Abstract
Cells require nucleotides to support DNA replication and repair damaged DNA. In addition to de novo synthesis, cells recycle nucleotides from the DNA of dying cells or from cellular material ingested through the diet. Salvaged nucleosides come with the complication that they can contain epigenetic modifications. Because epigenetic inheritance of DNA methylation mainly relies on copying of the modification pattern from parental strands1,2,3, random incorporation of pre-modified bases during replication could have profound implications for epigenome fidelity and yield adverse cellular phenotypes. Although the salvage mechanism of 5-methyl-2′deoxycytidine (5mdC) has been investigated before4,5,6, it remains unknown how cells deal with the recently identified oxidized forms of 5mdC: 5-hydroxymethyl-2′deoxycytidine (5hmdC), 5-formy-2′deoxycytidine (5fdC) and 5-carboxyl-2′deoxycytidine (5cadC)7,8,9,10. Here we show that enzymes of the nucleotide salvage pathway display substrate selectivity, effectively protecting newly synthesized DNA from the incorporation of epigenetically modified forms of cytosine. Thus, cell lines and animals can tolerate high doses of these modified cytidines without any deleterious effects on physiology. Notably, by screening cancer cell lines for growth defects after exposure to 5hmdC, we unexpectedly identify a subset of cell lines in which 5hmdC or 5fdC administration leads to cell lethality. Using genomic approaches, we show that the susceptible cell lines overexpress cytidine deaminase (CDA). CDA converts 5hmdC and 5fdC into variants of uridine that are incorporated into DNA, resulting in accumulation of DNA damage, and ultimately, cell death. Our observations extend current knowledge of the nucleotide salvage pathway by revealing the metabolism of oxidized epigenetic bases, and suggest a new therapeutic option for cancers, such as pancreatic cancer, that have CDA overexpression and are resistant to treatment with other cytidine analogues11.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Riggs, A. D. X inactivation, differentiation, and DNA methylation. Cytogenet. Cell Genet. 14, 9–25 (1975)
Wigler, M., Levy, D. & Perucho, M. The somatic replication of DNA methylation. Cell 24, 33–40 (1981)
Gruenbaum, Y., Cedar, H. & Razin, A. Substrate and sequence specificity of a eukaryotic DNA methylase. Nature 295, 620–622 (1982)
Vilpo, J. A. & Vilpo, L. M. Nucleoside monophosphate kinase may be the key enzyme preventing salvage of DNA 5-methylcytosine. Mutat. Res. 286, 217–220 (1993)
Jekunen, A., Puukka, M. & Vilpo, J. Exclusion of exogenous 5-methyl-2′-deoxycytidine from DNA in human leukemic cells. A study with [2(-14)C]- and [methyl-14C]5-methyl-2′-deoxycytidine. Biochem. Pharmacol. 32, 1165–1168 (1983)
Jekunen, A. & Vilpo, J. A. 5-Methyl-2′-deoxycytidine. Metabolism and effects on cell lethality studied with human leukemic cells in vitro. Mol. Pharmacol. 25, 431–435 (1984)
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009)
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009)
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011)
Pfaffeneder, T. et al. The discovery of 5-formylcytosine in embryonic stem cell DNA. Angew. Chem. 50, 7008–7012 (2011)
Frese, K. K. et al. nab-Paclitaxel potentiates gemcitabine activity by reducing cytidine deaminase levels in a mouse model of pancreatic cancer. Cancer Disov. 2, 260–269 (2012)
Loffler, M., Fairbanks, L. D., Zameitat, E., Marinaki, A. M. & Simmonds, H. A. Pyrimidine pathways in health and disease. Trends Mol. Med. 11, 430–437 (2005)
Hershey, H. V., Stieber, J. F. & Mueller, G. C. DNA synthesis in isolated HeLa nuclei. A system for continuation of replication in vivo. Eur. J. Biochem. 34, 383–394 (1973)
Arner, E. S. & Eriksson, S. Mammalian deoxyribonucleoside kinases. Pharmacol. Ther. 67, 155–186 (1995)
Boissan, M. et al. The mammalian Nm23/NDPK family: from metastasis control to cilia movement. Mol. Cell. Biochem. 329, 51–62 (2009)
Xu, Y., Johansson, M. & Karlsson, A. Human UMP-CMP kinase 2, a novel nucleoside monophosphate kinase localized in mitochondria. J. Biol. Chem. 283, 1563–1571 (2008)
Lin, K. T., Momparler, R. L. & Rivard, G. E. High-performance liquid chromatographic analysis of chemical stability of 5-aza-2′-deoxycytidine. J. Pharm. Sci. 70, 1228–1232 (1981)
