Abstract
The development of targeted anti-cancer therapies through the study of cancer genomes is intended to increase survival rates and decrease treatment-related toxicity. We treated a transposon–driven, functional genomic mouse model of medulloblastoma with ‘humanized’ in vivo therapy (microneurosurgical tumour resection followed by multi-fractionated, image-guided radiotherapy). Genetic events in recurrent murine medulloblastoma exhibit a very poor overlap with those in matched murine diagnostic samples (<5%). Whole-genome sequencing of 33 pairs of human diagnostic and post-therapy medulloblastomas demonstrated substantial genetic divergence of the dominant clone after therapy (<12% diagnostic events were retained at recurrence). In both mice and humans, the dominant clone at recurrence arose through clonal selection of a pre-existing minor clone present at diagnosis. Targeted therapy is unlikely to be effective in the absence of the target, therefore our results offer a simple, proximal, and remediable explanation for the failure of prior clinical trials of targeted therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hovestadt, V. et al. Decoding the regulatory landscape of medulloblastoma using DNA methylation sequencing. Nature 510, 537–541 (2014)
Jones, D. T. et al. Dissecting the genomic complexity underlying medulloblastoma. Nature 488, 100–105 (2012)
Kool, M. et al. Genome sequencing of SHH medulloblastoma predicts genotype-related response to smoothened inhibition. Cancer Cell 25, 393–405 (2014)
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014)
Northcott, P. A. et al. Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 488, 49–56 (2012)
Pugh, T. J. et al. Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations. Nature 488, 106–110 (2012)
Wu, X. et al. Clonal selection drives genetic divergence of metastatic medulloblastoma. Nature 482, 529–533 (2012)
Moxon-Emre, I. et al. Impact of craniospinal dose, boost volume, and neurologic complications on intellectual outcome in patients with medulloblastoma. J. Clin. Oncol. 32, 1760–1768 (2014)
Northcott, P. A., Korshunov, A., Pfister, S. M. & Taylor, M. D. The clinical implications of medulloblastoma subgroups. Nature Rev. Neurol. 8, 340–351 (2012)
Northcott, P. A. et al. Medulloblastomics: the end of the beginning. Nature Rev. Cancer 12, 818–834 (2012)
Ramaswamy, V. et al. Recurrence patterns across medulloblastoma subgroups: an integrated clinical and molecular analysis. Lancet Oncol. 14, 1200–1207 (2013)
Anderson, K. et al. Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature 469, 356–361 (2011)
Burrell, R. A., McGranahan, N., Bartek, J. & Swanton, C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 501, 338–345 (2013)
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012)
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012)
Landau, D. A. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, 714–726 (2013)
Mullighan, C. G. et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 322, 1377–1380 (2008)
Johnson, B. E. et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343, 189–193 (2014)
Shah, S. P. et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 461, 809–813 (2009)
Gerlinger, M. & Swanton, C. How Darwinian models inform therapeutic failure initiated by clonal heterogeneity in cancer medicine. Br. J. Cancer 103, 1139–1143 (2010)
Eirew, P. et al. Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature 518, 422–426 (2015)
Kreso, A. et al. Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 339, 543–548 (2013)
Notta, F. et al. Evolution of human BCR–ABL1 lymphoblastic leukaemia-initiating cells. Nature 469, 362–367 (2011)
Mumert, M. et al. Functional genomics identifies drivers of medulloblastoma dissemination. Cancer Res. 72, 4944–4953 (2012)
Brett, B. T. et al. Novel molecular and computational methods improve the accuracy of insertion site analysis in Sleeping Beauty-induced tumors. PLoS ONE 6, e24668 (2011)
