Abstract
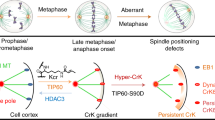
Cell division requires the precise coordination of chromosome segregation and cytokinesis. This coordination is achieved by the recruitment of an actomyosin regulator, Ect2, to overlapping microtubules at the centre of the elongating anaphase spindle1. Ect2 then signals to the overlying cortex to promote the assembly and constriction of an actomyosin ring between segregating chromosomes1. Here, by studying division in proliferating Drosophila and human cells, we demonstrate the existence of a second, parallel signalling pathway, which triggers the relaxation of the polar cell cortex at mid anaphase. This is independent of furrow formation, centrosomes and microtubules and, instead, depends on PP1 phosphatase and its regulatory subunit Sds22 (refs 2, 3). As separating chromosomes move towards the polar cortex at mid anaphase, kinetochore-localized PP1–Sds22 helps to break cortical symmetry by inducing the dephosphorylation and inactivation of ezrin/radixin/moesin proteins at cell poles. This promotes local softening of the cortex2,3, facilitating anaphase elongation and orderly cell division. In summary, this identifies a conserved kinetochore-based phosphatase signal and substrate, which function together to link anaphase chromosome movements to cortical polarization, thereby coupling chromosome segregation to cell division.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Green, R. A., Paluch, E. & Oegema, K. Cytokinesis in animal cells. Annu. Rev. Cell Dev. Biol. 28, 29–58 (2012)
Kunda, P. et al. PP1-mediated moesin dephosphorylation couples polar relaxation to mitotic exit. Curr. Biol. 22, 231–236 (2012)
Roubinet, C. et al. Molecular networks linked by Moesin drive remodeling of the cell cortex during mitosis. J. Cell Biol. 195, 99–112 (2011)
Matthews, H. K. et al. Changes in Ect2 localization couple actomyosin-dependent cell shape changes to mitotic progression. Dev. Cell 23, 371–383 (2012)
Sedzinski, J. et al. Polar actomyosin contractility destabilizes the position of the cytokinetic furrow. Nature 476, 462–466 (2011)
Roegiers, F., Younger-Shepherd, S., Jan, L. Y. & Jan, Y. N. Two types of asymmetric divisions in the Drosophila sensory organ precursor cell lineage. Nature Cell Biol. 3, 58–67 (2001)
Georgiou, M. & Baum, B. Polarity proteins and Rho GTPases cooperate to spatially organise epithelial actin-based protrusions. J. Cell Sci. 123, 1089–1098 (2010)
Hickson, G. R., Echard, A. & O'Farrell, P. H. Rho-kinase controls cell shape changes during cytokinesis. Curr. Biol. 16, 359–370 (2006)
Mishima, M., Kaitna, S. & Glotzer, M. Central spindle assembly and cytokinesis require a kinesin-like protein/RhoGAP complex with microtubule bundling activity. Dev. Cell 2, 41–54 (2002)
Lekomtsev, S. et al. Centralspindlin links the mitotic spindle to the plasma membrane during cytokinesis. Nature 492, 276–279 (2012)
Tse, Y. C., Piekny, A. & Glotzer, M. Anillin promotes astral microtubule-directed cortical myosin polarization. Mol. Biol. Cell 22, 3165–3175 (2011)
Fededa, J. P. & Gerlich, D. W. Molecular control of animal cell cytokinesis. Nature Cell Biol. 14, 440–447 (2012)
Murthy, K. & Wadsworth, P. Dual role for microtubules in regulating cortical contractility during cytokinesis. J. Cell Sci. 121, 2350–2359 (2008)
Werner, M., Munro, E. & Glotzer, M. Astral signals spatially bias cortical myosin recruitment to break symmetry and promote cytokinesis. Curr. Biol. 17, 1286–1297 (2007)
Blachon, S. et al. Drosophila asterless and vertebrate Cep152 Are orthologs essential for centriole duplication. Genetics 180, 2081–2094 (2008)
Hu, C. K., Coughlin, M., Field, C. M. & Mitchison, T. J. Cell polarization during monopolar cytokinesis. J. Cell Biol. 181, 195–202 (2008)
Lancaster, O. M. et al. Mitotic rounding alters cell geometry to ensure efficient bipolar spindle formation. Dev. Cell 25, 270–283 (2013)
