Abstract
Because autism spectrum disorders are neurodevelopmental disorders and patients typically display symptoms before the age of three1, one of the key questions in autism research is whether the pathology is reversible in adults. Here we investigate the developmental requirement of Shank3 in mice, a prominent monogenic autism gene that is estimated to contribute to approximately 1% of all autism spectrum disorder cases2,3,4,5,6. SHANK3 is a postsynaptic scaffold protein that regulates synaptic development, function and plasticity by orchestrating the assembly of postsynaptic density macromolecular signalling complex7,8,9. Disruptions of the Shank3 gene in mouse models have resulted in synaptic defects and autistic-like behaviours including anxiety, social interaction deficits, and repetitive behaviour10,11,12,13. We generated a novel Shank3 conditional knock-in mouse model, and show that re-expression of the Shank3 gene in adult mice led to improvements in synaptic protein composition, spine density and neural function in the striatum. We also provide behavioural evidence that certain behavioural abnormalities including social interaction deficit and repetitive grooming behaviour could be rescued, while anxiety and motor coordination deficit could not be recovered in adulthood. Together, these results reveal the profound effect of post-developmental activation of Shank3 expression on neural function, and demonstrate a certain degree of continued plasticity in the adult diseased brain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Amaral, D., Geschwind, D. & Dawson, G. Autism Spectrum Disorders 1st edn (Oxford Univ. Press, 2011)
Gauthier, J. et al. Novel de novo SHANK3 mutation in autistic patients. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 150B, 421–424 (2009)
El-Fishawy, P. & State, M. W. The genetics of autism: key issues, recent findings, and clinical implications. Psychiatr. Clin. North Am. 33, 83–105 (2010)
Leblond, C. S. et al. Meta-analysis of SHANK mutations in autism spectrum disorders: a gradient of severity in cognitive impairments. PLoS Genet. 10, e1004580 (2014)
Moessner, R. et al. Contribution of SHANK3 mutations to autism spectrum disorder. Am. J. Hum. Genet. 81, 1289–1297 (2007)
Durand, C. M. et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nature Genet. 39, 25–27 (2007)
Ebert, D. H. & Greenberg, M. E. Activity-dependent neuronal signalling and autism spectrum disorder. Nature 493, 327–337 (2013)
Kim, E. et al. GKAP, a novel synaptic protein that interacts with the guanylate kinase- like domain of the PSD-95/SAP90 family of channel clustering molecules. J. Cell Biol. 136, 669–678 (1997)
Takeuchi, M. et al. SAPAPs. A family of PSD-95/SAP90-associated proteins localized at postsynaptic density. J. Biol. Chem. 272, 11943–11951 (1997)
Wang, X. et al. Synaptic dysfunction and abnormal behaviors in mice lacking major isoforms of Shank3. Hum. Mol. Genet. 20, 3093–3108 (2011)
Bozdagi, O. et al. Haploinsufficiency of the autism-associated Shank3 gene leads to deficits in synaptic function, social interaction, and social communication. Mol. Autism 1, 15 (2010)
Peça, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011)
Yang, M. et al. Reduced excitatory neurotransmission and mild autism-relevant phenotypes in adolescent Shank3 null mutant mice. J. Neurosci. 32, 6525–6541 (2012)
Sheng, M. & Kim, E. The Shank family of scaffold proteins. J. Cell Sci. 113, 1851–1856 (2000)
Boeckers, T. M. et al. Proline-rich synapse-associated proteins ProSAP1 and ProSAP2 interact with synaptic proteins of the SAPAP/GKAP family. Biochem. Biophys. Res. Commun. 264, 247–252 (1999)
