Abstract
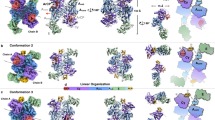
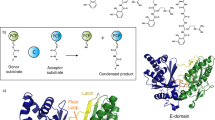
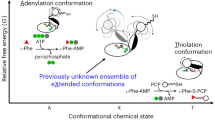
Many important natural products are produced by multidomain non-ribosomal peptide synthetases (NRPSs)1,2,3,4. During synthesis, intermediates are covalently bound to integrated carrier domains and transported to neighbouring catalytic domains in an assembly line fashion5. Understanding the structural basis for catalysis with non-ribosomal peptide synthetases will facilitate bioengineering to create novel products. Here we describe the structures of two different holo-non-ribosomal peptide synthetase modules, each revealing a distinct step in the catalytic cycle. One structure depicts the carrier domain cofactor bound to the peptide bond-forming condensation domain, whereas a second structure captures the installation of the amino acid onto the cofactor within the adenylation domain. These structures demonstrate that a conformational change within the adenylation domain guides transfer of intermediates between domains. Furthermore, one structure shows that the condensation and adenylation domains simultaneously adopt their catalytic conformations, increasing the overall efficiency in a revised structural cycle. These structures and the single-particle electron microscopy analysis demonstrate a highly dynamic domain architecture and provide the foundation for understanding the structural mechanisms that could enable engineering of novel non-ribosomal peptide synthetases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Koglin, A. & Walsh, C. T. Structural insights into nonribosomal peptide enzymatic assembly lines. Nat. Prod. Rep. 26, 987–1000 (2009)
Fischbach, M. A. & Walsh, C. T. Antibiotics for emerging pathogens. Science 325, 1089–1093 (2009)
Walsh, C. T. The chemical versatility of natural-product assembly lines. Acc. Chem. Res. 41, 4–10 (2008)
Walsh, C. T. & Fischbach, M. A. Natural products version 2.0: connecting genes to molecules. J. Am. Chem. Soc. 132, 2469–2493 (2010)
Strieker, M., Tanovic´, A. & Marahiel, M. A. Nonribosomal peptide synthetases: structures and dynamics. Curr. Opin. Struct. Biol. 20, 234–240 (2010)
Mercer, A. C. & Burkart, M. D. The ubiquitous carrier protein--a window to metabolite biosynthesis. Nat. Prod. Rep. 24, 750–773 (2007)
Gulick, A. M. Conformational dynamics in the Acyl-CoA synthetases, adenylation domains of non-ribosomal peptide synthetases, and firefly luciferase. ACS Chem. Biol. 4, 811–827 (2009)
Conti, E., Stachelhaus, T., Marahiel, M. A. & Brick, P. Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S. EMBO J. 16, 4174–4183 (1997)
Mitchell, C. A., Shi, C., Aldrich, C. C. & Gulick, A. M. Structure of PA1221, a nonribosomal peptide synthetase containing adenylation and peptidyl carrier protein domains. Biochemistry 51, 3252–3263 (2012)
Sundlov, J. A., Shi, C., Wilson, D. J., Aldrich, C. C. & Gulick, A. M. Structural and functional investigation of the intermolecular interaction between NRPS adenylation and carrier protein domains. Chem. Biol. 19, 188–198 (2012)
Goodrich, A. C., Harden, B. J. & Frueh, D. P. Solution structure of a nonribosomal peptide synthetase carrier protein loaded with its substrate reveals transient, well-defined contacts. J. Am. Chem. Soc. 137, 12100–12109 (2015)
Jaremko, M. J., Lee, D. J., Opella, S. J. & Burkart, M. D. Structure and substrate sequestration in the pyoluteorin type II peptidyl carrier protein PltL. J. Am. Chem. Soc. 137, 11546–11549 (2015)
Tanovic, A., Samel, S. A., Essen, L. O. & Marahiel, M. A. Crystal structure of the termination module of a nonribosomal peptide synthetase. Science 321, 659–663 (2008)
Clemmer, K. M., Bonomo, R. A. & Rather, P. N. Genetic analysis of surface motility in Acinetobacter baumannii. Microbiology 157, 2534–2544 (2011)
