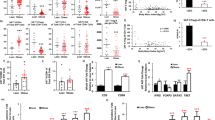
Abstract
Age-associated insulin resistance (IR) and obesity-associated IR are two physiologically distinct forms of adult-onset diabetes. While macrophage-driven inflammation is a core driver of obesity-associated IR1,2,3,4,5,6, the underlying mechanisms of the obesity-independent yet highly prevalent age-associated IR7 are largely unexplored. Here we show, using comparative adipo-immune profiling in mice, that fat-resident regulatory T cells, termed fTreg cells, accumulate in adipose tissue as a function of age, but not obesity. Supporting the existence of two distinct mechanisms underlying IR, mice deficient in fTreg cells are protected against age-associated IR, yet remain susceptible to obesity-associated IR and metabolic disease. By contrast, selective depletion of fTreg cells via anti-ST2 antibody treatment increases adipose tissue insulin sensitivity. These findings establish that distinct immune cell populations within adipose tissue underlie ageing- and obesity-associated IR, and implicate fTreg cells as adipo-immune drivers and potential therapeutic targets in the treatment of age-associated IR.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ferrante, A. W. Jr. Macrophages, fat, and the emergence of immunometabolism. J. Clin. Invest. 123, 4992–4993 (2013)
Lumeng, C. N. & Saltiel, A. R. Inflammatory links between obesity and metabolic disease. J. Clin. Invest. 121, 2111–2117 (2011)
Mathis, D. Immunological goings-on in visceral adipose tissue. Cell Metab. 17, 851–859 (2013)
Osborn, O. & Olefsky, J. M. The cellular and signaling networkslinking the immune system and metabolism in disease. Nature Med. 18, 363–374 (2012)
Weisberg, S. P. et al. Obesity is associated with macrophage accumulation in adipose tissue. J. Clin. Invest. 112, 1796–1808 (2003)
Xu, H. Chronic inflammation in fat plays a crucial role in the development of obesity-related insulin resistance. J. Clin. Invest. 112, 1821–1830 (2003)
CDC. 2014 National Diabetes Statistics Report; http://www.cdc.gov/diabetes/data/statistics/2014statisticsreport.html (2014)
Wu, D. et al. Eosinophils sustain adipose alternatively activated macrophages associated with glucose homeostasis. Science 332, 243–247 (2011)
Qiu, Y. et al. Eosinophils and type 2 cytokine signaling in macrophages orchestrate development of functional beige fat. Cell 157, 1292–1308 (2014)
Rao, R. R. et al. Meteorin-like is a hormone that regulates immune-adipose interactions to increase beige fat thermogenesis. Cell 157, 1279–1291 (2014)
Odegaard, J. I. et al. Macrophage-specific PPARγ controls alternative activation and improves insulin resistance. Nature 447, 1116–1120 (2007)
Fujisaka, S. et al. Regulatory mechanisms for adipose tissue m1 and m2 macrophages in diet-induced obese mice. Diabetes 58, 2574–2582 (2009)
Feuerer, M. et al. Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nature Med. 15, 930–939 (2009)
Lumeng, C. N. et al. Aging is associated with an increase in T cells and inflammatory macrophages in visceral adipose tissue. J. Immunol. 187, 6208–6216 (2011)
Cipolletta, D. et al. PPAR-γ is a major driver of the accumulation and phenotype of adipose tissue Treg cells. Nature 486, 549–553 (2012)
Houtkooper, R. H. et al. The metabolic footprint of aging in mice. Sci. Rep. 1, 134 (2011)
Sun, K. et al. Dichotomous effects of VEGF-A on adipose tissue dysfunction. Proc. Natl Acad. Sci. USA 109, 5874–5879 (2012)
Khan, T. et al. Metabolic dysregulation and adipose tissue fibrosis: role of collagen VI. Mol. Cell. Biol. 29, 1575–1591 (2009)
Steppan, C. M. et al. The hormone resistin links obesity to diabetes. Nature 409, 307–312 (2001)
Schwartz, D. R. & Lazar, M. A. Human resistin: found in translation from mouse to man. Trends Endocrinol. Metab. 22, 259–265 (2011)