Collier, L. S., Carlson, C. M., Ravimohan, S., Dupuy, A. J. & Largaespada, D. A. Cancer gene discovery in solid tumours using transposon-based somatic mutagenesis in the mouse. Nature 436, 272–276 (2005)
Dupuy, A. J., Akagi, K., Largaespada, D. A., Copeland, N. G. & Jenkins, N. A. Mammalian mutagenesis using a highly mobile somatic Sleeping Beauty transposon system. Nature 436, 221–226 (2005)
Zhukova, N. et al. Subgroup-specific prognostic implications of TP53 mutation in medulloblastoma. J. Clin. Oncol. 31, 2927–2935 (2013)
Zhu, S. et al. The bHLH repressor Deadpan regulates the self-renewal and specification of Drosophila larval neural stem cells independently of Notch. PLoS ONE 7, e46724 (2012)
Ollmann, M. et al. Drosophila p53 is a structural and functional homolog of the tumor suppressor p53. Cell 101, 91–101 (2000)
Fraley, C., Raftery, E. A., Murphy, T. B. & Scrucca, L. mclust Version 4 for R: Normal Mixture Modeling for Model-Based Clustering, Classification, and Density Estimation. Technical Report No. 597 (2012)
Remke, M. et al. TERT promoter mutations are highly recurrent in SHH subgroup medulloblastoma. Acta Neuropathol. 126, 917–929 (2013)
Govind, S. K. et al. ShatterProof: operational detection and quantification of chromothripsis. BMC Bioinformatics 15, 78 (2014)
Griffith, M. et al. DGIdb: mining the druggable genome. Nature Methods 10, 1209–1210 (2013)
McGranahan, N. & Swanton, C. Perspective biological and therapeutic impact of intratumor heterogeneity in cancer evolution. Cancer Cell 27, 15–26 (2015)
Andor, N., Harness, J. V., Müller, S., Mewes, H. W. & Petritsch, C. EXPANDS: expanding ploidy and allele frequency on nested subpopulations. Bioinformatics 30, 50–60 (2014)
Maley, C. C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nature Genet. 38, 468–473 (2006)
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nature Methods 11, 396–398 (2014)
Hill, R. M. et al. Combined Myc and p53 defects emerge at medulloblastoma relapse and define rapidly progressive, therapeutically targetable disease. Cancer Cell 27, 72–84 (2015)
Wang, X. et al. Medulloblastoma subgroups remain stable across primary and metastatic compartments. Acta Neuropathol. 129, 449–457 (2015)
Aparicio, S. & Caldas, C. The implications of clonal genome evolution for cancer medicine. N. Engl. J. Med. 368, 842–851 (2013)
Maley, C. C., Reid, B. J. & Forrest, S. Cancer prevention strategies that address the evolutionary dynamics of neoplastic cells: simulating benign cell boosters and selection for chemosensitivity. Cancer Epidemiol. Biomarkers Prev. 13, 1375–1384 (2004)
Clarkson, R. et al. Characterization of image quality and image-guidance performance of a preclinical microirradiator. Med. Phys. 38, 845–856 (2011)
Ma, C. M. et al. AAPM protocol for 40–300 kV X-ray beam dosimetry in radiotherapy and radiobiology. Med. Phys. 28, 868–893 (2001)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Cingolani, P. et al. Using Drosophila melanogaster as a model for genotoxic chemical mutational studies with a new program, SnpSift. Front. Genet. 3, http://dx.doi.org/10.3389/fgene.2012.00035 (2012)
Saunders, C. T. et al. Strelka: Accurate somatic small-variant calling from sequenced tumour-normal sample pairs. Bioinformatics 28, 1811–1817 (2012)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Fischer, A., Illingworth, C. J., Campbell, P. J. & Mustonen, V. EMu: probabilistic inference of mutational processes and their localization in the cancer genome. Genome Biol. 14, R39 (2013)
Jones, S. J. et al. Evolution of an adenocarcinoma in response to selection by targeted kinase inhibitors. Genome Biol. 11, R82 (2010)
Shah, S. P. et al. Integrating copy number polymorphisms into array CGH analysis using a robust HMM. Bioinformatics 22, 431–439 (2006)
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425 (2012)
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386 (2000)
