Abstract
HIV-1 Nef, a protein important for the development of AIDS, has well-characterized effects on host membrane trafficking and receptor downregulation. By an unidentified mechanism, Nef increases the intrinsic infectivity of HIV-1 virions in a host-cell-dependent manner. Here we identify the host transmembrane protein SERINC5, and to a lesser extent SERINC3, as a potent inhibitor of HIV-1 particle infectivity that is counteracted by Nef. SERINC5 localizes to the plasma membrane, where it is efficiently incorporated into budding HIV-1 virions and impairs subsequent virion penetration of susceptible target cells. Nef redirects SERINC5 to a Rab7-positive endosomal compartment and thereby excludes it from HIV-1 particles. The ability to counteract SERINC5 was conserved in Nef encoded by diverse primate immunodeficiency viruses, as well as in the structurally unrelated glycosylated Gag from murine leukaemia virus. These examples of functional conservation and convergent evolution emphasize the fundamental importance of SERINC5 as a potent anti-retroviral factor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Sequence Read Archive
Referenced accessions
GenBank/EMBL/DDBJ
Data deposits
RNA-seq fatsq data have been deposited in NCBI Sequence Read Archive (SRA) under accession code SRP062444.
References
Kestler, H. W. Importance of the nef gene for maintenance of high virus loads and for development of AIDS. Cell 65, 651–662 (1991)
Deacon, N. J. et al. Genomic structure of an attenuated quasi species of HIV-1 from a blood transfusion donor and recipients. Science 270, 988–991 (1995)
Kirchhoff, F., Greenough, T. C., Brettler, D. B., Sullivan, J. L. & Desrosiers, R. C. Brief report: absence of intact nef sequences in a long-term survivor with nonprogressive HIV-1 infection. N. Engl. J. Med. 332, 228–232 (1995)
Landi, A., Iannucci, V., Nuffel, A. V., Meuwissen, P. & Verhasselt, B. One protein to rule them all: modulation of cell surface receptors and molecules by HIV Nef. Curr. HIV Res. 9, 496–504 (2011)
Baur, A. S. et al. HIV-1 Nef leads to inhibition or activation of T cells depending on its intracellular localization. Immunity 1, 373–384 (1994)
Schrager, J. A. & Marsh, J. W. HIV-1 Nef increases T cell activation in a stimulus-dependent manner. Proc. Natl Acad. Sci. USA 96, 8167–8172 (1999)
Alexander, L., Du, Z., Rosenzweig, M., Jung, J. U. & Desrosiers, R. C. A role for natural simian immunodeficiency virus and human immunodeficiency virus type 1 nef alleles in lymphocyte activation. J. Virol. 71, 6094–6099 (1997)
Simmons, A., Aluvihare, V. & McMichael, A. Nef triggers a transcriptional program in T cells imitating single-signal T cell activation and inducing HIV virulence mediators. Immunity 14, 763–777 (2001)
Stolp, B. et al. HIV-1 Nef interferes with host cell motility by deregulation of Cofilin. Cell Host Microbe 6, 174–186 (2009)
Chowers, M. Y. et al. Optimal infectivity in vitro of human immunodeficiency virus type 1 requires an intact nef gene. J. Virol. 68, 2906–2914 (1994)
Munch, J. et al. Nef-mediated enhancement of virion infectivity and stimulation of viral replication are fundamental properties of primate lentiviruses. J. Virol. 81, 13852–13864 (2007)
Carl, S. et al. Modulation of different human immunodeficiency virus type 1 Nef functions during progression to AIDS. J. Virol. 75, 3657–3665 (2001)
Aiken, C. & Trono, D. Nef stimulates human immunodeficiency virus type 1 proviral DNA synthesis. J. Virol. 69, 5048–5056 (1995)
Chowers, M. Y., Pandori, M. W., Spina, C. A., Richman, D. D. & Guatelli, J. C. The growth advantage conferred by HIV-1 nef is determined at the level of viral DNA formation and is independent of CD4 downregulation. Virology 212, 451–457 (1995)
