog. 10, e1004257 (2014)
Moyes, D. L., Richardson, J. P. & Naglik, J. R. Candida albicans—epithelial interactions and pathogenicity mechanisms: scratching the surface. Virulence 6, 338–346 (2015)
Birse, C. E., Irwin, M. Y., Fonzi, W. A. & Sypherd, P. S. Cloning and characterization of ECE1, a gene expressed in association with cell elongation of the dimorphic pathogen Candida albicans. Infect. Immun. 61, 3648–3655 (1993)
Röhm, M. et al. A family of secreted pathogenesis-related proteins in Candida albicans. Mol. Microbiol. 87, 132–151 (2013)
Kamai, Y., Kubota, M., Hosokawa, T., Fukuoka, T. & Filler, S. G. New model of oropharyngeal candidiasis in mice. Antimicrob. Agents Chemother. 45, 3195–3197 (2001)
Brothers, K. M. et al. NADPH oxidase-driven phagocyte recruitment controls Candida albicans filamentous growth and prevents mortality. PLoS Pathog. 9, e1003634 (2013)
Gratacap, R. L., Rawls, J. F. & Wheeler, R. T. Mucosal candidiasis elicits NF-κB activation, proinflammatory gene expression and localized neutrophilia in zebrafish. Dis. Model. Mech. 6, 1260–1270 (2013)
Bader, O., Krauke, Y. & Hube, B. Processing of predicted substrates of fungal Kex2 proteinases from Candida albicans, C. glabrata, Saccharomyces cerevisiae and Pichia pastoris. BMC Microbiol. 8, 116 (2008)
Newport, G. & Agabian, N. KEX2 influences Candida albicans proteinase secretion and hyphal formation. J. Biol. Chem. 272, 28954–28961 (1997)
Liu, P., Huang, X., Zhou, R. & Berne, B. J. Observation of a dewetting transition in the collapse of the melittin tetramer. Nature 437, 159–162 (2005)
Bechinger, B. & Salnikov, E. S. The membrane interactions of antimicrobial peptides revealed by solid-state NMR spectroscopy. Chem. Phys. Lipids 165, 282–301 (2012)
Pieta, P., Mirza, J. & Lipkowski, J. Direct visualization of the alamethicin pore formed in a planar phospholipid matrix. Proc. Natl Acad. Sci. USA 109, 21223–21227 (2012)
Bischofberger, M., Iacovache, I. & van der Goot, F. G. Pathogenic pore-forming proteins: function and host response. Cell Host Microbe 12, 266–275 (2012)
Los, F. C., Randis, T. M., Aroian, R. V. & Ratner, A. J. Role of pore-forming toxins in bacterial infectious diseases. Microbiol. Mol. Biol. Rev. 77, 173–207 (2013)
Oren, Z. & Shai, Y. Selective lysis of bacteria but not mammalian cells by diastereomers of melittin: structure-function study. Biochemistry 36, 1826–1835 (1997)
Walev, I. et al. Delivery of proteins into living cells by reversible membrane permeabilization with streptolysin-O. Proc. Natl Acad. Sci. USA 98, 3185–3190 (2001)
Schmitt, M. J. & Breinig, F. Yeast viral killer toxins: lethality and self-protection. Nature Rev. Microbiol. 4, 212–221 (2006)
Zakikhany, K. et al. In vivo transcript profiling of Candida albicans identifies a gene essential for interepithelial dissemination. Cell. Microbiol. 9, 2938–2954 (2007)
Wächtler, B. et al. Candida albicans-epithelial interactions: dissecting the roles of active penetration, induced endocytosis and host factors on the infection process. PLoS ONE 7, e36952 (2012)
Rupniak, H. T. et al. Characteristics of four new human cell lines derived from squamous cell carcinomas of the head and neck. J. Natl. Cancer Inst. 75, 621–635 (1985)
Mayer, F. L. et al. The novel Candida albicans transporter Dur31 is a multi-stage pathogenicity factor. PLoS Pathog. 8, e1002592 (2012)
Gillum, A. M., Tsay, E. Y. & Kirsch, D. R. Isolation of the Candida albicans gene for orotidine-5′-phosphate decarboxylase by complementation of S. cerevisiae ura3 and E. coli pyrF mutations. Mol. Gen. Genet. 198, 179–182 (1984)
Citiulo, F. et al. Candida albicans scavenges host zinc via Pra1 during endothelial invasion. PLoS Pathog. 8, e1002777 (2012)