Webster, K. E. et al. In vivo expansion of T reg cells with IL-2-mAb complexes: induction of resistance to EAE and long-term acceptance of islet allografts without immunosuppression. J. Exp. Med. 206, 751–760 (2009)
Feuerer, M. et al. Genomic definition of multiple ex vivo regulatory T cell subphenotypes. Proc. Natl Acad. Sci. USA 107, 5919–5924 (2010)
Josefowicz, S. Z., Lu, L.-F. & Rudensky, A. Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012)
Ohkura, N., Kitagawa, Y. & Sakaguchi, S. Development and maintenance of regulatory T cells. Immunity 38, 414–423 (2013)
Wing, K. et al. CTLA-4 control over Foxp3+ regulatory T cell function. Science 322, 271–275 (2008)
Fontenot, J. D. et al. A function for interleukin 2 in Foxp3-expressing regulatory T cells. Nature Immunol. 6, 1142–1151 (2005)
Vasanthakumar, A. et al. The transcriptional regulators IRF4, BATF and IL-33 orchestrate development and maintenance of adipose tissue–resident regulatory T cells. Nature Immunol. 16, 276–285 (2015)
Schiering, C. et al. The alarmin IL-33 promotes regulatory T-cell function in the intestine. Nature 513, 564–568 (2014)
Asterholm, I. W. et al. Adipocyte inflammation is essential for healthy adipose tissue expansion and remodeling. Cell Metab. 20, 103–118 (2014)
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008)
He, W. et al. Adipose-specific peroxisome proliferator-activated receptor gamma knockout causes insulin resistance in fat and liver but not in muscle. Proc. Natl Acad. Sci. USA 100, 15712–15717 (2003)
Liston, A. et al. Differentiation of regulatory Foxp3+ T cells in the thymic cortex. Proc. Natl Acad. Sci. USA 105, 11903–11908 (2008)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nature Biotechnol. 31, 46–53 (2012)
Roberts, A., Pimentel, H., Trapnell, C. & Pachter, L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27, 2325–2329 (2011)
Monticelli, L. A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nature Immunol. 12, 1045–1054 (2011)
Acknowledgements
We would like to thank L. Chong, J. Alvarez, Y. Dai, S. Kaufman and B. Collins for technical assistance, L. Ong and C. Brondos for administrative assistance, and J. Simon for assistance with graphics. S.P.B. is supported by National Institutes of Health (NIH) grants F30 DK096828 and T32 GM007198. C.L. and M.D. are funded by grants from the National Health and Medical Research Council of Australia Project grants 512354, 632886 and 1043199. R.M.E. is an Investigator of the Howard Hughes Medical Institute (HHMI) at the Salk Institute and March of Dimes Chair in Molecular and Developmental Biology, and is supported by NIH grants DK057978, DK090962, HL088093, HL105278 and ES010337, the Glenn Foundation for Medical Research, the Leona M. and Harry B. Helmsley Charitable Trust, Ipsen/Biomeasure, California Institute for Regenerative Medicine and The Ellison Medical Foundation. Y. Zheng is supported by the Nomis Foundation, the Rita Allen Foundation, the Emerald Foundation, the Hearst Foundation, the National Multiple Sclerosis Society, and National Institutes of Health (AI099295 and AI107027). This work was also supported by National Cancer Institute funded Salk Institute Cancer Center core facilities (CA014195) and the James B. Pendleton Charitable Trust.
Author information
Authors and Affiliations
Contributions
S.P.B., J.M.S., M.D., R.M.E. and Y. Zheng designed and supervised the research. S.P.B., J.M.S., S.F., S.L., Y. Zhang, A.C., C.Z. and Y.L. performed research. S.P.B., J.M.S., M.L., C.L., A.R.A., R.T.Y., M.D., R.M.E. and Y. Zheng analysed data. S.P.B., J.M.S., S.L., A.R.A., R.T.Y., M.D., R.M.E. and Y. Zheng wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.B., R.M.E., Y. Zheng and M.D. are co-inventors of technologies related to methods of targeting fat Treg cells to prevent insulin resistance.