Petitjean, A. et al. Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum. Mutat. 28, 622–629 (2007)
Ross, D. T. et al. Systematic variation in gene expression patterns in human cancer cell lines. Nature Genet. 24, 227–235 (2000)
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012)
Nabel, C. S. et al. AID/APOBEC deaminases disfavor modified cytosines implicated in DNA demethylation. Nature Chem. Biol. 8, 751–758 (2012)
Chung, S. J., Fromme, J. C. & Verdine, G. L. Structure of human cytidine deaminase bound to a potent inhibitor. J. Med. Chem. 48, 658–660 (2005)
Boorstein, R. J., Chiu, L. N. & Teebor, G. W. A mammalian cell line deficient in activity of the DNA repair enzyme 5-hydroxymethyluracil-DNA glycosylase is resistant to the toxic effects of the thymidine analog 5-hydroxymethyl-2'-deoxyuridine. Mol. Cell. Biol. 12, 5536–5540 (1992)
Weizman, N. et al. Macrophages mediate gemcitabine resistance of pancreatic adenocarcinoma by upregulating cytidine deaminase. Oncogene 33, 3812–3819 (2013)
Eliopoulos, N., Cournoyer, D. & Momparler, R. L. Drug resistance to 5-aza-2'-deoxycytidine, 2',2'-difluorodeoxycytidine, and cytosine arabinoside conferred by retroviral-mediated transfer of human cytidine deaminase cDNA into murine cells. Cancer Chemother. Pharmacol. 42, 373–378 (1998)
Qin, T. et al. Mechanisms of resistance to decitabine in the myelodysplastic syndrome. PLoS ONE 6, e23372 (2011)
Pei, H. et al. FKBP51 affects cancer cell response to chemotherapy by negatively regulating Akt. Cancer Cell 16, 259–266 (2009)
Shin, G. et al. GENT: gene expression database of normal and tumor tissues. Cancer Inform. 10, 149–157 (2011)
Gad, H. et al. MTH1 inhibition eradicates cancer by preventing sanitation of the dNTP pool. Nature 508, 215–221 (2014)
Huber, K. V. et al. Stereospecific targeting of MTH1 by (S)-crizotinib as an anticancer strategy. Nature 508, 222–227 (2014)
Ramsahoye, B. H. Nearest-neighbor analysis. Methods Mol. Biol. 200, 9–15 (2002)
Van Rompay, A. R., Johansson, M. & Karlsson, A. Phosphorylation of deoxycytidine analog monophosphates by UMP-CMP kinase: molecular characterization of the human enzyme. Mol. Pharmacol. 56, 562–569 (1999)
Vincenzetti, S., Cambi, A., Neuhard, J., Garattini, E. & Vita, A. Recombinant human cytidine deaminase: expression, purification, and characterization. Protein Expr. Purif. 8, 247–253 (1996)
Irwin, J. J., Sterling, T., Mysinger, M. M., Bolstad, E. S. & Coleman, R. G. ZINC: a free tool to discover chemistry for biology. J. Chem. Inf. Model. 52, 1757–1768 (2012)
Grosdidier, A., Zoete, V. & Michielin, O. SwissDock, a protein-small molecule docking web service based on EADock DSS. Nucleic Acids Res. 39, W270–W277 (2011)
Planck, S. R. & Mueller, G. C. DNA chain growth in isolated HeLa nuclei. Biochemistry 16, 2778–2782 (1977)
Kemmerich, K., Dingler, F. A., Rada, C. & Neuberger, M. S. Germline ablation of SMUG1 DNA glycosylase causes loss of 5-hydroxymethyluracil- and UNG-backup uracil-excision activities and increases cancer predisposition of Ung−/−Msh2−/− mice. Nucleic Acids Res. 40, 6016–6025 (2012)
Dietmair, S., Timmins, N. E., Gray, P. P., Nielsen, L. K. & Kromer, J. O. Towards quantitative metabolomics of mammalian cells: development of a metabolite extraction protocol. Anal. Biochem. 404, 155–164 (2010)
MacKenzie, C. J. & Shioda, T. COS-1 cells as packaging host for production of lentiviruses. Curr. Protocols Cell Biol. Chapter 26, Unit–26.27 (2011)
Pfister, T. D. et al. Topoisomerase I levels in the NCI-60 cancer cell line panel determined by validated ELISA and microarray analysis and correlation with indenoisoquinoline sensitivity. Mol. Cancer Ther. 8, 1878–1884 (2009)
Acknowledgements
We acknowledge Ludwig Cancer Research and BBSRC for funding; R. Klose, T. Milne, G. Bond, M. Muers and members of the Kriaucionis laboratory for the discussions and critical reading of the manuscript. M. Shipman for his technical assistance in image acquisition and analysis; I. Ratnayaka and R. Lisle for assistance with histology; S. Laird and J. Tanner for help with animal experiments; X. Lu, S. Lunardi and G. Bond for providing cell lines; P. Filippakopoulos for advice on molecular docking. B.M.K. was supported by the John Fell Fund 133/075 (BMK) and the Wellcome Trust 097813/Z/11/Z.