Han, K. et al. SHANK3 overexpression causes manic-like behaviour with unique pharmacogenetic properties. Nature 503, 72–77 (2013)
Schnütgen, F. et al. A directional strategy for monitoring Cre-mediated recombination at the cellular level in the mouse. Nature Biotechnol. 21, 562–565 (2003)
Karayannis, T. et al. Cntnap4 differentially contributes to GABAergic and dopaminergic synaptic transmission. Nature 511, 236–240 (2014)
Guo, C., Yang, W. & Lobe, C. G. A Cre recombinase transgene with mosaic, widespread tamoxifen-inducible action. Genesis 32, 8–18 (2002)
Rothwell, P. E. et al. Autism-associated neuroligin-3 mutations commonly impair striatal circuits to boost repetitive behaviors. Cell 158, 198–212 (2014)
Welch, J. M. et al. Cortico-striatal synaptic defects and OCD-like behaviours in Sapap3-mutant mice. Nature 448, 894–900 (2007)
Chao, H.-T. et al. Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 468, 263–269 (2010)
Dölen, G. et al. Social reward requires coordinated activity of nucleus accumbens oxytocin and serotonin. Nature 501, 179–184 (2013)
Gross, C. et al. Serotonin1A receptor acts during development to establish normal anxiety-like behaviour in the adult. Nature 416, 396–400 (2002)
De Zeeuw, C. I. & Ten Brinke, M. M. Motor learning and the cerebellum. Cold Spring Harb. Perspect. Biol. 7, a021683 (2015)
Poirier, M. C. & Schild, L. J. The genotoxicity of tamoxifen: extent and consequences, Kona, Hawaii, January 23, 2003. Mutagenesis 18, 395–399 (2003)
Guy, J., Gan, J., Selfridge, J., Cobb, S. & Bird, A. Reversal of neurological defects in a mouse model of Rett syndrome. Science 315, 1143–1147 (2007)
Clement, J. P. et al. Pathogenic SYNGAP1 mutations impair cognitive development by disrupting maturation of dendritic spine synapses. Cell 151, 709–723 (2012)
Hancock, J. F., Cadwallader, K., Paterson, H. & Marshall, C. J. A CAAX or a CAAL motif and a second signal are sufficient for plasma-membrane targeting of ras proteins. EMBO J. 10, 4033–4039 (1991)
Acknowledgements
We thank T. Dalia, A. Lim, S. Feng, K. Han, W. Stockton, H. Zaniewski and B. Clear for technical support. We thank Q. Zhang for designing the pAAV-hSYN1-EGFP-P2A-EGFPf-WPRE-HGHpA construct. We thank all members of the Feng laboratory for their support and discussions. Y.M. would like to thank T. Littleton, Y. Lin and K. Tye. P.M. would like to thank C. Duarte and the late S. Chaterjee, and acknowledge support from the ‘Programa Doutoral em Biologia Experimental e Biomedicina’ (CNC, Coimbra, Portugal). This work was funded by the National Science Foundation Graduate Fellowship and Integrative Neuronal Systems to Y.M.; the Stanley Center for Psychiatric Research at the Broad Institute of MIT and Harvard and a doctoral fellowship from the Portuguese Foundation for Science and Technology to P.M. (SFRH/BD/33894/2009). Y.Z. is supported by postdoc fellowships from the Simons Center for the Social Brain at MIT, Nancy Lurie Marks Family Foundation and Shenzhen Overseas Innovation Team Project (no. KQTD20140630180249366). X.G. was supported by the Stanley Center for Psychiatric Research at the Broad Institute of MIT and Harvard and a graduate fellowship from China Scholarship Council. Z.F. is supported by Stanley Center for Psychiatric Research at Broad Institute of MIT and Harvard and NARSAD Young Investigator Grant from the Brain & Behavior Research Foundation. Research in the Feng laboratory is supported by the Poitras Center for Affective Disorders Research at MIT, Stanley Center for Psychiatric Research at Broad Institute of MIT and Harvard, National Institute of Health (NIMH R01MH097104), Nancy Lurie Marks Family Foundation, Simons Foundation Autism Research Initiative (SFARI) and Simons Center for the Social Brain at MIT.