Rumbo-Feal, S. et al. Whole transcriptome analysis of Acinetobacter baumannii assessed by RNA-sequencing reveals different mRNA expression profiles in biofilm compared to planktonic cells. PLoS One 8, e72968 (2013)
Giles, S. K., Stroeher, U. H., Eijkelkamp, B. A. & Brown, M. H. Identification of genes essential for pellicle formation in Acinetobacter baumannii. BMC Microbiol . 15, 116 (2015)
Bloudoff, K., Rodionov, D. & Schmeing, T. M. Crystal structures of the first condensation domain of CDA synthetase suggest conformational changes during the synthetic cycle of nonribosomal peptide synthetases. J. Mol. Biol. 425, 3137–3150 (2013)
Marahiel, M. A., Stachelhaus, T. & Mootz, H. D. Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem. Rev. 97, 2651–2674 (1997)
Frueh, D. P. et al. Dynamic thiolation-thioesterase structure of a non-ribosomal peptide synthetase. Nature 454, 903–906 (2008)
Liu, Y., Zheng, T. & Bruner, S. D. Structural basis for phosphopantetheinyl carrier domain interactions in the terminal module of nonribosomal peptide synthetases. Chem. Biol. 18, 1482–1488 (2011)
Qiao, C., Wilson, D. J., Bennett, E. M. & Aldrich, C. C. A mechanism-based aryl carrier protein/thiolation domain affinity probe. J. Am. Chem. Soc. 129, 6350–6351 (2007)
Miller, B. R., Sundlov, J. A., Drake, E. J., Makin, T. A. & Gulick, A. M. Analysis of the linker region joining the adenylation and carrier protein domains of the modular nonribosomal peptide synthetases. Proteins 82, 2691–2702 (2014)
Gaudelli, N. M., Long, D. H. & Townsend, C. A. β-Lactam formation by a non-ribosomal peptide synthetase during antibiotic biosynthesis. Nature 520, 383–387 (2015)
Maruyama, C. et al. A stand-alone adenylation domain forms amide bonds in streptothricin biosynthesis. Nature Chem. Biol. 8, 791–797 (2012)
Dutta, S. et al. Structure of a modular polyketide synthase. Nature 510, 512–517 (2014)
Adams, M. D. et al. Comparative genome sequence analysis of multidrug-resistant Acinetobacter baumannii. J. Bacteriol. 190, 8053–8064 (2008)
Kapust, R. B. et al. Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Protein Eng. 14, 993–1000 (2001)
Doublié, S. Preparation of selenomethionyl proteins for phase determination. Methods Enzymol. 276, 523–530 (1997)
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D 67, 271–281 (2011)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nature Protocols 3, 1171–1179 (2008)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Urzhumtsev, A., Afonine, P. V. & Adams, P. D. TLS from fundamentals to practice. Crystallogr. Rev . 19, 230–270 (2013)
Roche, E. D. & Walsh, C. T. Dissection of the EntF condensation domain boundary and active site residues in nonribosomal peptide synthesis. Biochemistry 42, 1334–1344 (2003)
Luft, J. R. et al. A deliberate approach to screening for initial crystallization conditions of biological macromolecules. J. Struct. Biol. 142, 170–179 (2003)
Sundlov, J. A. & Gulick, A. M. Structure determination of the functional domain interaction of a chimeric nonribosomal peptide synthetase from a challenging crystal with noncrystallographic translational symmetry. Acta Crystallogr. D 69, 1482–1492 (2013)
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D 62, 1002–1011 (2006)
Ohi, M., Li, Y., Cheng, Y. & Walz, T. Negative staining and image classification - powerful tools in modern electron microscopy. Biol. Proced. Online 6, 23–34 (2004)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Yang, Z., Fang, J., Chittuluru, J., Asturias, F. J. & Penczek, P. A. Iterative stable alignment and clustering of 2D transmission electron microscope images. Structure 20, 237–247 (2012)
Ikeuchi, H., Meyer, M. E., Ding, Y., Hiratake, J. & Richards, N. G. A critical electrostatic interaction mediates inhibitor recognition by human asparagine synthetase. Bioorg. Med. Chem. 17, 6641–6650 (2009)