Author information
Authors and Affiliations
Contributions
M.Z. performed all experiments, with the following exceptions: mass spectrometry was done by G.B., M.-L.T., K.M.P. and B.M.K.; gene expression analysis was done by S.K.; analysis of tissue pathology was done by R.G. S.K. conceived the study, S.K. and M.Z. designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 DNA polymerase and nucleoside kinase activities on modified nucleosides.
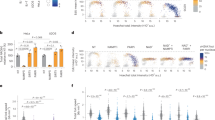
a, Mass spectrometry confirmation of 5hmdC, 5fdC and 5cadC in the purchased nucleosides. b, HPLC–UV chromatogram of nucleosides from DNA extracted from H1299 cells transfected with 5hmdCTP. The abundance of 5hmdC relative to dG is illustrated in the right panel (n = 3). n.d., not detected. Error bars denote s.d. c, Coomassie-stained SDS–PAGE gel of recombinant purified DCK and CMPK1 enzymes used in the study. d, Two-dimensional TLC images of DCK reaction products. Dotted lines indicate reference points, which aid in tracking the migration localization of the nucleosides. The monophosphate in each reaction is circled in red (representative picture, n = 3). e, Schematic map of nucleoside migration on two-dimensional TLC plate (asterisk indicates a background spot coming from ATP and used as a reference point)
Extended Data Figure 2 Stability of the nucleosides and CDA activity.
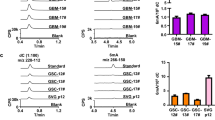
a, b, Quantification of nucleosides by HPLC–UV during 10 days of incubation in water (a) and DMEM (b) at 37 °C (n = 3). c, Representative HPLC–UV chromatograms at days 0, 2 and 10 with retention times indicated above each peak. d, Cell lines used in the study and their characteristics. e, Western blot showing knockdown of CDA by shRNA in the SN12C cell line. Right panel illustrates the growth of the cell line during treatment with 10 μM 5hmdC (n = 3). f, Western blot showing expression of CDA in wild-type and lentivirally transduced MCF7 cell line. Growth curve after treatment with 10 μM 5hmdC is shown on the right (n = 3). g, Coomassie-stained SDS–PAGE gel of recombinant purified CDA enzyme used in this study. h, HPLC–UV chromatograms showing the retention times and identity of substrates and CDA-catalysed products. i, List of Km, kcat and vmax values of catalytic activity of CDA catalysing the deamination of cytidine variants. All error bars denote s.d.