Schwartz, O., Marechal, V., Danos, O. & Heard, J. M. Human immunodeficiency virus type 1 Nef increases the efficiency of reverse transcription in the infected cell. J. Virol. 69, 4053–4059 (1995)
Pizzato, M. MLV glycosylated-Gag is an infectivity factor that rescues Nef-deficient HIV-1. Proc. Natl Acad. Sci. USA 107, 9364–9369 (2010)
Usami, Y., Popov, S. & Gottlinger, H. G. The Nef-Like effect of murine leukemia virus glycosylated Gag on HIV-1 infectivity is mediated by its cytoplasmic domain and depends on the AP-2 adaptor complex. J. Virol. 88, 3443–3454 (2014)
Grossman, T. R., Luque, J. M. & Nelson, N. Identification of a ubiquitous family of membrane proteins and their expression in mouse brain. J. Exp. Biol. 203, 447–457 (2000)
Xu, J. et al. Cloning and expression of a novel human C5orf12 gene, a member of the TMS_TDE family. Mol. Biol. Rep. 30, 47–52 (2003)
Inuzuka, M., Hayakawa, M. & Ingi, T. Serinc, an activity-regulated protein family, incorporates serine into membrane lipid synthesis. J. Biol. Chem. 280, 35776–35783 (2005)
Craig, H. M., Pandori, M. W. & Guatelli, J. C. Interaction of HIV-1 Nef with the cellular dileucine-based sorting pathway is required for CD4 down-regulation and optimal viral infectivity. Proc. Natl Acad. Sci. USA 95, 11229–11234 (1998)
Pizzato, M. et al. Dynamin 2 is required for the enhancement of HIV-1 infectivity by Nef. Proc. Natl Acad. Sci. USA 104, 6812–6817 (2007)
Saksela, K., Cheng, G. & Baltimore, D. Proline-rich (PxxP) motifs in HIV-1 Nef bind to SH3 domains of a subset of Src kinases and are required for the enhanced growth of Nef+ viruses but not for down-regulation of CD4. EMBO J. 14, 484–491 (1995)
Piguet, V. et al. Nef-induced CD4 degradation: a diacidic-based motif in Nef functions as a lysosomal targeting signal through the binding of β-COP in endosomes. Cell 97, 63–73 (1999)
Miller, M. D., Warmerdam, M. T., Page, K. A., Feinberg, M. B. & Greene, W. C. Expression of the human immunodeficiency virus type 1 (HIV-1) nef gene during HIV-1 production increases progeny particle infectivity independently of gp160 or viral entry. J. Virol. 69, 579–584 (1995)
Aiken, C. Pseudotyping human immunodeficiency virus type 1 (HIV-1) by the glycoprotein of vesicular stomatitis virus targets HIV-1 entry to an endocytic pathway and suppresses both the requirement for Nef and the sensitivity to cyclosporin A. J. Virol. 71, 5871–5877 (1997)
Chazal, N., Singer, G., Aiken, C., Hammarskjold, M. L. & Rekosh, D. Human immunodeficiency virus type 1 particles pseudotyped with envelope proteins that fuse at low pH no longer require Nef for optimal infectivity. J. Virol. 75, 4014–4018 (2001)
Pizzato, M., Popova, E. & Gottlinger, H. G. Nef can enhance the infectivity of receptor-pseudotyped human immunodeficiency virus type 1 particles. J. Virol. 82, 10811–10819 (2008)
Luo, T., Douglas, J. L., Livingston, R. L. & Garcia, J. V. Infectivity enhancement by HIV-1 Nef is dependent on the pathway of virus entry: implications for HIV-based gene transfer systems. Virology 241, 224–233 (1998)
Lai, R. P. et al. Nef decreases HIV-1 sensitivity to neutralizing antibodies that target the membrane-proximal external region of TMgp41. PLoS Pathog. 7, e1002442 (2011)
Usami, Y. & Gottlinger, H. HIV-1 Nef responsiveness is determined by env variable regions involved in trimer association and correlates with neutralization sensitivity. Cell Rep. 5, 802–812 (2013)
Campbell, E. M., Nunez, R. & Hope, T. J. Disruption of the actin cytoskeleton can complement the ability of Nef to enhance human immunodeficiency virus type 1 infectivity. J. Virol. 78, 5745–5755 (2004)
