Abstract
Influenza pandemics occur unpredictably when zoonotic influenza viruses with novel antigenicity acquire the ability to transmit amongst humans1. Host range breaches are limited by incompatibilities between avian virus components and the human host. Barriers include receptor preference, virion stability and poor activity of the avian virus RNA-dependent RNA polymerase in human cells2. Mutants of the heterotrimeric viral polymerase components, particularly PB2 protein, are selected during mammalian adaptation, but their mode of action is unknown3,4,5,6. We show that a species-specific difference in host protein ANP32A accounts for the suboptimal function of avian virus polymerase in mammalian cells. Avian ANP32A possesses an additional 33 amino acids between the leucine-rich repeats and carboxy-terminal low-complexity acidic region domains. In mammalian cells, avian ANP32A rescued the suboptimal function of avian virus polymerase to levels similar to mammalian-adapted polymerase. Deletion of the avian-specific sequence from chicken ANP32A abrogated this activity, whereas its insertion into human ANP32A, or closely related ANP32B, supported avian virus polymerase function. Substitutions, such as PB2(E627K), were rapidly selected upon infection of humans with avian H5N1 or H7N9 influenza viruses, adapting the viral polymerase for the shorter mammalian ANP32A. Thus ANP32A represents an essential host partner co-opted to support influenza virus replication and is a candidate host target for novel antivirals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Accessions
ArrayExpress
Data deposits
The microarray data have been submitted to The European Bioinformatics Institute (EBI) (http://www.ebi.ac.uk) ArrayExpress under accession number E-MTAB-3643.
References
Neumann, G. & Kawaoka, Y. Transmission of influenza A viruses. Virology 479-480, 234–246 (2015)
Cauldwell, A. V., Long, J. S., Moncorgé, O. & Barclay, W. S. Viral determinants of influenza A virus host range. J. Gen. Virol. 95, 1193–1210 (2014)
Almond, J. W. A single gene determines the host range of influenza virus. Nature 270, 617–618 (1977)
Subbarao, E. K., London, W. & Murphy, B. R. A single amino acid in the PB2 gene of influenza A virus is a determinant of host range. J. Virol. 67, 1761–1764 (1993)
Naffakh, N., Tomoiu, A., Rameix-Welti, M.-A. & van der Werf, S. Host restriction of avian influenza viruses at the level of the ribonucleoproteins. Annu. Rev. Microbiol. 62, 403–424 (2008)
Mänz, B., Schwemmle, M. & Brunotte, L. Adaptation of avian influenza A virus polymerase in mammals to overcome the host species barrier. J. Virol. 87, 7200–7209 (2013)
Eisfeld, A. J., Neumann, G. & Kawaoka, Y. At the centre: influenza A virus ribonucleoproteins. Nature Rev. Microbiol. 13, 28–41 (2015)
Steel, J., Lowen, A. C., Mubareka, S. & Palese, P. Transmission of influenza virus in a mammalian host is increased by PB2 amino acids 627K or 627E/701N. PLoS Pathog. 5, e1000252 (2009)
Van Hoeven, N. et al. Human HA and polymerase subunit PB2 proteins confer transmission of an avian influenza virus through the air. Proc. Natl Acad. Sci. USA 106, 3366–3371 (2009)
Moncorgé, O., Mura, M. & Barclay, W. S. Evidence for avian and human host cell factors that affect the activity of influenza virus polymerase. J. Virol. 84, 9978–9986 (2010)
Long, J. S. et al. The effect of the PB2 mutation 627K on highly pathogenic H5N1 avian influenza virus is dependent on the virus lineage. J. Virol. 87, 9983–9996 (2013)
Mehle, A. & Doudna, J. A. An inhibitory activity in human cells restricts the function of an avian-like influenza virus polymerase. Cell Host Microbe 4, 111–122 (2008)
Massin, P., van der Werf, S. & Naffakh, N. Residue 627 of PB2 is a determinant of cold sensitivity in RNA replication of avian influenza viruses. J. Virol. 75, 5398–5404 (2001)