Extended Data Figure 3 Mechanism of CDA catalysed deamination of epigenetic nucleosides, their cytotoxicity and dUTPase activity.
a, Molecular docking of dC, 5hmdC and 5fdC on the CDA active site (Protein Data Bank (PDB) accession 1MQ0). The detailed view of the catalytic pocket is shown with the modified nucleoside in the centre. Chains A, B and C indicate units of the tetramer, which CDA forms to deaminate four nucleosides. Thin yellow lines show compatible distances for the formation of hydrogen bonds. b, Growth curves of H1299 and MCF7 cell lines treated with 10 and 1 μM of dC, 5hmdU and 5fdU over a period of 10 days (n = 3). c, Coomassie-stained gel demonstrating recombinant purified DUT (molecular mass, 18 kDa) and in vitro measurements of dUTPase activity using non-canonical uridine triphosphates (n = 3). d, Extracted ion chromatogram of nucleoside standards analysed by HPLC–QTOF mass spectrometry. Each nucleoside intensity was measured using the merged m/z values of the [M+H]+, [M+Na]+, [M+H]+, [2M+H]+ and [base+H]+ and a symmetric single m/z expansion of ±0.02. e, The most prominent ion of 5hmdU was identified in 5hmdC-treated MDA-MB-231 cells. All error bars denote s.d.
Extended Data Figure 4 Mass spectrometry identification of 5fUra and ultraviolet quantification of 5hmdU in the DNA.
a, Extracted ion chromatogram of nucleoside standards with 5fdU analysed by HPLC–QTOF mass spectrometry (as in Extended Data Fig. 3d). b, Weak, but consistent signal of 5fUra is identified in DNA of 5fdC-treated MDA-MB-231 cells, but not dC-treated cells or buffer alone. Two representative examples are shown. c, Relative quantification of 5fUra signal from three biological mass spectrometry replicates. d, Relationship between measured 5hmdU/T in the DNA of cell lines treated with 10 μM 5hmdC for 3 days and CDA expression levels. The cell lines used in this study are in coloured font (n = 3). All error bars denote s.d.
Extended Data Figure 5 Effect of 5hmdC administration on the cell cycle and DNA damage.
a, b, Propidium iodide FACS assay of the cell cycle. Shown are two representative plots of MDA-MB-231 cells at day 3 of treatment with dC and 5hmdC (10 μM) (a) and quantification for all the cell lines analysed (n = 3) (b). Two-way ANOVA: P = 0.0027 (S: 5hmdC versus dC MDA-MB-231), P = 0.0149 (G2-M: 5hmdC versus dC MDA-MB-231). HOP-92 P < 0.0001, P = 0.0005 (S: 5hmdC versus dC Capan-2), P < 0.0001 (G2-M: 5hmdC versus dC Capan-2) (n = 3; 10,000 events acquired). c, γH2AX immunofluorescence in MDA-MB-231 and H1299 cell lines at day 3 after treatment with 10 μM 5hmdC or dC. Scale bar, 50 μm. d, Fraction of cells showing a γH2AX signal above background (n = 3). ANOVA with Sidak correction for multiple comparisons: P = 0.0208 (5hmdC versus dC MDA-MB-231), P = 0.0135 (5hmdC versus dC HOP-92). Error bars denote s.d.
Extended Data Figure 6 Quantification of intracellular nucleotides by ion-pair HPLC and SMUG1 glycosylase activity.
a, Illustrative chromatogram of all standards indicated in b mixed together. b, Retention times of nucleotides were determined by analysing each standard separately and are indicated in the table. c, An average relative abundance of NTP and dNTP levels in cells treated with dC, 5hmdC and 5fdC (n = 3). d, Representative chromatograms of indicated experiments (blue) overlaid with standards separated on the same run (red). e, Typical image of denaturing PAGE electrophoresis of DNA incubated with SMUG1 and cleaved with APE1. f, Quantification of the DNA oligonucleotides with excised bases. g, Expression of SMUG1 and uracil DNA glycosylase (UNG) in MDA_MB_231, SN12C and Capan-2 cell lines (Genevestigator). Error bars denote s.d.
Extended Data Figure 7 CDA expression in human cancer and normal tissues, and toxicity evaluation of 5hmdC and 5fdC in mice.