Rusnak, F., Faraci, W. S. & Walsh, C. T. Subcloning, expression, and purification of the enterobactin biosynthetic enzyme 2,3-dihydroxybenzoate-AMP ligase: demonstration of enzyme-bound (2,3-dihydroxybenzoyl)adenylate product. Biochemistry 28, 6827–6835 (1989)
Horswill, A. R. & Escalante-Semerena, J. C. Characterization of the propionyl-CoA synthetase (PrpE) enzyme of Salmonella enterica: residue Lys592 is required for propionyl-AMP synthesis. Biochemistry 41, 2379–2387 (2002)
Reger, A. S., Carney, J. M. & Gulick, A. M. Biochemical and crystallographic analysis of substrate binding and conformational changes in acetyl-CoA synthetase. Biochemistry 46, 6536–6546 (2007)
Wilson, D. J. & Aldrich, C. C. A continuous kinetic assay for adenylation enzyme activity and inhibition. Anal. Biochem. 404, 56–63 (2010)
Reuter, D. C., McIntosh, J. E., Guinn, A. C. & Madera, A. M. Synthesis of vinyl sulfonamides using the Horner reaction. Synthesis 2003, 2321–2324 (2003)
Acknowledgements
We thank R. Sanishvili for assistance with data collection. This work was funded in part by National Institutes of Health GM-068440 (to A.M.G.) and GM-115601 (to G.S.), and Award W81XWH-11-2-0218 from the Telemedicine and Advanced Technology Research Center of the US Army Medical Research and Materiel Command (A.M.G.). Data were collected at the GM/CA beamline of the Advanced Photon Source, which is funded by the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006) under Department of Energy contract number DE-AC02-06CH11357 to A.P.S. A Stafford Fellowship (to B.R.M.) and support from the Hauptman-Woodward Institute is acknowledged.
Author information
Authors and Affiliations
Contributions
C.L.A. characterized activity of and initially crystallized AB3403. J.A.S. initially crystallized EntF. E.J.D. and B.R.M. optimized crystal, and solved and refined the models of AB3403 and EntF, respectively. C.S. and C.C.A. designed and synthesized the mechanism-based inhibitor. J.T.T. and G.S. performed and analysed the single-particle electron microscopy. A.M.G., E.J.D., B.R.M., G.S., J.T.T., C.C.A., and C.S. analysed the results and wrote the manuscript. All authors saw and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Structure-based alignment of EntF, AB3403, and SrfA-C.
Condensation, adenylation, PCP, and thioesterase domains are represented with bars in grey, pink, green–cyan, and blue. Conserved motifs and catalytically important residues are highlighted with the same colours, including the HHxxxD motif of the condensation domains, the aspartic acid hinge that separates the N- and C-terminal subdomains of the adenylation domain, the GGHS motif that is the site of pantetheinylation in the PCP, and the catalytic nucleophile of the thioesterase domain. The SrfA-C, AB3403, and EntF proteins share approximately 26% sequence identity. The adenylation and PCP domains are more well-conserved, sharing ~35% identity, whereas the condensation (21%) and thioesterase (25%) domains are less well conserved. Domain boundaries are described in the table below.
Extended Data Figure 2 Substrate specificity of full-length AB3403.
Amino-acid specificity of AB3403 was recorded for all 20 proteinogenic amino acids, as well as 4-chlorobenzoate (4CB) and 4-hydroxybenzoate (4HB). Average values and standard deviations are shown for three replicates with each substrate; results were recorded as micromoles of radiolabelled ATP incorporated per minute per milligram of enzyme. Apparent kinetic constants are also shown for ATP and glycine calculated from duplicate measurements for four to six substrate concentrations.
Extended Data Figure 3 Stereo representations of electron density figures shown in Fig. 2.