Bussey, K. A., Bousse, T. L., Desmet, E. A., Kim, B. & Takimoto, T. PB2 residue 271 plays a key role in enhanced polymerase activity of influenza A viruses in mammalian host cells. J. Virol. 84, 4395–4406 (2010)
Mehle, A. & Doudna, J. A. Adaptive strategies of the influenza virus polymerase for replication in humans. Proc. Natl Acad. Sci. USA 106, 21312–21316 (2009)
Pflug, A., Guilligay, D., Reich, S. & Cusack, S. Structure of influenza A polymerase bound to the viral RNA promoter. Nature 516, 355–360 (2014)
Morisson, M. et al. ChickRH6: a chicken whole-genome radiation hybrid panel. Genet. Sel. Evol. 34, 521–533 (2002)
Bortz, E. et al. Host- and strain-specific regulation of influenza virus polymerase activity by interacting cellular proteins. MBio 2, e00151–11 (2011)
Gabriel, G. et al. Differential use of importin-α isoforms governs cell tropism and host adaptation of influenza virus. Nature Commun. 2, 156 (2011)
Reilly, P. T., Yu, Y., Hamiche, A. & Wang, L. Cracking the ANP32 whips: important functions, unequal requirement, and hints at disease implications. BioEssays 36, 1062–1071 (2014)
Bradel-Tretheway, B. G. et al. Comprehensive proteomic analysis of influenza virus polymerase complex reveals a novel association with mitochondrial proteins and RNA polymerase accessory factors. J. Virol. 85, 8569–8581 (2011)
Watanabe, T. et al. Influenza virus-host interactome screen as a platform for antiviral drug development. Cell Host Microbe 16, 795–805 (2014)
Sugiyama, K., Kawaguchi, A., Okuwaki, M. & Nagata, K. pp32 and APRIL are host cell-derived regulators of influenza virus RNA synthesis from cRNA. eLife 4, e08939 (2015)
Shinya, K. et al. Ostrich involvement in the selection of H5N1 influenza virus possessing mammalian-type amino acids in the PB2 protein. J. Virol. 83, 13015–13018 (2009)
Rameix-Welti, M.-A., Tomoiu, A., Dos Santos Afonso, E., van der Werf, S. & Naffakh, N. Avian Influenza A virus polymerase association with nucleoprotein, but not polymerase assembly, is impaired in human cells during the course of infection. J. Virol. 83, 1320–1331 (2009)
Crescenzo-Chaigne, B., van der Werf, S. & Naffakh, N. Differential effect of nucleotide substitutions in the 3′ arm of the influenza A virus vRNA promoter on transcription/replication by avian and human polymerase complexes is related to the nature of PB2 amino acid 627. Virology 303, 240–252 (2002)
Paterson, D., te Velthuis, A. J. W., Vreede, F. T. & Fodor, E. Host restriction of influenza virus polymerase activity by PB2 627E is diminished on short viral templates in a nucleoprotein-independent manner. J. Virol. 88, 339–344 (2014)
Weber, M. et al. Influenza virus adaptation PB2-627K modulates nucleocapsid inhibition by the pathogen sensor RIG-I. Cell Host Microbe 17, 309–319 (2015)
Resa-Infante, P. et al. The host-dependent interaction of α-importins with influenza PB2 polymerase subunit is required for virus RNA replication. PLoS One 3, e3904 (2008)
Whiteley, A. et al. Generation of candidate human influenza vaccine strains in cell culture - rehearsing the European response to an H7N1 pandemic threat. Influenza Other Respir. Viruses 1, 157–166 (2007)
Iqbal, M., Yaqub, T., Mukhtar, N., Shabbir, M. Z. & McCauley, J. W. Infectivity and transmissibility of H9N2 avian influenza virus in chickens and wild terrestrial birds. Vet. Res. 44, 100 (2013)
Elleman, C. J. & Barclay, W. S. The M1 matrix protein controls the filamentous phenotype of influenza A virus. Virology 321, 144–153 (2004)
Neumann, G. et al. Generation of influenza A viruses entirely from cloned cDNAs. Proc. Natl Acad. Sci. USA 96, 9345–9350 (1999)
Hoffmann, E., Neumann, G., Kawaoka, Y., Hobom, G. & Webster, R. G. A DNA transfection system for generation of influenza A virus from eight plasmids. Proc. Natl Acad. Sci. USA 97, 6108–6113 (2000)