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nature Biotechnol. 31, 213–219 (2013)
Vanner, R. J. et al. Quiescent Sox2+ cells drive hierarchical growth and relapse in sonic hedgehog subgroup medulloblastoma. Cancer Cell 26, 33–47 (2014)
Shih, D. J. H. et al. Cytogenetic prognostication within medulloblastoma subgroups. J. Clin. Oncol. 32, 886–896 (2014)
Cho, Y.-J. et al. Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J. Clin. Oncol. 29, 1424–1430 (2011)
Butterfield, Y. S. et al. JAGuaR: junction alignments to genome for RNA-seq reads. PLoS ONE 9, e102398 (2014)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Acknowledgements
The MAGIC project is financially supported by: Genome Canada, Genome BC, Terry Fox Research Institute, Ontario Institute for Cancer Research, Pediatric Oncology Group Ontario, Funds from ‘The Family of Kathleen Lorette’ and the Clark H. Smith Brain Tumour Centre, Montreal Children’s Hospital Foundation, Hospital for Sick Children: Sonia and Arthur Labatt Brain Tumour Research Centre, Chief of Research Fund, Cancer Genetics Program, Garron Family Cancer Centre, B.R.A.I.N. Child, and BC Childhood Cancer Parents Association. M.D.T. is also supported by a Stand Up To Cancer St. Baldrick’s Pediatric Dream Team Translational Research Grant (SU2C-AACR-DT1113). Stand Up To Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research. M.D.T. is supported by The Canadian Cancer Society Research Institute, The Garron Family Chair in Childhood Cancer Research, and grants from the Cure Search for Children’s Cancer Foundation, the National Institutes of Health (R01CA148699 R01CA159859), The Pediatric Brain Tumour Foundation, The Terry Fox Research Institute, Brainchild and The McLaughlin Centre at the University of Toronto. M.D.T. is also supported by the Swifty Foundation. L.G. was supported by the Davis M. Ferguson Memorial Fund at ABTA. Alex’s Lemonade Stand Young Investigator Award supported V.R. This study was conducted with the support of the Ontario Institute for Cancer Research through funding provided by the Government of Ontario. This work was also supported by a Program Project Grant from the Terry Fox Research Institute, and a Grand Challenge Award from CureSearch for Children’s Cancer. Additionally, this work was supported by the PedBrain Tumour Project contributing to the International Cancer Genome Consortium, funded by German Cancer Aid (109252) and by the German Federal Ministry of Education and Research (BMBF, grants 01KU1201A, MedSys 0315416C and NGFNplus 01GS0883). Funding by the German Childhood Cancer Foundation (Deutsche Kinderkrebsstiftung) to S.M.P., G.F. and T.P. The study was also financed by the Hungarian Brain Research Program Grant No. KTIA_13_NAP-A-V/3. and NAP-A-II/7. A.K. was supported by the János Bolyai scholarship of the Hungarian Academy of Sciences. E.G.V.M. was supported by NIH R01 grants CA163722 and NS096236, and St. Baldrick’s and Cure Childhood Cancer Foundations. We would like to acknowledge R. P. Hill (Ontario Cancer Institute), the Labatt Brain Tumour Research Centre Tumour and Tissue Repository, which is supported by B.R.A.I.N. Child and Megan’s Walk. M.R. is supported by a fellowship from the Dr. Mildred Scheel Foundation for Cancer Research/German Cancer Aid. F.M.G.C. is supported by the Stephen Buttrum Brain Tumour Research Fellowship, granted by Brain Tumour Foundation of Canada. V.R. is supported by a CIHR fellowship and an Alberta Innovates-Health Solutions Clinical Fellowship. We would like to thank the Toronto Centre for Phenogenomics for animal housing and veterinary support, and the Preclinical Core II and animal research facility at STTARR (Spatiotemporal Targeting and Amplification of Radiation Response) in Toronto for assistance with CT-guided radiation experiments. We would like to thank Z. Wang for technical help with IHC, S. Archer for technical writing and C. Smith for artwork.