To better visualize the active sites and electron density quality, stereo figures are included in the extended data. In all panels, density is shown with coefficients of the form (Fo − Fc) calculated before inclusion of ligands and contoured at 3σ. a, Stereo representation of electron density of AB3403 condensation domain shows the phosphopantethine on Ser1006 approaching His145 within the condensation domain pocket. Inhibitor carbon atoms in green, carbons of residues within 5 Å of inhibitor in grey, nitrogen in blue, oxygen in red, sulphur in yellow, and water in light blue. b, Electron density of the nucleotide binding pocket of AB3403 bound to glycine and AMP. Stereo representation of electron density shows the AMP, glycine, and Mg+ present in the active site of the adenylation domain. Ligand carbon atoms are in green, carbons of residues within 5 Å of inhibitor in grey, nitrogen in blue, oxygen in red, phosphorus in orange, and the Mg+ cofactor in purple. c, Stereo representation of the electron density shows the phosphopantethine on Ser1006 covalently attached to the Ser-AVS inhibitor in the active site of the adenylation domain. Inhibitor carbon atoms in green, carbons of residues within 4 Å of inhibitor in grey, nitrogen in blue, oxygen in red, phosphorus in orange, sulphur in yellow, and water in light blue.
Extended Data Figure 4 Comparison of AB3403 and SrfA-C PCP-condensation domain interaction.
Stereo representation illustrating different orientations of the PCP domains of SrfA-C and AB3403 relative to the condensation domains with which they interact. AB3403 is shown with a white condensation domain and a green-cyan PCP. SrfA-C is shown with a yellow condensation domain and a pale blue PCP. The pantetheine of AB3403 is shown bound to Ser1006. The position of Ser1003, mutated to an alanine residue in SrfA-C, is also highlighted.
Extended Data Figure 5 Comparison of AB3403 thioesterase domain to the functional PCP–thioesterase interaction.
Stereo representation of the thioesterase (blue) domain of AB3403 interacts with the back face of the PCP domain in AB3403. The functional interaction between the EntF thioesterase domain and its holo-PCP, trapped crystallographically, illustrates that the same face of the thioesterase domain interacts functionally (PDB 3TEJ). A 28-residue insertion of AB3403 is coloured yellow.
Extended Data Figure 6 Synthesis of Ser-AVS.
The Ser-AVS probe was synthesized following similar protocols described elsewhere41,46. Garner’s aldehyde 1 was coupled with 2 using LiHMDS to exclusively furnish the (E)-vinylsulfonamide 3. Mitsunobu coupling of 3 with bis-Boc adenosine 4 afforded 5, which was globally deprotected using 80% aqueous trifluoroacetic acid to yield Ser-AVS.
Extended Data Figure 7 Electrophoretic mobility of EntF.
a, Native gel electrophoresis. Lane 1: EntF. Lane2: EntF incubated with fourfold molar excess of Ser-AVS inhibitor. Lane 3: EntF Crystals. Lane 4: novex NativeMark labelled in kilodaltons. b, Denaturing gel electrophoresis using loading buffer with SDS and β-mercaptoethanol. Gel lane 1: EntF. Lane 2: EntF incubated four times with Ser-AVS inhibitor. Lane 3: Life Technologies Mark12 labelled in kilodaltons. The native gel shows the inhibited EntF in a more compact conformation compared with EntF without the inhibitor.
Extended Data Figure 8 Negative-stain electron microscopy analysis of EntF.
a, Raw electron microscopy image of negative-stained EntF. b, Class averages of EntF particles.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion. (PDF 114 kb)
Rights and permissions
About this article
Cite this article
Drake, E., Miller, B., Shi, C. et al. Structures of two distinct conformations of holo-non-ribosomal peptide synthetases. Nature 529, 235–238 (2016). https://doi.org/10.1038/nature16163
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16163
This article is cited by
-
Subdomain dynamics enable chemical chain reactions in non-ribosomal peptide synthetases
Nature Chemistry (2024)
-
Advances in the adenylation domain: discovery of diverse non-ribosomal peptides
Applied Microbiology and Biotechnology (2023)
-
Structural insights into the substrate-bound condensation domains of non-ribosomal peptide synthetase AmbB
Scientific Reports (2022)
-
Bifurcation drives the evolution of assembly-line biosynthesis
Nature Communications (2022)
-
Rational inhibitor design for Pseudomonas aeruginosa salicylate adenylation enzyme PchD
JBIC Journal of Biological Inorganic Chemistry (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.