Schaeffer, E., Geleziunas, R. & Greene, W. C. Human immunodeficiency virus type 1 Nef functions at the level of virus entry by enhancing cytoplasmic delivery of virions. J. Virol. 75, 2993–3000 (2001)
Zhou, J. & Aiken, C. Nef enhances human immunodeficiency virus type 1 infectivity resulting from intervirion fusion: evidence supporting a role for Nef at the virion envelope. J. Virol. 75, 5851–5859 (2001)
Tobiume, M., Lineberger, J. E., Lundquist, C. A., Miller, M. D. & Aiken, C. Nef does not affect the efficiency of human immunodeficiency virus type 1 fusion with target cells. J. Virol. 77, 10645–10650 (2003)
Cavrois, M., Neidleman, J., Yonemoto, W., Fenard, D. & Greene, W. C. HIV-1 virion fusion assay: uncoating not required and no effect of Nef on fusion. Virology 328, 36–44 (2004)
Day, J. R., Munk, C. & Guatelli, J. C. The membrane-proximal tyrosine-based sorting signal of human immunodeficiency virus type 1 gp41 is required for optimal viral infectivity. J. Virol. 78, 1069–1079 (2004)
Cavrois, M., De Noronha, C. & Greene, W. C. A sensitive and specific enzyme-based assay detecting HIV-1 virion fusion in primary T lymphocytes. Nature Biotechnol. 20, 1151–1154 (2002)
van der Aa, L. M. et al. A large new subset of TRIM genes highly diversified by duplication and positive selection in teleost fish. BMC Biol. 7, 7 (2009)
Duggal, N. K. & Emerman, M. Evolutionary conflicts between viruses and restriction factors shape immunity. Nature Rev. Immunol. 12, 687–695 (2012)
Briggs, J. A., Wilk, T., Welker, R., Kräusslich, H. G. & Fuller, S. D. Structural organization of authentic, mature HIV‐1 virions and cores. EMBO J. 22, 1707–1715 (2003)
Cohen, F. S. & Melikyan, G. B. The energetics of membrane fusion from binding, through hemifusion, pore formation, and pore enlargement. J. Membr. Biol. 199, 1–14 (2004)
Razinkov, V. I. & Cohen, F. S. Sterols and sphingolipids strongly affect the growth of fusion pores induced by the hemagglutinin of influenza virus. Biochemistry 39, 13462–13468 (2000)
Ciechonska, M. & Duncan, R. Lysophosphatidylcholine reversibly arrests pore expansion during syncytium formation mediated by diverse viral fusogens. J. Virol. 88, 6528–6531 (2014)
Chen, A. et al. Fusion-pore expansion during syncytium formation is restricted by an actin network. J. Cell Sci. 121, 3619–3628 (2008)
Zufferey, R., Nagy, D., Mandel, R. J., Naldini, L. & Trono, D. Multiply attenuated lentiviral vector achieves efficient gene delivery in vivo. Nature Biotechnol. 15, 871–875 (1997)
Pizzato, M. et al. A one-step SYBR Green I-based product-enhanced reverse transcriptase assay for the quantitation of retroviruses in cell culture supernatants. J. Virol. Methods 156, 1–7 (2009)
Simon, J. H., Gaddis, N. C., Fouchier, R. A. & Malim, M. H. Evidence for a newly discovered cellular anti-HIV-1 phenotype. Nature Med. 4, 1397–1400 (1998)
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature Methods 5, 621–628 (2008)
Acknowledgements
We thank the Centre for AIDS Reagents, NIBSC, and NIH AIDS Research and Reference Reagent Program, Division of AIDS, for cell lines, plasmids and antibodies. We thank V. Adami and the CIBIO high-throughput screening and the Advanced Imaging facilities staff for technical assistance, G. De Silvestro, G. Mattiuzzo, C. Reinhard and L. Conti for reagents, G. Melikian, S. Basmaciogullari, P. Cherepanov, O. Fackler, N. Segata, F. Demichelis, A. Marcello, T. Fedrizzi and A. Helander for critical discussions. This work was supported by the Biotechnology Program of University of Trento, FP7 Marie Curie Career Integration grant number 322130 and Caritro ‘Ricerca Biomedica’ grant number 2013.0248 to M.P., National Institute of Health grant DP1DA034990 to J.L. and European Research Council grant 249968 to S.E.A.