Gola, S., Martin, R., Walther, A., Dunkler, A. & Wendland, J. New modules for PCR-based gene targeting in Candida albicans: rapid and efficient gene targeting using 100 bp of flanking homology region. Yeast 20, 1339–1347 (2003)
Wilson, R. B., Davis, D. & Mitchell, A. P. Rapid hypothesis testing with Candida albicans through gene disruption with short homology regions. J. Bacteriol. 181, 1868–1874 (1999)
Walther, A. & Wendland, J. An improved transformation protocol for the human fungal pathogen Candida albicans. Curr. Genet. 42, 339–343 (2003)
Murad, A. M., Lee, P. R., Broadbent, I. D., Barelle, C. J. & Brown, A. J. CIp10, an efficient and convenient integrating vector for Candida albicans. Yeast 16, 325–327 (2000)
Gratacap, R. L., Rawls, J. F. & Wheeler, R. T. Mucosal candidiasis elicits NF-κB activation, proinflammatory gene expression and localized neutrophilia in zebrafish. Dis. Model. Mech. 6, 1260–1270 (2013)
Moyes, D. L. et al. A biphasic innate immune MAPK response discriminates between the yeast and hyphal forms of Candida albicans in epithelial cells. Cell Host Microbe 8, 225–235 (2010)
Moyes, D. L. et al. Candida albicans yeast and hyphae are discriminated by MAPK signaling in vaginal epithelial cells. PLoS ONE 6, e26580 (2011)
Wächtler, B., Wilson, D., Haedicke, K., Dalle, F. & Hube, B. From attachment to damage: defined genes of Candida albicans mediate adhesion, invasion and damage during interaction with oral epithelial cells. PLoS ONE 6, e17046 (2011)
Trinh L, A. et al. A versatile gene trap to visualize and interrogate the function of the vertebrate proteome. Genes Dev. 25, 2306–2320 (2011)
Solis, N. V. & Filler, S. G. Mouse model of oropharyngeal candidiasis. Nature Protocols 7, 637–642 (2012)
Cranfield, C., Carne, S., Martinac, B. & Cornell, B. The assembly and use of tethered bilayer lipid membranes (tBLMs). Methods Mol. Biol. 1232, 45–53 (2015)
Cranfield, C. G. et al. Transient potential gradients and impedance measures of tethered bilayer lipid membranes: pore-forming peptide insertion and the effect of electroporation. Biophys. J. 106, 182–189 (2014)
Schromm, A. B. et al. Lipopolysaccharide-binding protein mediates CD14-independent intercalation of lipopolysaccharide into phospholipid membranes. FEBS Lett. 399, 267–271 (1996)
Chen, G. C. & Yang, J. T. 2-Point calibration of circular dichrometer with D-10-camphorsulfonic acid. Anal. Lett. 10, 1195–1207 (1977)
Montal, M. & Mueller, P. Formation of bimolecular membranes from lipid monolayers and a study of their electrical properties. Proc. Natl Acad. Sci. USA 69, 3561–3566 (1972)
Gutsmann, T., Heimburg, T., Keyser, U., Mahendran, K. R. & Winterhalter, M. Protein reconstitution into freestanding planar lipid membranes for electrophysiological characterization. Nature Protocols 10, 188–198 (2015)
Gillum, A. M., Tsay, E. Y. & Kirsch, D. R. Isolation of the Candida albicans gene for orotidine-5′-phosphate decarboxylase by complementation of S. cerevisiae ura3 and E. coli pyrF mutations. Mol. Gen. Genet. 198, 179–182 (1984)
Wilson, R. B., Davis, D. & Mitchell, A. P. Rapid hypothesis testing with Candida albicans through gene disruption with short homology regions. J. Bacteriol. 181, 1868–1874 (1999)
Fonzi, W. A. & Irwin, M. Y. Isogenic strain construction and gene mapping in Candida albicans. Genetics 134, 717–728 (1993)
Davis, D., Wilson, R. B. & Mitchell, A. P. RIM101-dependent and-independent pathways govern pH responses in Candida albicans. Mol. Cell. Biol. 20, 971–978 (2000)
Braun, B. R. & Johnson, A. D. TUP1, CPH1 and EFG1 make independent contributions to filamentation in Candida albicans. Genetics 155, 57–67 (2000)
Lo, H. J. et al. Nonfilamentous C. albicans mutants are avirulent. Cell 90, 939–949 (1997)