Author information
Authors and Affiliations
Contributions
A.S.M., L.G., and M.D.T. led the study. L.G. planned and carried out in vivo and in vitro experiments and analyses, and performed a subset of bioinformatic analyses. A.S.M. supervised the RNA-seq and WGS experiments, led and executed bioinformatic analyses. D.J.H.S. performed bioinformatics analysis of mutation signatures. S.Z. developed and implemented the computational method of finding initiating events in mouse tumours. X.H. developed the Drosophila brain tumour model and performed imaging of Drosophila brains. P.S. assisted with mouse library preparation and bioinformatics analysis. M.R. and V.R. performed bioinformatics analyses on DYNC1H1 and 14q loss. F.M.G.C. generated visualizations of structural rearrangements. P.E.L. and S.J. developed the radiotherapy schedule for the mouse model and designed the custom made collimators, beds, and stages for mouse CSI. K.Z. assisted with library preparation. B.L. extracted nucleic acids, managed the biobanking, and maintained the patient database. N.T., Y.M., and K.L.M. supervised bioinformatics analyses at the Genome Sciences Center, including sequence alignment, copy number analysis, and SNV and structural variant calling. Y.L., C.M. and E.M. performed bioinformatics analysis of human sequencing and deep-sequencing data. K.T. and T.Z. supervised and implemented the targeted deep-sequencing work. K.S. performed PyClone analysis. A.J.L.R. and S.S. designed and implemented PyClone, and supervised its use. H.F., S.M-L., J.R., and T.P. assisted with bioinformatic analyses. J.L., and L.Q., assisted with animal care, and N.K., B.L.H., J.J.Y.L., L.K.D., Xin W., S.C.M., A.M., K.A.M., C.N., John P., A.R., and Y.Y.T. provided technical support. Xiaochong W. generated the transgenic mouse model and offered technical advice. A.A., M.B., Y.S.N.B., R.C., Y.C., E.C., R.C., N.D., A.H., D.L., H.I.L., W.L., M.M., P.P., J.Q.Q., J.E.S., A.T., T.W., I.B., and Y.Z., led and performed RNA-seq and WGS library preparation and sequencing experiments and performed data analyses. A.K., D.T.W.J., M.K., P.A.N., and S.M.P. at DKFZ performed the sequencing of four patients’ sets. C.C.F., José P., S.N., T.S., M.G., I.F.P., R.L.H., X.-N.Li., A.E.B., D.W.F., A.W.W., T.K., T.T., V.P.C., Y.-J.C., C.H., D.L., J.H.W., J.H.G. Jr, D.S.S., L.M., U.S., J.S., K.Z., S.P., O.A., S.E.D., D.P.C.T., C.G.C., H.W., A.R.H., W.I., T.J.M., J.J.O., E.G.V.M., J.-Y.L., K.-C.W., S.-K.K., B.-K.C., Y.S.R., S.B., J.C.L., S.C.C., C.G.E., M.K.C., R.J.P., M.M., M.L.G., N.J., and S.M.P. obtained the patient samples and clinical details that made the study possible. T.P., G.F., S.T., U.B., U.T., C.E.H., P.D., E.B., J.T.R., R.J.W.-R., W.A.W., L.S.C., A.J.D., A.K., D.T.W.J., M.K., P.A.N., S.M.P., D.A.L., A.J.M., R.A.M., N.J., G.D.B., S.J.M.J., and D.M. provided valuable input regarding study design, data analysis, and interpretation of results. A.S.M., L.G., M.R., S.Z., G.D.B., M.A.M. and M.D.T. wrote the manuscript. M.A.M. and M.D.T. provided financial and technical infrastructure and oversaw the study. M.A.M. and M.D.T. are joint senior authors and project co-leaders.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Microneurosurgical resection and CT guided multi-fractionated craniospinal radiotherapy in a Shh mousemodel of medulloblastoma.
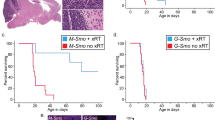
a, Under general anaesthesia, Ptch+/−/Math1-SB11/T2Onc mice with symptomatic medulloblastoma underwent microneurosurgical posterior fossa craniotomy and subtotal tumour removal (n = 38), followed by post-operative care and monitoring. b, Subsequently, post-operative mice are recurrently anaesthetized, and receive multi-fractionated cranial and spinal cord irradiation in 18 fractions for a total of 36 Gy over a period of four weeks. Radiation is delivered under computed tomography (CT) guidance using custom-made mouse beds and collimators in order to precisely target the entire craniospinal axis. c, Mice that completed the entire course of craniospinal radiation were cured of disease in 39% of cases (7/18), while the remainder had to be euthanized as they recurred locally and/or with leptomeningeal metastases (61%, 11/18). Histology (haematoxylin and eosin staining) at the time of autopsy is shown. d, Extent of overlap of primary, local recurrences and metastatic recurrences initiator genes as predicted by a per-mouse driver modelling approach. e, Clonal transposon insertions in Trp53, Tcf4 and Arid1b disrupt the coding sequence of the gene. Sense orientation insertions are illustrated in green, antisense insertions in red. f, Insertion-site end-point PCR demonstrates Trp53 insertions that are clonal in the recurrence, but present only in a subclone of the matched primary tumour or completely absent. Three levels of input DNA were used for each sample (1×, 5× and 25×). g, Mice treated with microneurosurgical resection and craniospinal radiation, whose tumours show Trp53 gCIS insertions in the local recurrence show a trend for a shorter survival than similarly treated mice without Trp53 insertions (log-rank test; P = 0.054; n = 10). h, Drosophila brain tumours are induced by expressing dpn in the neural stem cell lineage using insc-Gal4. In response to a systemic 40 Gy irradiation at late third instar stage, overexpressing a dominant negative form of Drosophila p53, p53R159N, resulted in moderately increased mitosis in tumour cells labelled by the membrane GFP (mCD8–GFP), scale bar, 50 μm.