Author information
Authors and Affiliations
Contributions
Y.M., P.M. and G.F. designed the experiments and wrote the paper. Y.M., P.M., Y.Z., J.-A.K., X.G. and Z.F. performed the experiments and analysed the data. Y.M., P.M., Y.Z., J.-A.K., X.G. and Z.F. interpreted the results.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Generation of Shank3 conditional knock-in mice.
a, i–ii, Schematic of the Shank3 gene and the target locus. a, iii, Targeted exons 13–16. a, iv, Neo-cassette excision via breeding with germline Flp mice. a, v, Exons 13–16 re-inversion via Cre-expressing mice. b, PCR genotyping showing the bands for Fx (knockout), rescue and WT. c, Western blot showing rescue of Shank3 expression upon germline Cre recombination, with the exception of the putative SHANK3 γ isoform. This is probably due to the disruption of a putative intronic promoter by the introduction of the loxP sites. d, Tamoxifen-inducible Cre strategy leads to broad reporter expression. Sagittal sections from pCAGGS-CreER+/−:Rosa-floxstop-tdTomato+/− mice after feeding with tamoxifen (left) or corn oil (right). Results show widespread induction of tdTomato reporter expression after tamoxifen-induced Cre activation, but not in the absence of Cre activity (corn oil feeding); the pCAGGS promoter consists of the CMV early enhancer with chicken β-actin promoter19. See Supplementary Fig. 1 for gel source data.
Extended Data Figure 2 Additional measurements of dorsal striatum synaptic function in TM rescue.
a, Representative traces (left) and bar graph (right) for the AMPAR/NMDAR ratio in WT, KO and TM groups (n = 30 WT, n = 33 KO and n = 33 TM MSNs). AMPAR/NMDAR ratio calculated as the ratio of the EPSC peak amplitude at −70 mV (AMPAR EPSC) to the amplitude of the EPSC recorded at +40 mV, 50 ms after afferent stimulation. b, Representative traces (left) and bar graph (right) for pharmacologically isolated AMPAR/NMDAR ratio (n = 20 WT, n = 20 KO and n = 23 TM MSNs). Dual-component evoked EPSC at +40 mV recorded before and after AP5 bath application. c, Representative traces (left) and bar graph (right) for the NR2B/NMDAR ratio in WT, KO and TM groups (n = 16 WT, n = 16 KO and n = 22 TM MSNs). Dual-component evoked EPSC at +40 mV recorded before and after ifenprodil bath application. N.S., not significant (Kruskal–Wallis test, with Dunn’s multiple comparison test (a), one-way ANOVA with Bonferroni post-hoc test (b, c)). Data are mean ± s.e.m.
Extended Data Figure 3 Additional measurements of synaptic function in dorsal striatum by cortical and corpus callosum evoked stimulation.
a, IR-DIC images showing representative placement of the stimulation electrode in the cortex (left) or corpus callosum (cc; right) to evoke EPSCs in dorsal striatum, using parasagittal (left) and coronal (right) slices. b, Representative traces (right) and summary bar graphs for the paired-pulse ratios (PPR) of evoked EPSCs in dorsal striatum MSNs, showing similar magnitude in parasagittal (left; n = 23 WT , n = 20 KO and n = 20 TM MSNs) and coronal (right; n = 9 WT, n = 11 KO and n = 10 TM MSNs) slices. One-way ANOVA with Bonferroni post-hoc test. c, Representative traces (right) and summary bar graphs of AMPAR/NMDAR ratios evoked in parasagittal (left; n = 17 WT, n = 15 KO and n = 16 TM MSNs) and coronal (right; n = 20 WT, n = 16 KO and n = 15 TM MSNs) slices from all three genotypes. AMPAR/NMDAR ratio calculated as the ratio of the EPSC peak amplitude at −70 mV (AMPAR EPSC) to the amplitude of the EPSC recorded at +40 mV, 50 ms after afferent stimulation (c, right). Kruskal–Wallis test, with Dunn’s multiple comparison test. Data are mean ± s.e.m.