Moyes, D. L. et al. A biphasic innate immune MAPK response discriminates between the yeast and hyphal forms of Candida albicans in epithelial cells. Cell Host Microbe 8, 225–235 (2010)
Zakikhany, K. et al. In vivo transcript profiling of Candida albicans identifies a gene essential for interepithelial dissemination. Cell. Microbiol. 9, 2938–2954 (2007)
Cao, F. et al. The Flo8 transcription factor is essential for hyphal development and virulence in Candida albicans. Mol. Biol. Cell 17, 295–307 (2006)
Bockmühl, D. P., Krishnamurthy, S., Gerads, M., Sonneborn, A. & Ernst, J. F. Distinct and redundant roles of the two protein kinase A isoforms Tpk1p and Tpk2p in morphogenesis and growth of Candida albicans. Mol. Microbiol. 42, 1243–1257 (2001)
Sonneborn, A. et al. Protein kinase A encoded by TPK2 regulates dimorphism of Candida albicans. Mol. Microbiol. 35, 386–396 (2000)
Palmer, G. E., Cashmore, A. & Sturtevant, J. Candida albicans VPS11 is required for vacuole biogenesis and germ tube formation. Eukaryot. Cell 2, 411–421 (2003)
Zou, H., Fang, H. M., Zhu, Y. & Wang, Y. Candida albicans Cyr1, Cap1 and G-actin form a sensor/effector apparatus for activating cAMP synthesis in hyphal growth. Mol. Microbiol. 75, 579–591 (2010)
Bates, S. et al. Outer chain N-glycans are required for cell wall integrity and virulence of Candida albicans. J. Biol. Chem. 281, 90–98 (2006)
Murciano, C. et al. Candida albicans cell wall glycosylation may be indirectly required for activation of epithelial cell proinflammatory responses. Infect. Immun. 79, 4902–4911 (2011)
Newport, G. & Agabian, N. KEX2 influences Candida albicans proteinase secretion and hyphal formation. J. Biol. Chem. 272, 28954–28961 (1997)
Murad, A. M. et al. NRG1 represses yeast-hypha morphogenesis and hypha-specific gene expression in Candida albicans. EMBO J. 20, 4742–4752 (2001)
Liu, H., Kohler, J. & Fink, G. R. Suppression of hyphal formation in Candida albicans by mutation of a STE12 homolog. Science 266, 1723–1726 (1994)
Lane, S., Zhou, S., Pan, T., Dai, Q. & Liu, H. The basic helix-loop-helix transcription factor Cph2 regulates hyphal development in Candida albicans partly via TEC1. Mol. Cell. Biol. 21, 6418–6428 (2001)
White, S. J. et al. Self-regulation of Candida albicans population size during GI colonization. PLoS Pathog. 3, e184 (2007)
Brown, D. H., Jr, Giusani, A. D., Chen, X. & Kumamoto, C. A. Filamentous growth of Candida albicans in response to physical environmental cues and its regulation by the unique CZF1 gene. Mol. Microbiol. 34, 651–662 (1999)
Kadosh, D. & Johnson, A. D. Rfg1, a protein related to the Saccharomyces cerevisiae hypoxic regulator Rox1, controls filamentous growth and virulence in Candida albicans. Mol. Cell. Biol. 21, 2496–2505 (2001)
San José, C., Monge, R. A., Perez-Diaz, R., Pla, J. & Nombela, C. The mitogen-activated protein kinase homolog HOG1 gene controls glycerol accumulation in the pathogenic fungus Candida albicans. J. Bacteriol. 178, 5850–5852 (1996)
Firon, A. et al. The SUN41 and SUN42 genes are essential for cell separation in Candida albicans. Mol. Microbiol. 66, 1256–1275 (2007)
de Boer, A. D. et al. The Candida albicans cell wall protein Rhd3/Pga29 is abundant in the yeast form and contributes to virulence. Yeast 27, 611–624 (2010)
Mühlschlegel, F. A. & Fonzi, W. A. PHR2 of Candida albicans encodes a functional homolog of the pH-regulated gene PHR1 with an inverted pattern of pH-dependent expression. Mol. Cell. Biol. 17, 5960–5967 (1997)
Martin, R. et al. A core filamentation response network in Candida albicans is restricted to eight genes. PLoS ONE 8, e58613 (2013)