Extended data figures and tables
Extended Data Figure 1 Schematic outlining study premise and selected gating strategies used to generate AIPs.
a, Schematic outlining study premise. b–d, AIPs were generated through the use of several distinct antibody cocktails. Here, using Foxp3Cre (Foxp3-IRES-YFP-Cre) reporter mice, we show how the stromal vascular fraction of VAT was analysed by flow cytometry to identify several T cell subtypes (b), macrophage subsets (c) and eosinophils and neutrophils (d).
Extended Data Figure 2 AIPs of aged fTreg knockout and control mice in VAT, SAT and spleen.
a, AIPs of aged (~14 months) fTreg knockout and control male mice depicting immune cell abundance, expressed as percentage of CD45.2+ cells. Entirety of immune compartment (top) is further divided into pan-macrophage (middle) and non-macrophage (bottom) pie charts (n = 9 mice per group). b–d, Immune cell abundance between fTreg knockout and control mice, expressed as cells per gram of VAT (b), SAT (c) and spleen (d) (n = 9 mice per group). Data are mean ± s.e.m. #, false discovery rate < 2%.
Extended Data Figure 3 Aged fTreg knockout mice do not show signs of systemic autoimmunity or breakdown in peripheral tolerance.
a, Percentage of splenic naive CD4+ T cells as defined by CD62hi CD44lo relative to total CD4+ CD25− Foxp3Cre population (n = 9 mice per group). b, Representative histology of gastrointestinal tract—duodenum, jejunum, ileum and colon (left to right) (n = 3 mice per group). There were no significant lesions observed or differences in inflammation, epithelial changes, or mucosal architecture between the two groups (H&E, original magnification, ×100). Scale bar, 50 μm. c, Histopathology score in the small intestine and colon of fTreg knockout and control mice. The severity and extent of inflammation and epithelial changes as well as mucosal architecture were each graded on a score of 1 (minimal) to 5 (severe) and added to obtain an overall score over 20. There were minimal inflammatory changes with no significant differences between groups. d, Multiplex inflammation panel of serum from fTreg knockout and control mice (n = 4 pooled samples (3 mice per sample) per group). Data are mean ± s.e.m. *P < 0.05, ***P < 0.001, Student’s t-test.
Extended Data Figure 4 Weight-matched cohorts’ body weights and adipocyte size frequency in VAT of aged control and fTreg knockout mice.
a, Body weights of fTreg knockout and control male mice used in weight-matched metabolic studies in young (12 week; control n = 9; fTreg KO n = 9), aged (36 week; control, n = 9 mice; fTreg KO, n = 11 mice) and obese (diet-induced obese, 12 weeks of HFD starting at 12 weeks; control n = 10; fTreg KO n = 10) settings. b, Frequency of small (0–5,000 μm2), medium (5,000–10,000 μm2) and large (>10,000 μm2) adipocytes in VAT of aged control and fTreg knockout mice (n = 3 mice per group, 850 adipocytes counted from control mice, 269 adipocytes counted from fTreg knockout adipose). Data are mean ± s.e.m.
Extended Data Figure 5 VAT AIPs of obese fTreg knockout and control mice.
a, AIPs of diet-induced obese (16 weeks high fat diet started at 12 weeks) control (n = 6 mice) and fTreg knockout (n = 8 mice) male mice depicting immune cell abundance, expressed as percentage of CD45.2+ cells. Entirety of immune compartment (top) is further divided into pan-macrophage (middle) and non-macrophage (bottom) pie charts. b, Immune cell abundance between fTreg knockout and control mice, expressed as cells per gram of VAT (n = 9 mice per group). Data are mean ± s.e.m.
Extended Data Figure 6 fTreg cells are dispensable for TZDs to exert their therapeutic insulin-sensitizing effect.