Extended Data Figure 2 Subclonal events in primary mouse tumours become clonal at recurrence.
a, Naive tumours from Ptch+/−, Ptch+/−/Trp53+/− or Ptch+/−/Trp53−/− germline mutant mice were analysed by immunohistochemical staining for nuclear p21 (upper panels), demonstrating decreased nuclear p21 expression due to Trp53 pathway dysfunction. Tumours with Trp53 damaging gCIS insertions at recurrence (03-04-11 and 06-28-11) also show decreased immunohistochemical staining for nuclear p21 staining (lower panels), when compared to a recurrent tumour without gCIS Trp53 insertions (02-23-11w) (scale bars, 25 μm and 50 μm as indicated). b, Relative dominance of driver events is shown in one individual tumour where Tead1 is detectable in both primary tumour sample and at recurrence. c, d, Clonal insertions in the local and metastatic recurrences that were found at a subclonal level in the matching primary tumours are shown by mouse. In each case, the number of insertions with evidence for expansion from a subclone of the primary is shown as a proportion (red bar) of the total number of considered events. Green and blue bars depict the proportion of the total number of considered events that were found in local and metastatic recurrences, respectively. The grey bar indicates the proportion of insertions that are also found in an unrelated Sleeping Beauty library of similar depth. We narrowed the analysis on matching primary recurrences with at least 1 clonal insertion in common. This excluded 3 local recurrence cases and 6 metastatic recurrence cases that had no overlap between clonal insertions in the primary and clonal insertions in the matching local recurrences (black stars). c, Local recurrences display statistical support for subclonal derivation from the primary tumours (P = 0.041; n = 7; Mann–Whitney U-test). d, Metastatic recurrences instead show a limited extent of overlap with the matching primaries that does not reach statistical significance (P = 0.298; n = 5; Mann–Whitney U-test). e, Box plot comparing the extent of overlap between primary/local recurrences versus primary/metastatic recurrences, local recurrences (with at least 1 clonal insertion in common with the primary) show a trend for higher evidence of subclonal derivation from their matched primaries than metastatic recurrences (P = 0.051, Mann–Whitney U-test, centre lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles, samples are represented by dots. n = 7 and 5 sample points).
Extended Data Figure 3 Subclonal events in primary human tumours become clonal at recurrence.
The proportion of somatic SNVs in the primary and recurrent disease compartments of 15 patients with matched germline is shown as a function of clonality. Black indicates homozygous events, purple indicates clonal SNVs, and subclonal SNVs are shown in green, where lighter shades correspond to less abundant subpopulations. On average, we observe a 1.9-fold increase in the proportion of clonal and homozygous events across the cohort (Student’s t-test; P value = 8.7 × 10−3, n = 15).
Extended Data Figure 4 Altered spectra of somatic SNVs when comparing therapy-naive to recurrent tumours.
a, Mutations in each tumour sample (n = 15) were classified based on their sequence context, and clustered into signatures that represent four known mutational processes. Signature A is the age-related signature observed in most tumour types (deamination of methyl-C). Signature B is characterized by C > A and C > T mutations without a strict context requirement. Signatures C and D respectively resemble the MSI-L and MSI-H signatures that correlate with low (MSI-L) or high (MSI-H) microsatellite instability. b, The contribution of each mutational process to each primary and recurrent tumour is summarized by patient. Recurrent tumours show a shift away from signature A, and an increased prevalence of signature B and signature D. ***P < 0.001, chi-squared test denotes significantly different distributions; NS denotes not significant. All tumours shifted mutational signatures at the time of recurrence, for a and b, n = 15. c, d, The number c, and frequency d, of transversion mutations is summarized in therapy-naive and recurrent samples. Significant increases in the number and frequency of transversions is most strongly observed in local recurrences, and to a lesser extent in metastatic recurrences. P < 0.05, Wilcoxon rank-sum test. e, Breakdown of transversion (Tv) and transition (Ts) mutations in therapy-naive and recurrent samples does not show a significant trend in specific nucleotide changes. Centre lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend up to 1.5 times the interquartile range (from the 25th to the 75th percentiles), and data points beyond the whiskers are outliers represented by dots. For c, d and e, n = 13, n = 7 and n = 6, respectively.