Birse, C. E., Irwin, M. Y., Fonzi, W. A. & Sypherd, P. S. Cloning and characterization of ECE1, a gene expressed in association with cell elongation of the dimorphic pathogen Candida albicans. Infect. Immun. 61, 3648–3655 (1993)
Navarro-García, F., Sanchez, M., Pla, J. & Nombela, C. Functional characterization of the MKC1 gene of Candida albicans, which encodes a mitogen-activated protein kinase homolog related to cell integrity. Mol. Cell. Biol. 15, 2197–2206 (1995)
Hausauer, D. L., Gerami-Nejad, M., Kistler-Anderson, C. & Gale, C. A. Hyphal guidance and invasive growth in Candida albicans require the Ras-like GTPase Rsr1p and its GTPase-activating protein Bud2p. Eukaryot. Cell 4, 1273–1286 (2005)
Sentandreu, M., Elorza, M. V., Sentandreu, R. & Fonzi, W. A. Cloning and characterization of PRA1, a gene encoding a novel pH-regulated antigen of Candida albicans. J. Bacteriol. 180, 282–289 (1998)
Pardini, G. et al. The CRH family coding for cell wall glycosylphosphatidylinositol proteins with a predicted transglycosidase domain affects cell wall organization and virulence of Candida albicans. J. Biol. Chem. 281, 40399–40411 (2006)
Braun, B. R., Head, W. S., Wang, M. X. & Johnson, A. D. Identification and characterization of TUP1-regulated genes in Candida albicans. Genetics 156, 31–44 (2000)
Fradin, C. et al. Granulocytes govern the transcriptional response, morphology and proliferation of Candida albicans in human blood. Mol. Microbiol. 56, 397–415 (2005)
Staab, J. F., Bradway, S. D., Fidel, P. L. & Sundstrom, P. Adhesive and mammalian transglutaminase substrate properties of Candida albicans Hwp1. Science 283, 1535–1538 (1999)
Bailey, D. A., Feldmann, P. J., Bovey, M., Gow, N. A. & Brown, A. J. The Candida albicans HYR1 gene, which is activated in response to hyphal development, belongs to a gene family encoding yeast cell wall proteins. J. Bacteriol. 178, 5353–5360 (1996)
Sandini, S., La Valle, R., De Bernardis, F., Macri, C. & Cassone, A. The 65 kDa mannoprotein gene of Candida albicans encodes a putative β-glucanase adhesin required for hyphal morphogenesis and experimental pathogenicity. Cell. Microbiol. 9, 1223–1238 (2007)
Csank, C. et al. Roles of the Candida albicans mitogen-activated protein kinase homolog, Cek1p, in hyphal development and systemic candidiasis. Infect. Immun. 66, 2713–2721 (1998)
Hube, B. et al. Disruption of each of the secreted aspartyl proteinase genes SAP1, SAP2, and SAP3 of Candida albicans attenuates virulence. Infect. Immun. 65, 3529–3538 (1997)
Taylor, B. N. et al. Induction of SAP7 correlates with virulence in an intravenous infection model of candidiasis but not in a vaginal infection model in mice. Infect. Immun. 73, 7061–7063 (2005)
Schild, L. et al. Proteolytic cleavage of covalently linked cell wall proteins by Candida albicans Sap9 and Sap10. Eukaryot. Cell 10, 98–109 (2011)
Zhao, X. et al. ALS3 and ALS8 represent a single locus that encodes a Candida albicans adhesin; functional comparisons between Als3p and Als1p. Microbiol . 150, 2415–2428 (2004)
Murciano, C. et al. Evaluation of the role of Candida albicans agglutinin-like sequence (Als) proteins in human oral epithelial cell interactions. PLoS ONE 7, e33362 (2012)
Zhao, X., Oh, S. H., Yeater, K. M. & Hoyer, L. L. Analysis of the Candida albicans Als2p and Als4p adhesins suggests the potential for compensatory function within the Als family. Microbiol. 151, 1619–1630 (2005)
Zhao, X., Oh, S. H. & Hoyer, L. L. Deletion of ALS5, ALS6 or ALS7 increases adhesion of Candida albicans to human vascular endothelial and buccal epithelial cells. Med. Mycol. 45, 429–434 (2007)
Zhao, X., Oh, S. H. & Hoyer, L. L. Unequal contribution of ALS9 alleles to adhesion between Candida albicans and human vascular endothelial cells. Microbiol. 153, 2342–2350 (2007)