Zhang, D. & Nicklas, R. B. 'Anaphase' and cytokinesis in the absence of chromosomes. Nature 382, 466–468 (1996)
Lesage, B., Qian, J. & Bollen, M. Spindle checkpoint silencing: PP1 tips the balance. Curr. Biol. 21, R898–R903 (2011)
Posch, M. et al. Sds22 regulates aurora B activity and microtubule-kinetochore interactions at mitosis. J. Cell Biol. 191, 61–74 (2010)
Wurzenberger, C. et al. Sds22 and Repo-Man stabilize chromosome segregation by counteracting Aurora B on anaphase kinetochores. J. Cell Biol. 198, 173–183 (2012)
Qian, J., Lesage, B., Beullens, M., Van Eynde, A. & Bollen, M. PP1/Repo-man dephosphorylates mitotic histone H3 at T3 and regulates chromosomal aurora B targeting. Curr. Biol. 21, 766–773 (2011)
Liu, D. et al. Regulated targeting of protein phosphatase 1 to the outer kinetochore by KNL1 opposes Aurora B kinase. J. Cell Biol. 188, 809–820 (2010)
Kennedy, M. J. et al. Rapid blue-light-mediated induction of protein interactions in living cells. Nature Methods 7, 973–975 (2010)
Turlier, H., Audoly, B., Prost, J. & Joanny, J. F. Furrow constriction in animal cell cytokinesis. Biophys. J. 106, 114–123 (2014)
Kiyomitsu, T. & Cheeseman, I. M. Cortical dynein and asymmetric membrane elongation coordinately position the spindle in anaphase. Cell 154, 391–402 (2013)
Dehapiot, B. & Halet, G. Ran GTPase promotes oocyte polarization by regulating ERM (Ezrin/Radixin/Moesin) inactivation. Cell Cycle 12, 1672–1678 (2013)
Edwards, K. A., Demsky, M., Montague, R. A., Weymouth, N. & Kiehart, D. P. GFP-moesin illuminates actin cytoskeleton dynamics in living tissue and demonstrates cell shape changes during morphogenesis in Drosophila. Dev. Biol. 191, 103–117 (1997)
Mummery-Widmer, J. L. et al. Genome-wide analysis of Notch signalling in Drosophila by transgenic RNAi. Nature 458, 987–992 (2009)
Matsumoto, K., Toh-e, A. & Oshima, Y. Genetic control of galactokinase synthesis in Saccharomyces cerevisiae: evidence for constitutive expression of the positive regulatory gene gal4 . J. Bacteriol. 134, 446–457 (1978)
Jeong, J. Y. et al. One-step sequence- and ligation-independent cloning as a rapid and versatile cloning method for functional genomics studies. Appl. Environ. Microbiol. 78, 5440–5443 (2012)
Hickson, G. R. & O'Farrell, P. H. Rho-dependent control of anillin behavior during cytokinesis. J. Cell Biol. 180, 285–294 (2008)
Théry, M. et al. The extracellular matrix guides the orientation of the cell division axis. Nature Cell Biol. 7, 947–953 (2005)
Lénárt, P. et al. The small-molecule inhibitor BI 2536 reveals novel insights into mitotic roles of polo-like kinase 1. Curr. Biol. 17, 304–315 (2007)
Liu, T., Sims, D. & Baum, B. Parallel RNAi screens across different cell lines identify generic and cell type-specific regulators of actin organization and cell morphology. Genome Biol. 10, R26 (2009)
Acknowledgements
N.T.L.R., S.L. and B.B. thank Cancer Research UK, and J.K.-V. the Medical Research Council for funding. S.J. and G.R.X.H. were funded by the Canadian Institutes of Health Research, the Canada Foundation for Innovation and a salary award from the Fonds de Recherche du Québec-Santé, and G.R.X.H. thanks the Cole Foundation for a Transition award. This study benefited from support from INCa and the BBSRC (BB/K009001/1). We thank M. Lam, E. Paluch, M. Petronczki, G. Salbreux and members of the Baum laboratory for input and critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
N.T.L.R. designed and conducted all experiments using flies and helped analyse human cell data with the aid of J.K.-V. S.L. designed and conducted all experiments using human cells. S.J. and G.R.X.H. conducted all experiments in fly cell culture. B.B. oversaw the project, which was conceived by N.T.L.R. and B.B. N.T.L.R., S.L. and B.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Depletion of RacGAP1 in SOP cells affects neither polar relaxation nor anaphase cell elongation.