Extended Data Figure 4 Whole-cell measurements of excitatory synaptic function in nucleus accumbens.
a, Representative traces (top) and summary bar graphs (bottom) of mEPSCs in nucleus accumbens (NAc) MSNs (n = 22 WT, n = 16 KO and n = 18 GR MSNs). b, Representative traces (top) and summary bar graph (bottom) of the AMPAR/NMDAR ratio in NAc MSNs. The AMPAR/NMDAR ratio was calculated as the ratio of the EPSC peak amplitude at −70 mV (AMPAR EPSC) to the amplitude of the EPSC recorded at +40 mV, 50 ms after afferent stimulation (n = 19 WT, n = 16 KO and n = 15 GR MSNs). c, Representative traces (top) and summary bar graph (bottom) of paired-pulse ratios in NAc MSNs (n = 25 WT, n = 22 KO and n = 19 GR MSNs). One-way ANOVA Bonferroni post-hoc test (a, c), Kruskal–Wallis test, with Dunn’s multiple comparison test (b). Data are mean ± s.e.m.
Extended Data Figure 5 Western blots of synaptosomal preparations from the cortex of the adult treated mice show minimal difference across genotypes.
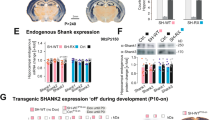
a, Representative western blot of SHANK3 in the cortex of adult WT, KO and TM mice, showing that most major SHANK3 isoforms are restored in the TM mice. b, Representative western blots of synaptic proteins including scaffolding proteins and neurotransmitter receptors in the adult cortex across genotypes. c, Quantification of multiple synaptic proteins in the cortex of the adult treated mice. All data, n = 3 WT, n = 3 TM and n = 3 KO mice; each sample is from tissue taken from two animals. Student’s two-tailed unpaired t-test. Data are mean ± s.e.m. See Supplementary Fig. 1 for gel source data.
Extended Data Figure 6 Western blots of synaptosomal preparations from the cerebellum of the adult treated mice show minimal difference across genotypes.
a, Representative western blot of SHANK3 in the cerebellum of the adult mice treated with TM, showing that SHANK3 isoforms are restored in the TM mice. b, Representative western blots of synaptic proteins in the adult cerebellum across genotypes. c, Quantification of multiple synaptic proteins in the cerebellum of the adult treated mice. All data, n = 3 WT, n = 3 TM and n = 3 KO mice; each sample is from tissue taken from a single animal. Student’s two-tailed unpaired t-test. Data are as mean ± s.e.m. See Supplementary Fig. 1 for gel source data.
Extended Data Figure 7 Whole-cell measurements of excitatory synaptic function in the cortex.
a, b, Summary bar graphs of mEPSC frequency (a) and amplitude (b) in the prefrontal cortex of the adult WT and KO mice (n = 13 WT and n = 10 KO cells; Student’s two-tailed unpaired t-test). Data are mean ± s.e.m. c, Representative traces from mEPSC recordings in the adult prefrontal cortex of the WT and KO mice.
Extended Data Figure 8 Electrophysiological measurements in the dorsal striatum of germline rescue mice.
a, b, Normal pop spike amplitude (a) and NP1 (b) in germline rescued mice (insets show representative field traces). c, d, Reduced mEPSC frequency in KO compared to WT and GR mice (c). A reduction in mEPSC peak current amplitude is observed between KO and WT group only (d) (n = 30 WT, n = 24 KO and n = 26 GR MSNs). e, Representative traces (left) and bar graph (right) for the pharmacologically isolated AMPAR/NMDAR ratio (n = 9 WT, n = 9 KO and n = 8 GR MSNs). Dual-component evoked EPSC at +40 mV recorded before and after DL-AP5 bath application. f, Representative traces (left) and bar graph (right) for NR2B/NMDAR ratio in WT, KO and GR groups (n = 6 WT, n = 5 KO and n = 7 GR MSNs). Dual-component evoked EPSC at +40 mV recorded before and after ifenprodil bath application. *P < 0.05; **P < 0.01; ***P < 0.001 (two-way (a, b) and one-way (c–f) ANOVA with Bonferroni post-hoc test). Data are mean ± s.e.m.