Timpel, C., Strahl-Bolsinger, S., Ziegelbauer, K. & Ernst, J. F. Multiple functions of Pmt1p-mediated protein O-mannosylation in the fungal pathogen Candida albicans. J. Biol. Chem. 273, 20837–20846 (1998)
Bates, S. et al. Candida albicans Pmr1p, a secretory pathway P-type Ca2+/Mn2+-ATPase, is required for glycosylation and virulence. J. Biol. Chem. 280, 23408–23415 (2005)
Hobson, R. P. et al. Loss of cell wall mannosylphosphate in Candida albicans does not influence macrophage recognition. J. Biol. Chem. 279, 39628–39635 (2004)
Southard, S. B., Specht, C. A., Mishra, C., Chen-Weiner, J. & Robbins, P. W. Molecular analysis of the Candida albicans homolog of Saccharomyces cerevisiae MNN9, required for glycosylation of cell wall mannoproteins. J. Bacteriol. 181, 7439–7448 (1999)
Munro, C. A. et al. Mnt1p and Mnt2p of Candida albicans are partially redundant α-1,2-mannosyltransferases that participate in O-linked mannosylation and are required for adhesion and virulence. J. Biol. Chem. 280, 1051–1060 (2005)
Mio, T. et al. Role of three chitin synthase genes in the growth of Candida albicans. J. Bacteriol. 178, 2416–2419 (1996)
Mille, C. et al. Inactivation of CaMIT1 inhibits Candida albicans phospholipomannan β-mannosylation, reduces virulence, and alters cell wall protein β-mannosylation. J. Biol. Chem. 279, 47952–47960 (2004)
Mille, C. et al. Identification of a new family of genes involved in β-1,2-mannosylation of glycans in Pichia pastoris and Candida albicans. J. Biol. Chem. 283, 9724–9736 (2008)
Mille, C. et al. Members 5 and 6 of the Candida albicans BMT family encode enzymes acting specifically on β-mannosylation of the phospholipomannan cell-wall glycosphingolipid. Glycobiol . 22, 1332–1342 (2012)
Mio, T. et al. Cloning of the Candida albicans homolog of Saccharomyces cerevisiae GSC1/FKS1 and its involvement in β-1,3-glucan synthesis. J. Bacteriol. 179, 4096–4105 (1997)
Mio, T. et al. Isolation of the Candida albicans homologs of Saccharomyces cerevisiae KRE6 and SKN1: expression and physiological function. J. Bacteriol. 179, 2363–2372 (1997)
Staab, J. F. & Sundstrom, P. URA3 as a selectable marker for disruption and virulence assessment of Candida albicans genes. Trends Microbiol. 11, 69–73 (2003)
Murad, A. M., Lee, P. R., Broadbent, I. D., Barelle, C. J. & Brown, A. J. CIp10, an efficient and convenient integrating vector for Candida albicans. Yeast 16, 325–327 (2000)
Acknowledgements
We thank S. Gaffen, B. Klein, C. Hertweck, A. Tucker, J. Green and S. Challacombe for comments on the manuscript. For experimental assistance, we thank S. Bevan and D. Andersson (calcium assays), D. Nayar (histology), D. Rahman and M. Mistry (murine model), M. Nilan (zebrafish model), S. Groth (FRET spectroscopy), N. Gebauer (Impedance experiments), D. Schulz (kex1∆/∆ strain) and our colleagues for supplying fungal mutant strains. This work was supported by grants from the Medical Research Council (MR/J008303/1, MR/M011372/1), Biotechnology & Biological Sciences Research Council (BB/J015261/1), FP7-PEOPLE-2013-Initial Training Network (606786) to J.R.N.; Wellcome Trust Strategic Award for Medical Mycology and Fungal Immunology (097377/Z/11/Z) to J.R.N. and D.W.; Sir Henry Dale Fellowship jointly funded by the Wellcome Trust and the Royal Society (102549/Z/13/Z) to D.W.; Deutsche Forschungsgemeinschaft CRC/TR124 FungiNet Project C1 and Z2, Deutsche Forschungsgemeinschaft SPP 1580 (Hu 528/17-1) and CSCC, German Federal Ministry of Education and Health (BMBF) 01EO1002 to B.Hu.; Cluster of Excellence ‘Inflammation at interfaces’ and Deutsche Forschungsgemeinschaft SPP1580 project GU 568/5-1 to T.G.; National Institutes of Health (R15AI094406) and the Burroughs Wellcome Fund to R.T.W.