Pleschka, S. et al. A plasmid-based reverse genetics system for influenza A virus. J. Virol. 70, 4188–4192 (1996)
Moncorgé, O. et al. Investigation of influenza virus polymerase activity in pig cells. J. Virol. 87, 384–394 (2013)
Flick, R. & Pettersson, R. F. Reverse genetics system for Uukuniemi virus (Bunyaviridae): RNA polymerase I-catalyzed expression of chimeric viral RNAs. J. Virol. 75, 1643–1655 (2001)
Howard, W. et al. Development of a reverse genetics system enabling the rescue of recombinant avian influenza virus A/Turkey/England/50-92/91 (H5N1). Avian Dis. 51 (Suppl. 1), 393–395 (2007)
Cauldwell, A. V., Moncorgé, O. & Barclay, W. S. Unstable polymerase-nucleoprotein interaction is not responsible for avian influenza virus polymerase restriction in human cells. J. Virol. 87, 1278–1284 (2013)
Naldini, L., Blömer, U., Gage, F. H., Trono, D. & Verma, I. M. Efficient transfer, integration, and sustained long-term expression of the transgene in adult rat brains injected with a lentiviral vector. Proc. Natl Acad. Sci. USA 93, 11382–11388 (1996)
Ulm, J. W., Perron, M., Sodroski, J. & C Mulligan, R. Complex determinants within the Moloney murine leukemia virus capsid modulate susceptibility of the virus to Fv1 and Ref1-mediated restriction. Virology 363, 245–255 (2007)
Obayashi, E. et al. The structural basis for an essential subunit interaction in influenza virus RNA polymerase. Nature 454, 1127–1131 (2008)
Kawakami, E. et al. Strand-specific real-time RT-PCR for distinguishing influenza vRNA, cRNA, and mRNA. J. Virol. Methods 173, 1–6 (2011)
Acknowledgements
We thank G. Maertens, J. Stech, R. Fouchier, A. Cauldwell, G. Roche, J. McCauley, D. Huntley, A. Vaughan, V. Nair and H. Shelton for provision of reagents, advice and discussions. This work was funded by BBSRC sLoLa BB/K002465/1 “Developing Rapid Responses to Emerging Virus Infections of Poultry (DDREVIP)” which funds J.S.L. and E.S.G., B.M. was funded by a Wellcome Trust studentship. R.F. and O.M. were funded by a Wellcome Trust Programme Grant (087039/Z/08/Z). O.M. was funded by MRC (G0600006). M.I. was funded by a BBSRC Avian Diseases Programme Grant (BBS/E/I/00001708).
Author information
Authors and Affiliations
Contributions
J.S.L. designed and performed the experiments and wrote the manuscript. E.S.G. performed microarrays and analysed data. O.M. generated plasmids for polymerase assays and wrote the manuscript. R.F., E.S.G. and B.M. performed qRT–PCR analysis. J.J. generated plasmids for polymerase assays. M.I. supplied UDL/08 reverse genetics system. A.V. and M.M. supplied radiation hybrid clones. M.A.S. analysed data, designed microarray experiments and wrote the manuscript. W.S.B. designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Analysis of mRNAs by PCA mapping reveals diversity of the radiation hybrid clones and their genetic instability during cell passage.
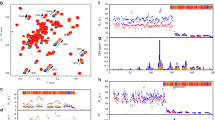
Each sphere represents a microarray sample. The percentage values in the axes parentheses designate proportion of overall variance as described by each PC. PC1 principal component 1 (x-axis); PC2 principal component 2 (y-axis); PC3 principal component 3 (z-axis). PC1 describes the predominant amount of variance (15.6%). Selection of negative clones (red), parent Wg3H cells (blue) and positive clones: 377 (purple), 386 (orange) and 365 (green) and 476 (cyan) arrays are distinguished by colour, and passage numbers 1 and 12 are distinguished by the size of spheres. Negative arrays are dispersed, while parent cells are accumulated further to the right of PC1 and upwards of PC2. Positive clones show distinct variability in their location while passaging reduced their separation from parent cells. This analysis accompanies Fig. 1.
Extended Data Figure 2 Confirmation of chANP32A and chANP32B expression in RH clones by qRT–PCR.