Extended Data Figure 9 Expression of SHANK3 at P20–P21 rescues some behavioural measurements.
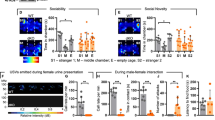
a, Representative western blots showing efficient SHANK3 re-expression in the cortex, striatum and cerebellum in mice that were treated with tamoxifen at P20–P21. b, The total distance travelled as measured by the open-field test was not improved in the TM condition compared to the KO condition. One-way ANOVA with Bonferroni post-hoc test. c, The open-field total distance plotted across 5-min time bins, showing that there is no difference between KO and TM conditions across time. d, Rearing activity measured by open-field plotted across time, showing that the TM mice perform in between the WT and KO mice for most of the 30-min test. e, Rearing time measured by open-field test plotted across time, also showing the intermediate performance of TM mice between that of WT and KO mice (b–e: n = 21 WT, n = 36 KO and n = 30 TM mice; two-way ANOVA with Bonferroni post-hoc test). f, Activity on the zero maze indicates that the TM condition shows significantly reduced anxiety compared to that of the KO condition (n = 18 WT, n = 25 KO and n = 30 TM mice; outliers were removed using Iglewicz and Hoaglin’s test (two-sided); Kruskal–Wallis test, Dunn’s multiple comparison test). Data are mean ± s.e.m. See Supplementary Fig. 1 for gel source data.
Extended Data Figure 10 Expression of SHANK3 at P20–P21 improves motor coordination.
a, Summary data from trial 1 of the accelerating rotarod test on mice treated with TM at P20–P21 (n = 19 WT, n = 35 KO and n = 33 TM mice). b, Summary data from trial 2 of the same rotarod test on mice treated at P20–P21 (n = 18 WT, n = 33 KO and n = 33 TM mice). c, Summary data from trial 3 of the rotarod on mice treated at P20–P21 (n = 20 WT, n = 36 KO and n = 33 TM mice). Outliers in a–c were removed using Iglewicz and Hoaglin’s robust outlier test. One-way ANOVA with Bonferroni post-hoc test. d, Body weight from WT and KO mice that were unfed (UF) and mice that were fed with either corn oil or TM (n = 28 WT (UF), n = 14 WT (TM), n = 27 KO (UF), n = 25 KO (CO) and n = 26 KO (TM)). One-way ANOVA with Bonferroni post-hoc test. Data are mean ± s.e.m.
Supplementary information
Supplementary Information
This file contains full blots for all western blots used in the Main and Extended Data Figures. (PDF 1156 kb)
Rights and permissions
About this article
Cite this article
Mei, Y., Monteiro, P., Zhou, Y. et al. Adult restoration of Shank3 expression rescues selective autistic-like phenotypes. Nature 530, 481–484 (2016). https://doi.org/10.1038/nature16971
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16971
This article is cited by
-
Multi-animal 3D social pose estimation, identification and behaviour embedding with a few-shot learning framework
Nature Machine Intelligence (2024)
-
Hyperthermia elevates brain temperature and improves behavioural signs in animal models of autism spectrum disorder
Molecular Autism (2023)
-
The Sapap3−/− mouse reconsidered as a comorbid model expressing a spectrum of pathological repetitive behaviours
Translational Psychiatry (2023)
-
Disrupted extracellular matrix and cell cycle genes in autism-associated Shank3 deficiency are targeted by lithium
Molecular Psychiatry (2023)
-
Deficits in integrative NMDA receptors caused by Grin1 disruption can be rescued in adulthood
Neuropsychopharmacology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.