Author information
Authors and Affiliations
Contributions
D.L.M., J.P.R., S.X.T., M.R., C.M., M.B., S.I.I. and N.K. performed signalling, transcription factor, calcium and cytokine assays, and murine work; D.W., S.H., S.M., T.M.F., B.He., L.K. A.H., O.B. and O.Ku. created fungal strains and performed fluorescent microscopy, adhesion, invasion, gene expression and damage assays; R.L.G. and R.T.W. performed zebrafish experiments; J.W. and T.G. performed biophysical analysis with artificial membranes; J.R. performed whole patch clamp analysis; G.V. performed electron microscopy; S.T. performed histological analysis; S.M., T.L., T.K. and O.Kn. performed LC-MS analyses; J.R.N., B.Hu., D.L.M., J.P.R. and D.W. wrote the paper; J.R.N., B.Hu. and E.C. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 C. albicans ECE1 expression and phenotypic effects of ECE1 gene deletion.
a, Relative expression (vs t = 0) of ECE1 in C. albicans wild type over time after addition of yeast cells to TR146 epithelial cells as measured by RT-qPCR. b, Imaging confirmation of ECE1 expression over time within C. albicans wild-type strain. C. albicans cells expressing GFP under the control of the ECE1 5′ intragenic region, containing the ECE1 promoter, were grown on TR146 epithelial cells and stained with calcofluor white (CFW, post-permeabilization) to show cell wall chitin and Alexa-Fluor-647-labelled concanavalin A (ConA, pre-permeabilization) to show carbohydrates. A composite image showing CFW, ConA, GFP and the brightfield (BF) image is shown. c, Scanning electron micrographs (top panels, 5 h) and light microscopy (bottom panels, 24 h) showing no gross abnormalities in hypha formation between C. albicans wild-type (BWP17 + CIp30), ECE1-deletion (ece1Δ/Δ) and ECE1 re-integrant (ece1Δ/Δ + ECE1) strains after infection of TR146 epithelial cells. d, No difference in adhesion of C. albicans wild-type, ece1Δ/Δ and ece1Δ/Δ + ECE1 strains to TR146 epithelial cells after 60 min. e, No difference in invasion of C. albicans wild-type, ece1Δ/Δ and ece1Δ/Δ + ECE1 strains into TR146 epithelial cells after 3 h. f, Fluorescence staining of C. albicans wild-type and ece1Δ/Δ hyphae invading through TR146 epithelial cells. Fungal cells are stained with calcofluor white (CFW, post-permeabilization) and Alexa-Fluor-647-labelled concanavalin A (ConA, pre-permeabilization) to show cell wall chitin and carbohydrates, respectively, and to distinguish between invading hyphae (only stained after permeabilization) and non-invading hyphae (stained both pre- and post-permeabilization). Levels of chitin and β-glucan are comparable in both strains. White arrows indicate invasion into epithelial cells. Data shown are representative (b, c, f) or the mean (a, d, e) of three biological replicates. Error bars show ± s.e.m.
Extended Data Figure 2 C. albicans Ece1p is critical for mucosal virulence in vivo.