Extended Data Figure 5 Compartment-specific driver and druggable events in human tumours.
a, High-level TERT amplification in the primary tumour of patient MB-REC-14 is absent in the recurrent sample. b, Chromothripsis involving the MYC locus is specific to the recurrent tumour on patient MB-REC-09 (P value = 3.97 × 10−7). c, Genes with defined interactions to neoplastic drugs (DGIdb http://dgidb.genome.wustl.edu/). The majority of patients (n = 15; with matched or parental germline) have distinct druggable targets in the naive versus post-therapy tumour samples. Bolded gene names indicate the presence of damaging mutations that are clonal (versus subclonal events in lighter colours), underlined gene names indicate copy number aberrations (for example, loss at the TP53 locus), and italicized gene names indicate structural rearrangements.
Extended Data Figure 6 Clonal lineage evolution post-therapy in human tumours.
a, Subpopulations of cells in each primary and recurrent tumour were identified using the EXPANDS algorithm, based on somatic SNVs and copy number gains and losses in each sample. Each subpopulation is thus distinguished by (1) a unique combination of somatic aberrations, which are (2) present in a particular subset of cells. Phylogenetic relationships between the primary (lowercase red letter labels) and recurrent (uppercase blue letter labels) tumour subpopulations indicate that in a majority of cases the recurrent tumour lineages are derived from only one lineage in the primary tumour, while only a small proportion of recurrent tumours had a more intermediate similarity to the primary tumour. b, The Shannon Index (SI) of each tumour is calculated using the cellular prevalence of the subpopulations defined by EXPANDS. Increasing values between the primary versus recurrent compartments indicate an increase in tumour heterogeneity (two tailed, paired t-test; P value = 0.029, Black lines show the medians; white lines represent individual data points; polygons represent the estimated density of the data). c, Clonal evolution between therapy-naive and matched recurrent tumours was assayed through ultra-deep sequencing (>1,500×) of somatic mutations, and analysed using PyClone. Cellular frequencies of clones (y axis) are scaled by the number of mutations in each clone. Ancestral high-frequency clones present in both compartments indicate a common cell of origin in every case. Lower-frequency mutation clusters in the primary tumour indicate clones that subsequently expand to dominance in the recurrent tumour (blue lines). Higher frequency clusters in the primary tumour that are absent or extremely subclonal at the time of recurrence (red lines) indicate therapy-sensitive clones. The number of mutations studied that support each type of event are indicated in the inset box.
Extended Data Figure 7 Subclonal expansion of rare (<5%) SNVs in the primary tumour to clonal dominance in the recurrent compartment.
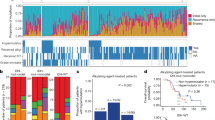
a, Deep amplicon sequencing was used to profile 20 patients with clonal SNVs restricted to their recurrent tumours as determined by 30× WGS data. Many ‘recurrence specific’ SNVs (blue) were found in a very minor subclone (<5%) of the primary tumour (red) when studied by deep amplicon sequencing. Clonal SNVs (allele frequency >15%) in recurrent tumours that had >1 read supporting an alternate base in the primary tumour are shown by patient. In each case, the number of events with evidence for expansion from a clone present at <5% is shown as a proportion (red bar) of the total number of considered events (blue bar). b, Evidence for clonal expansion at recurrence of clones present at <5% in the untreated tumour was observed in 16/20 patients, indicating that clonal selection is common after therapy for medulloblastoma. The extent of clonal selection (blue > red) varies across medulloblastoma cases, with prominent clonal selection in some cases (MB-REC-30), and more extreme divergence in others (MB-REC-23). c, d, Deep amplicon sequencing of clonal SNVs from both a first recurrence (dark blue), and a subsequent second recurrence (light blue) of patient MB-REC-31 reveals that clonal SNVs present at recurrence but absent from a 30× WGS profile of the untreated tumour (red) were indeed present at very low prevalence (~1/1,000) in the primary sample, indicating striking clonal expansion after initial treatment of the untreated tumour (c; AF, allele frequency; NA, not available). This is illustrated in panel d, which depicts the allelic frequency of a very low-prevalence PIK3CA mutation in the primary tumour that reaches clonal levels post-therapy.