a, CDA overexpression in pancreatic cancer (t-test, P < 0.0001). b, CDA expression across a panel of cancer (red) versus normal (green) tissues (GENT database). Arrows indicate cancer types with an evident difference between normal (N) and cancerous tissues (C). c, 5hmdC and 5fdC detection in the blood (mass spectrometry) of intraperitoneally injected mice at 30 min after injection. d, Label-free mass spectrometry quantification of 5hmdC in the blood of animals injected with doses of 25, 50 and 100 mg kg−1 (n = 3 (100 mg ml−1) and n = 4 (25 and 50 mg ml−1)). Error bars denote s.e.m. e, Immunohistochemistry showing CDA expression in the intestine. f, Haematoxylin and eosin staining of the intestine of mice injected with PBS and 100 mg kg−1 of 5hmdC and 5fdC. Tissue was removed 5 days after the injection. g, Immunofluorescence evaluation of proliferation (H3PS10) and DNA damage (γH2AX) in the intestine of mice treated with PBS and 100 mg kg−1 of 5hmdC and 5fdC 5 days after treatment. In parallel, the protocol was done on testis of irradiated mice, where positive signals for γH2AX were observed (data not shown). Scale bars, 50 μm (e–g). h, Weight of the mice plotted over the treatment period (n = 16 per group).
Extended Data Figure 8 Evaluation of wild-type SN12C cell line and CDA knockdown in a mouse xenograft model.
a, Schematic illustration of xenograft establishment and treatment with nucleoside variants. b, Tumour diameter was measured by Vernier caliper and volume calculated by assuming that tumours were spheres (n = 8, two-way ANOVA with repeated measures and Holm–Sidak correction, P < 0.0001). c, Photos of the dissected tumours (asterisks indicate dissected lymph nodes found after histological analysis). d, Western blot showing CDA expression in tumours extracted from mice. e, Quantification of proliferation (H3PS10) and DNA damage (γH2AX) using confocal microscopy and ImageJ of the central section of the tumour. Scale bar, 50 μm (n = 4, one-way ANOVA, SN12C H3PS10: P = 0.0033 (PBS versus 5hmdC), P = 0.0046 (PBS versus 5fdC); γH2AX: P = 0.0003 (PBS versus 5hmdC), P = 0.0436 (PBS versus 5fdC); SN12CshCDA_8: P = 0.0130 (PBS versus 5hmdC)). f, 5hmdU quantified from a HPLC–UV chromatogram of nucleosides from DNA extracted from tumours of mice treated with 5hmdC and PBS (n = 4, one-way ANOVA P = 0.0041). Error bars denote s.d.
Extended Data Figure 9 Identification and quantification of compounds' resulting peaks in HPLC–UV.
a, The abundance of molecule eluting at 5.1 min (5.7 min on the HPLC–QTOF) is not significantly different between dC- and 5hmdC-treated samples. It is a common component of DNA hydrolysis buffer. b, 5-methylcytosine in the DNA does not change after treatment with 5hmdC. Identity of 5mdC in the samples was confirmed by HPLC–QTOF mass spectrometry.
Extended Data Figure 10 Identification and quantification of compounds' resulting peaks in HPLC–UV.
Compound eluting at 4.5 min (5.0 min on the HPLC–QTOF) is an abundant component of DNA hydrolysis buffer, generating a m/z of 202.18.
Supplementary information
Supplementary Table 1
This table contains the gene expression analysis of 5hmdC sensitive and resistant cell lines. (XLSX 9160 kb)
Supplementary Table 2
This table contains the Mass spectrometry data of nucleosides. (XLSX 9 kb)
Rights and permissions
About this article
Cite this article
Zauri, M., Berridge, G., Thézénas, ML. et al. CDA directs metabolism of epigenetic nucleosides revealing a therapeutic window in cancer. Nature 524, 114–118 (2015). https://doi.org/10.1038/nature14948
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14948
This article is cited by
-
Upregulation of cytidine deaminase in NAT1 knockout breast cancer cells
Journal of Cancer Research and Clinical Oncology (2023)
-
FANCD2 maintains replication fork stability during misincorporation of the DNA demethylation products 5-hydroxymethyl-2’-deoxycytidine and 5-hydroxymethyl-2’-deoxyuridine
Cell Death & Disease (2022)
-
DNA methylome and single-cell transcriptome analyses reveal CDA as a potential druggable target for ALK inhibitor–resistant lung cancer therapy
Experimental & Molecular Medicine (2022)
-
Metabolic syndrome related gene signature predicts the prognosis of patients with pancreatic ductal carcinoma. A novel link between metabolic dysregulation and pancreatic ductal carcinoma
Cancer Cell International (2021)
-
Chemotherapy-induced CDA expression renders resistant non-small cell lung cancer cells sensitive to 5′-deoxy-5-fluorocytidine (5′-DFCR)
Journal of Experimental & Clinical Cancer Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.