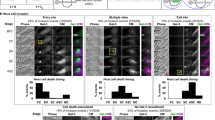
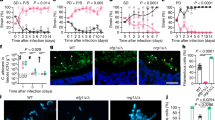
a, Fungal burdens recovered from the tongues of mice infected with C. albicans wild-type (BWP17 + CIp30) (number of mice(n) = 13), ECE1-deletion (ece1Δ/Δ) (n = 20) and ECE1 re-integrant (ece1Δ/Δ + ECE1) (n = 24) strains after a 2-day oropharyngeal infection. b, Average percentage of the entire tongue epithelium area infected in different groups of mice infected with the different C. albicans strains. c, Confocal imaging of 4-day post-fertilization (dpf) mpo-gfp transgenic zebrafish swimbladders infected with C. albicans wild-type (BWP17 + CIp30 + dTomato), ECE1-deletion (ece1Δ/Δ + dTomato) and ECE1 re-integrant (ece1Δ/Δ + ECE1 + dTomato) strains for 24 h. C. albicans cells appear red while neutrophils appear green. Red dots outline the swimbladder. Images are composites of maximum projections in the red and green channels (25 slices each, approximately 100 μm depth) with (left) or without (right) a single slice in the DIC channel overlay. Scale bars represent 100 μm. d, Confocal imaging of 4 dpf zebrafish swimbladders infected with C. albicans wild-type (BWP17 + CIp30 + dTomato), ECE1-deletion (ece1Δ/Δ + dTomato) and ECE1 re-integrant (ece1Δ/Δ + ECE1 + dTomato) strains for 24 h stained with the fluorescent exclusion dye Sytox Green. C. albicans cells appear red and damaged epithelial cells appear green. White dots outline the pronephros and red dots outline the swimbladder. Images are composites of maximum projections in the red and green channels (25 slices each, approximately 100 μm depth) with (left) or without (right) a single slice in the DIC channel overlay. High magnification images of the white boxes are shown. Scale bars (bottom right) represent 100 μm (low magnification) and 30 μm (high magnification). Data shown are the mean (a, b) or representative (c, d) of at least three biological replicates. Error bars show ± s.e.m. Data were analysed by Mann–Whitney U-test. ***P < 0.001.
Extended Data Figure 3 Ece1-III62–93 is the active region of Ece1p.
a, Amino acid sequence of Ece1p and a schematic of the protein, indicating the signal peptide (SP), lysine-arginine motifs (KR) at the C terminus of each peptide, and the processed peptides (Ece1-I–VIII) produced by Kex2p cleavage. b, Amino acid sequences of the processed peptides (Ece1-I–VIII) produced by Kex2p cleavage. c–e, Induction of GM-CSF (c), IL-1α (d) and IL-6 (e) secreted after stimulation of TR146 epithelial cells for 24 h with varying concentrations of Ece1-III62–93 (1.5–70 μM). f, Phosphorylation of MKP-1 and c-Fos production after 2 h treatment of TR146 epithelial cells with 15 μM of Ece1-III62–85 (hydrophobic region), Ece1-III86–93 (hydrophillic region), Ece1-III62–85 and Ece1-III86–93 together, or Ece1-III62–93 alone. g, Induction of G-CSF secretion after 24 h treatment of TR146 epithelial cells with 15 μM of Ece1-III62–85, Ece1-III86–93, Ece1-III62–85 and Ece1-III86–93 together, or Ece1-III62–93 alone. h, Fold change induction of LDH release after 24 h treatment of TR146 epithelial cells with 70 μM of Ece1-III62-85, Ece1-III86-93, Ece1-III62–85 and Ece1-III86–93 together, or Ece1-III62–93 alone. i, Induction of p-MKP-1 and c-Fos 2 h post-infection (p.i.) with the indicated C. albicans strains (MOI = 10). j, c-Fos DNA binding induction 3 h p.i. with indicated C. albicans strains (MOI = 10). k, G-CSF secretion and l, LDH release 24 h p.i. with indicated C. albicans strains (MOI = 0.01). Data shown are representative (f, i) or the mean (c–e, g, h, j–l) of three biological replicates. Error bars show ± s.e.m. Data were analysed by one-way ANOVA (c–e, g, h, k–l) or t-test (j). *P < 0.05, **P< 0.01, ***P < 0.001 (compared with vehicle control). For gel image, see Supplementary Fig. 1.
Extended Data Figure 4 Ece1-III62–93 is required for C. albicans mucosal infection.
a, Fungal burdens recovered from the tongues of mice infected with C. albicans wild-type (BWP17 + CIp30) (number of mice (n) = 13), ECE1-deletion (ece1Δ/Δ) (n = 20), ECE1 re-integrant (ece1Δ/Δ + ECE1) (n = 24) and Ece1-III62–93 deletion (ece1Δ/Δ +ECE1Δ184–279) (n = 10) strains after 2-day oropharyngeal infection. b, Average percentage of the entire tongue epithelium area infected in different groups of mice infected with the different C. albicans strains. c, Confocal imaging of 4 dpf zebrafish swimbladders infected with C. albicans Ece1-III62–93 deletion (ece1Δ/Δ + ECE1Δ184–279 + dTomato) and ECE1 re-integrant (ece1Δ/Δ + ECE1 + dTomato) strains for 24 h stained with the fluorescent exclusion dye Sytox Green. C. albicans cells appear red and damaged cells appear green. White dots outline the pronephros and red dots outline the swimbladder. Images are composites of maximum projections in the red and green channels (25 slices each, approximately 100 μm depth) with (left) or without (right) a single slice in the DIC channel overlay. Scale bars (bottom right) represent 100 μm. Data shown are the mean (a, b) or representative (c), of at least three biological replicates. Error bars show ± s.e.m. Data were analysed by Mann–Whitney U-test. **P < 0.01, ***P < 0.001.