Extended Data Figure 8 Base quality assessment of reference and alternate alleles at SNVs with clonal or rare allelic frequencies.
a, b, To determine whether low-frequency (<5%) base calls were SNVs or sequencing errors, we analysed the distribution of base quality (baseQ) values for each alternate base called. This plot shows the allele frequencies (AF; secondary y axis) and the proportion of supporting reads with baseQ values >30 (primary y axis) for a subset of SNVs in the recurrent tumour of MB-REC-03 (x axis). At all positions, and in both the recurrent (a), and primary (b), tumours, we observe a high proportion (~100%) of reads with baseQ >30 at both the mutant (black square) and wild-type allele (white squares). Grey squares indicate alleles categorized as sequencing errors. Errors have low allelic frequencies (in many cases are just one read) and a much smaller proportion of reads with baseQ values >30. In the primary tumour, the baseQ and AF values match the pattern observed in the recurrent tumour, indicating that these calls represent true SNVs present at very low frequencies. Sequencing errors in the primary sample have the same base distribution as sequencing errors in the recurrent tumour sample.
Extended Data Figure 9 Pathway enrichment results for genes recurrently aberrant in the primary tumour or recurrent tumour cohorts.
Pathway enrichment analysis of gene lists derived from the integrative analysis of CNVs (gain or loss of 2 or more copies), SNVs, indels, and structural variants specific to the primary or recurrent tumours of each patient was performed using g:Cocoa.
Extended Data Figure 10 Genetic events in recurrent human medulloblastoma converge on specific signalling pathways.
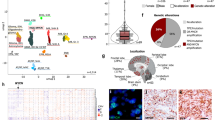
a, Copy number profile of MB-REC-12 therapy-naive (WT, green, left panel) and recurrent tumour (loss, blue, lower panel), showing recurrence-specific loss of chr14q. b, DYNC1H1 expression is reduced in Shh patients with chr14q loss (n = 18/80, Mann–Whitney test, P < 0.0001, centre lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles). c, Expression of the chr14 signature genes discriminating between chr14q balanced (n = 34) and chr14q loss (n = 18) in the Boston cohort of Shh medulloblastoma samples.
Supplementary information
Supplementary Information
This file contains full legends for Supplementary Tables 1-4, Supplementary Text and Data – see contents page for full details. (PDF 18306 kb)
Supplementary Table 1
This file contains driver genes prediction in mouse primary and recurrent tumours – see Supplementary Information document for full legend. (XLSX 48 kb)
Supplementary Table 2
This table contains patient information, analyses summary, and associated clinical data - see Supplementary Information document for full legend. (XLSX 17721 kb)
Supplementary Table 3
This table contains positions verified by deep sequencing – see Supplementary Information document for full legend. (XLS 1521 kb)
Supplementary Table 4
This table contains prognostic implications of chr14 loss in Shh MB – see Supplementary Information document for full legend. (XLSX 94 kb)
Rights and permissions
About this article
Cite this article
Morrissy, A., Garzia, L., Shih, D. et al. Divergent clonal selection dominates medulloblastoma at recurrence. Nature 529, 351–357 (2016). https://doi.org/10.1038/nature16478
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16478
This article is cited by
-
Comparison of transcriptome profiles between medulloblastoma primary and recurrent tumors uncovers novel variance effects in relapses
Acta Neuropathologica Communications (2023)
-
Molecular and functional profiling of chemotolerant cells unveils nucleoside metabolism-dependent vulnerabilities in medulloblastoma
Acta Neuropathologica Communications (2023)
-
Loss of phosphatase CTDNEP1 potentiates aggressive medulloblastoma by triggering MYC amplification and genomic instability
Nature Communications (2023)
-
ARF suppression by MYC but not MYCN confers increased malignancy of aggressive pediatric brain tumors
Nature Communications (2023)
-
Cytotoxicity of poly-guanidine in medulloblastoma cell lines
Investigational New Drugs (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.