a, Scheme used for longitudinal interventional study of control and fTreg knockout mice which indicates when particular assays were conducted and whose results are described in b–g, in which rosiglitazone (Rosi) was introduced in diet after firmly establishing obesity with a HFD alone for 12 weeks (n = 8 mice per group). b, Cohort weights during course of study. Black arrow indicates introduction of rosiglitazone to the diet. c, Homeostatic model assessment of IR (HOMA-IR). d, e, Glucose tolerance test (d) and glucose excursions of glucose tolerance test (e) described as area under curve (AUC). f, g, Insulin tolerance test (f) and bar-graph quantitation of relative serum glucose decrease during insulin tolerance test (g) described as area above curve (AAC). h, Scheme used for parallel prophylactic study of control and fTreg knockout mice, the results of which are described in i–l, in which mice were placed on a HFD or HFD with rosiglitazone for 12 weeks (n = 8 mice per group). i, Cohort weights at end of study. j, HOMA-IR. k, l, Glucose and insulin tolerance tests of control (k) or fTreg knockout (l) mice fed HFD or HFD with rosiglitazone. m, Scheme used to determine temporal relationship of TZD-induced fTreg expansion and TZD-induced insulin-sensitization in wild-type mice, the results of which are described in n–q, where mice were fed HFD or HFD with rosiglitazone for up to 11 weeks (n = 10 mice per group, 5 mice of each group were euthanized at 5 weeks after diet introduction and remaining 5 mice were euthanized at 11 weeks). n, HOMA-IR at 4 weeks. o, p, Glucose (o) and insulin (p) tolerance tests at 5 weeks. q, Relative fTreg cell enrichment of mice fed HFD with rosiglitazone versus mice fed HFD alone at 5 and at 11 weeks. Data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test.
Extended Data Figure 7 Increased TNFα levels and gene expression pattern of aged fTreg knockout adipose tissue is consistent with an improved adipose remodelling capacity.
a, TNFα levels quantified by ELISA of whole adipose lysate (~40 weeks, n = 6 per group). b–d, FPKM values of all differentially expressed genes (b), differentially expressed collagens (c) and differentially expressed extracellular matrix proteases (d) in VAT from aged fTreg knockout and control mice (~40 weeks, n = 3 mice per group). Data are mean ± s.e.m. ***P < 0.001, Student’s t-test.
Extended Data Figure 8 fTreg cell gene expression and depletion with anti-ST2 antibody treatment.
a, Expression of several canonical Treg cell genes across fat and splenic Treg and fat and splenic Tconv cells. Cells were pooled from 3 and 4 mice before isolating RNA for subsequent RNA-Seq analysis. b, Expression of St2 across all haematopoietic cells catalogued in the ImmGen database. Position of adipose CD4+ CD25+ T cells is marked. c, Total weight before beginning course of anti-ST2 or isotype control antibodies (day 0) and after terminal analysis (day 3) (n = 4 mice per group). d, e, Spleen weight (d) and percentage of splenic naive CD4+ T cells as defined by CD62hi CD44lo relative to total splenic CD45+ CD4+ CD25− T cell population (e) of mice after terminal analysis (day 3, n = 4 mice per group). Data are mean ± s.e.m.
Extended Data Figure 9 Aged fTreg knockout mice are resistant to short-term, but not persistent, HFD-induced weight gain and IR.
a–e, Aged control and fTreg knockout mice were placed on HFD and monitored throughout course of diet for weight (a), fasting glucose levels (b), fasting serum insulin levels (c), performance on glucose tolerance test (d), and on insulin tolerance test (e). (Control, n = 10; fTreg KO, n = 11; mice were aged 27–29 weeks and weight-matched before HFD was introduced.) Data are mean ± s.e.m. ns, non-significant, *P < 0.05, **P < 0.01, Student’s t-test.
Rights and permissions
About this article
Cite this article
Bapat, S., Myoung Suh, J., Fang, S. et al. Depletion of fat-resident Treg cells prevents age-associated insulin resistance. Nature 528, 137–141 (2015). https://doi.org/10.1038/nature16151
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16151
This article is cited by
-
Type 1 interferons and Foxo1 down-regulation play a key role in age-related T-cell exhaustion in mice
Nature Communications (2024)
-
Aging adipose tissue, insulin resistance, and type 2 diabetes
Biogerontology (2024)
-
Intermittent fasting promotes rejuvenation of immunosenescent phenotypes in aged adipose tissue
GeroScience (2024)
-
A Treg-specific long noncoding RNA maintains immune-metabolic homeostasis in aging liver
Nature Aging (2023)
-
Epithelial cell-derived cytokine TSLP activates regulatory T cells by enhancing fatty acid uptake
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.