Extended Data Figure 5 Ece1-III62–93 is a cytolytic α-helical peptide.
a, Circular dichroism spectra showing the α-helical conformation of Ece1-III62–93 in buffer (100 mM KCl, 5 mM HEPES, pH 7). Increasing the temperature from 25 °C to 40 °C did not affect the stability of the α-helical structure. b, Diagram to illustrate the amphipathic nature of Ece1-III62–93 (residues 62–78, left panel; residues 79–93, right panel). Residues with hydrophobic or polar/charged side chains are displayed with a blue and white background, respectively. Modified from output generated in PEPWHEEL (http://emboss.bioinformatics.nl/cgi-bin/emboss/pepwheel). c, Förster resonance energy transfer (FRET) experiments show the intercalation of Ece1-III62–93 into lipid liposomes (10 μM) composed of DOPC in the absence or presence of cholesterol. Peptide titration of Ece1-III62–93 to liposomes showed slightly enhanced intercalation for pure DOPC. d, Ece1-III62–93 induced the permeabilization of planar lipid membranes composed of DOPC. The graph shows heterogeneous and transient lesions leading finally to a rupture of the membrane. Ece1-III62–93 concentration was 0.125 μM. e, Induction of p-MKP-1 and c-Fos 2 h in TR146 cells post stimulation (p.s.) with Ece1-III62–93KR or Ece1-III62–93AA. f, Secretion of G-CSF from TR146 cells 24 h p.s. with Ece1-III62–93KR or Ece1-III62–93AA. Data shown are representative (a–e) or mean (f) of at least three biological replicates. Error bars show ± s.e.m. Data were analysed by one-way ANOVA (f). *P < 0.05, **P < 0.01, ***P < 0.001 (compared with vehicle control). For gel images, see Supplementary Fig. 1.
Extended Data Figure 6 Schematic of the role of Ece1-III in C. albicans infection of epithelial cells.
a–h, During early stage infection of the mucosal surface by C. albicans, Ece1-III (red α-helix) is secreted into the invasion pocket created by the invading hypha (a), Sub-lytic concentrations of Ece1-III trigger epithelial signal transduction through MAPK, p38/MKP-1 and c-Fos (b), resulting in the production of immune regulatory cytokines (c), As the severity of the infection increases, Ece1-III accumulates (d), and once lytic concentrations are reached, causes membrane damage and the release of lactate dehydrogenase from the host epithelium (e), concomitant with calcium influx (f). Epithelial signal transduction is maintained (g), and additionally induces the release of damage associated cytokines, such as IL-1α (h). Ece1-III may also have activity on the epithelial surface outside of the invasion pocket and on neighbouring cells not in contact with hyphae if Ece1-III is produced in sufficient concentrations.
Supplementary information
Supplementary Figure 1
This file contains uncropped scans of the source gels with size markers indicated. (PDF 199 kb)
Supplementary Table
This file contains Supplementary Data. (XLSX 133 kb)
Rights and permissions
About this article
Cite this article
Moyes, D., Wilson, D., Richardson, J. et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 532, 64–68 (2016). https://doi.org/10.1038/nature17625
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17625
This article is cited by
-
The hyphal-specific toxin candidalysin promotes fungal gut commensalism
Nature (2024)
-
A comprehensive guide to assess gut mycobiome and its role in pathogenesis and treatment of inflammatory bowel disease
Indian Journal of Gastroenterology (2024)
-
Dehydrocostus lactone inhibits Candida albicans growth and biofilm formation
AMB Express (2023)
-
Core-predominant gut fungus Kazachstania slooffiae promotes intestinal epithelial glycolysis via lysine desuccinylation in pigs
Microbiome (2023)
-
Candida albicans exploits N-acetylglucosamine as a gut signal to establish the balance between commensalism and pathogenesis
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.