RNA was extracted from the RH clones after testing for influenza polymerase activity and analysed by microarray for chicken transcripts. The same RNA was used to validate identification of ANP32A by confirming the level of expression of ANP32A (and ANP32B as control) in the parent Wg3h cells, positive clones, passaged positive clones and a selection of negative clones. a, Copy numbers of chANP32A mRNA were calculated by qRT–PCR against a standard curve generated with chANP32A cDNA using primers specific for chANP32A. b, Copy number of ANP32B mRNA were measured by qRT–PCR against a standard curve generated with chANP32B cDNA using primers specific for chANP32B. (n = 3 technical replicates; error as s.e.m.). This analysis accompanies Fig. 1.
Extended Data Figure 3 Knockdown of chANP32A in positive RH clone 476 diminished the ability to support avian influenza polymerase activity.
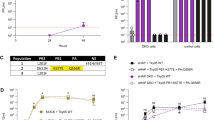
a, Positive RH clone 476 cells were transfected with 100 nM of siRNA targeting NP, chANP32A or no target (Allstars). After 48 h cells were transfected with mouse-polI-firefly minigenome reporter, avian influenza polymerase (H5N1 50–92) with either PB2 627E or 627K, Renilla control and either empty plasmid or codon optimised chANP32A (codon optimization according to algorithm by GeneArt with manual editing). (Data are luciferase activity measured after a further 24 h; n = 3 biological replicates; errors are displayed as s.e.m.). b, Knockdown of chANP32A was confirmed by qRT–PCR of RNA extracted from siRNA treated cells, calculated using a standard curve generated with chANP32A cDNA, using primers specific for chANP32A (n = 3 biological replicates; errors are displayed as s.e.m.). This analysis accompanies Fig. 1.
Extended Data Figure 4 Expression of chANP32A in human cells permits influenza polymerase activity of several avian influenza polymerases and an avianized human influenza polymerase and increases avian virus replication.
293T cells were transfected with empty vector, chANP32A or huANP32A. a, b, 20 h later, cells were transfected with pHOM1-firefly minigenome reporter, and the polymerase set from low pathogenicity avH1N1 (Bav) or H9N2 (UDL), highly pathogenic H5N1 (50-92), H5N1 (Ty05), or huH3N2 (Vic) viruses, with either PB2 627E (a) or 627K (b) and Renilla expression control. After a further 24 h luciferase activity was measured. (Data are mean PB2 627E or K polymerase activity normalized to Renilla; n = 3 biological replicates; error plotted as s.e.m. of the ratio; pattern of results consistent in at least three independent experiments). This analysis accompanies Fig. 1. c, 20 h after transfection with ch or huANP32A or empty vector, cells were infected with avian-like influenza virus (H5N1Ty05:PR8 6:2 recombinant virus) (MOI 0.1) bearing PB2 627E (black bars) or PB2 627K (grey bars). Infected cells were incubated at 37 °C and cell supernatant titrated for infectious virus at 24 h post infection on MDCK cells by plaque assay.(Data displayed as log10 plaque forming units per ml; n = 3 biological replicates; error plotted as s.e.m.; one-way ANOVA, comparisons to empty vector, NS= not significant, *P < 0.05 ***P < 0.001; pattern of results consistent in at least three independent experiments). This analysis accompanies Fig. 2.
Extended Data Figure 5 chANP32A does not alter expression or nuclear accumulation of avian PB2 protein in human cells.
293T cells were transfected with pHOM1-firefly minigenome, avian influenza polymerase and NP of H5N1 50–92 (PB2 627E) together with empty vector, chANP32A or chANP32AΔ33 or cells were untransfected (Mock). Cell monolayers were harvested after 24 h and lysed in 0.1% NP40 lysate buffer and total fractions taken before centrifugation to generate a nuclear pellet and cytoplasmic fraction. Nuclear pellets were resuspended in 1% NP40 buffer. a, Protein levels of vinculin (cytoplasmic marker) and lamin B (nuclear marker) and of PB2 in total, nuclear or cytoplasmic fractions were analysed by immunoblotting. b, Total lysates were immunoblotted for vinculin, PB2 and Flag peptide. c, Immunoblots were quantified using Image Studio Lite V5.2. The ratio of nuclear to cytoplasmic PB2 was calculated by dividing the ratio of PB2 to vinculin by the ratio of PB2 to lamin B from the cytoplasmic and nuclear fractions, respectively. Data are the mean ratios from three independent experiments (excepting chANP32Δ33 for which only 2 data points were available), error bars are s.e.m. Data are not statistically significantly different by one-way ANOVA. This analysis accompanies Fig. 2.