Author information
Authors and Affiliations
Contributions
A.R., A.C., S.Z., V.D.S., R.B., S.E.A., J.L., F.A.S. and M.P. designed the experiments. A.R., S.Z., A.C., V.D.S., R.B., S.L.G., S.M.M., A.N., F.A.S. and M.P. performed the experiments. All authors contributed to the assembly and writing of the manuscript. A.R., A.C. and S.Z. contributed equally to the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
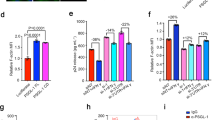
Extended Data Figure 1 SERINC5 is an inhibitor of HIV-1 infectivity.
a, Mapping of the INDELS in the genomic locus spanning SERINC5 exon 2 in JTAg cell clonal populations from Fig. 2a. b, Infectivity of HIV-1 from JTAg cells stably transduced with lentiCRISPR targeting GFP or SERINC5 in three different exons (n = 4, experiment replicated twice). c, Relative expression of SERINC5 in primary cells and in cell lines measured by qPCR normalized by expression of ACTB (n = 3). d, Infectivity of HIV-1 from the indicated cell lines expressing SERINC5 (n = 4, experiments were replicated twice). Mean ± s.d., unpaired two-tailed t-test, ***P < 0.001 e, Expression levels of the five SERINC genes in JTAg cells obtained from RNA-seq.
Extended Data Figure 2 Nef and glycoGag expression result in relocalization of SERINC5 to an endosomal compartment and prevent its incorporation into virions.
a, Single round Nef-defective NL4-3 produced by cotransfection of HEK293T cells with plasmids expressing Nef proteins or the empty vector control, and PBJ6-SERINC5–HA: immunoblotting of virions and cell lysates from producer cells. b, Immunofluorescence staining of JTAg cells transfected to express SERINC5–GFP, Nef–HA from HIV-1 isolate 97ZA012 (clade C), from SIVmac239, HA–glycoGag or an empty vector control. Scale bar, 10 μm.
Extended Data Figure 3 SERINC5 inhibits cytoplasmic delivery of virion content.
a, Immunodetection of Cre-recombinase (38 kDa) and p24 in HIV-1 particles. b, Effect of 1 μM AZT or 100 nM T20 on Cre-delivery and virus infectivity (TU, transducing units). c, Immunoblotting of HIV-1 virus particles produced from HEK293T expressing increasing levels of SERINC5–HA. d, Effect of SERINC5 on virus fusion measured with BLAM assay T20 served as a negative control. (n = 4, experiment replicated twice). e, Cre delivery by EBOV-GP pseudotyped HIV-1 particles. f, Inhibition of Cre delivery and counteraction by Nef on HIV-1 from HEK293T expressing SERINC5. Mean ± s.d., n = 4, unpaired two-tailed t-test, *P < 0.05, **P < 0.01, ***P < 0.001. Scale bar, 100 μm.
Extended Data Figure 4 SERINC3 and SERINC5 expression is not induced by interferon nor LPS treatments.
a–d, Relative gene expression levels of SERINC3, SERINC5 and CXCL10 in response to treatment with IFN-β and LPS in Jurkat (a), monocyte-derived dendritic cells from two donors (MDDC, b), CD4+ primary T cells unstimulated (c) or stimulated with PHA (d) from two donors. Expression of the housekeeping gene OAZ1 was used as a normalization control. Mean ± s.d., n = 3.
Rights and permissions
About this article
Cite this article
Rosa, A., Chande, A., Ziglio, S. et al. HIV-1 Nef promotes infection by excluding SERINC5 from virion incorporation. Nature 526, 212–217 (2015). https://doi.org/10.1038/nature15399
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15399
This article is cited by
-
HIV-1 subtype C Nef-mediated SERINC5 down-regulation significantly contributes to overall Nef activity
Retrovirology (2023)
-
HIV-1 restriction by SERINC5
Medical Microbiology and Immunology (2023)
-
Serinc2 deficiency causes susceptibility to sepsis-associated acute lung injury
Journal of Inflammation (2022)
-
Cul3-KLHL20 E3 ubiquitin ligase plays a key role in the arms race between HIV-1 Nef and host SERINC5 restriction
Nature Communications (2022)
-
Nef inhibits HIV transcription and gene expression in astrocytes and HIV transmission from astrocytes to CD4+ T cells
Journal of NeuroVirology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.