Extended Data Figure 6 Quantification of knockdown of chANP32A in chicken cells.
DF-1 cells were transduced with VSV-G lentiviral vectors that delivered a transgene expressing shRNA directed against chANP32A or a negative sequence and the puromycin gene. Puromycin selected cells were transfected with siRNA (100 nM) (underlined). RNA was extracted from untreated shRNA cells and siRNA-treated shRNA cells. Knockdown of chANP32A was quantified by qRT–PCR of the extracted RNA, calculated using a standard curve generated with chANP32A cDNA, using primers specific for chANP32A. Fold decrease of RNA copies is displayed compared to negative shRNA DF-1 or ALLstars treated chANP32A shRNA DF-1 cells. (n = 3 biological replicates; error displayed as s.e.m.). This analysis accompanies experiments in Fig. 3a–c.
Extended Data Figure 7 siRNA knockdown demonstrates that human-adapted influenza polymerase activity is dependent on huANP32A and huANP32B in human cells.
a, 293T cells were transfected with siRNA (100 nM) against NP, huANP32A, huANP32B or both huANP32A and huANP32B (50 nM each). After 48 h, cells were transfected with pHOM1-firefly minigenome, human-adapted avian influenza polymerase (H5N1 50–92 PB2 627K), and Renilla expression control. Luciferase activity was measured after a further 24 h. (Data are firefly activity normalized to Renilla, plotted as % of Allstars; n = 3 biological replicates; error as s.e.m.; one-way ANOVA comparisons to Allstars, ****P < 0.0001;). b, Knockdown of gene targets was verified by immunoblotting using antibody against vinculin, huANP32A and huANP32B. This analysis accompanies Fig. 3e, f.
Extended Data Figure 8 Alignment of ANP32A proteins reveals significant homology except for an extra 33 amino acid sequence in birds that is absent in mammals and ostrich and lacking from ANP32B family members.
The protein sequences of ANP32A for chicken, duck, zebra finch, turkey, ostrich, human, mouse and pig together with sequences of ANP32B for chicken and human were aligned using Geneious R6 software. chANP32A is set as the reference sequence, and colours represent similarity of amino acid identity (black, 100%; dark grey, 80–100%; light grey, 60–80%; white, <60%). Gaps are annotated by dashes. Residue numbers correspond to chANP32A. The 33 amino acid sequence found in avian species is situated between residues 176–208. This analysis accompanies Fig. 4.
Extended Data Figure 9 Expression of ANP32A and B proteins reduced human-adapted influenza polymerase activity in human cells.
293T cells were transfected with Flag-tagged ANP32 constructs and after 20 h transfected with pHOM1-firefly minigenome reporter, human-adapted influenza polymerase (H5N1 50–92 with PB2 627K, together with Renilla expression control. Cells were assayed for luciferase activity 24 h later. (Data are PB2 627K polymerase activity normalized to Renilla; n = 3 biological replicates; error plotted as s.e.m. of the ratio; one-way ANOVA, all constructs were significantly reduced compared to empty vector (P < 0.0001); pattern of results consistent in at least three independent experiments). These data relate to Fig. 4.
Rights and permissions
About this article
Cite this article
Long, J., Giotis, E., Moncorgé, O. et al. Species difference in ANP32A underlies influenza A virus polymerase host restriction. Nature 529, 101–104 (2016). https://doi.org/10.1038/nature16474
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16474
This article is cited by
-
Creating resistance to avian influenza infection through genome editing of the ANP32 gene family
Nature Communications (2023)
-
An Influenza A virus can evolve to use human ANP32E through altering polymerase dimerization
Nature Communications (2023)
-
Screening and identification of specific cluster miRNAs in N2a cells infected by H7N9 virus
Virus Genes (2023)
-
BTN3A3 evasion promotes the zoonotic potential of influenza A viruses
Nature (2023)
-
Are pigs overestimated as a source of zoonotic influenza viruses?
Porcine Health Management (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.