a–c, Time-lapse imaging of SOP cells in control and RacGAP1-depleted backgrounds was carried out to analyse the relative timing of polar relaxation and anaphase elongation. We analysed cells from five control animals and four RacGAP1 RNAi animals. Representative images are shown in a (scale bars, 5 µm). White arrowheads indicate actin clearance at the poles. Lifeact–GFP was used to label F-actin. For control and RacGAP1 RNAi cells that formed a furrow (n = 19 and n = 16, respectively), the period between anaphase onset and furrow initiation or furrow completion is plotted in b (black lines denote the median). Cell length is plotted in c for the control group (18 cells) and for RacGAP1 RNAi cells that were delayed in furrow formation (15 cells) or that failed to form a furrow (5 cells). d, Relative levels of polar actin were compared across movies of 12 control RNAi cells from four animals (same as seen in Fig. 2e) and from 12 RacGAP1 RNAi cells from three animals. Ana, anaphase; Met, metaphase. e, Graph shows the ratio of levels of cortical actin at poles versus the equator at mid anaphase for 20 control cells from five animals, and for 13 RacGAP1 RNAi cells from three animals. Data are shown as mean and s.d. in c and d, and as box-and-whisker plots with 10–90 percentiles in e. A two-tailed unpaired t-test was used to calculate statistical significance; P > 0.05 was deemed not significant (n.s.).
Extended Data Figure 2 Depletion of RacGAP1 impairs myosin II equatorial accumulation and furrow ingression, but does not affect actin clearance from the poles at mid anaphase.
a, Western blot showing RacGAP1 depletion in HeLa cells. b, Representative DIC stills from movies show HeLa cells at indicated times after the onset of anaphase. Images show control cells before and after furrow initiation and RacGAP1-depleted cells at mid anaphase, as all fail cytokinesis. Arrowheads point to blebs. c, d, Graphs show levels of cortical myosin II (c) and cortical actin (d) in the polar and equatorial regions of cells. Levels were measured in HeLa cells expressing myosin-II–GFP and Utrophin–Ruby treated with control siRNA (15 cells, 3 experiments) or siRacGAP1 siRNA (oligonucleotide no. 4, seen in a) (15 cells, 3 experiments). Data are shown as mean and s.d. e, f, Representative images and corresponding kymographs taken from 12 time-lapse movies of fly SOP cells, fluorescently labelled for both myosin II (Sqh–mCherry) and actin filaments (GMA), undergoing anaphase. Note that the same cells were used for the analysis in Extended Data Fig. 9a. Anaphase onset = 0 s. Asterisks mark the chromosomes. Kymographs of anaphase progression of the E–P perimeter section depicted in e. Note that actin and myosin II have distinct patterns of localization at the cortex during anaphase (also, see white arrowheads in e). g, Fly epithelial cell at mid anaphase immunostained for tubulin, DNA, p-myosin II and F-actin (phalloidin), representative of three cells. h, Fly epithelial cell at mid anaphase immunostained for p-moesin and DNA, representative of 15 cells. Scale bars in b, e, g, h, 5 µm. A two-tailed unpaired t-test was used to calculate statistical significance.
Extended Data Figure 3 Actin clearance from the poles is independent of centrosomes and astral microtubules.
a, SOP cell imaged at metaphase and anaphase (left) (representative of three cells imaged precisely in this way), together with a kymograph of the cross-section (yellow box). Cnn indicates centrosomin. Lifeact–GFP was used to label F-actin. b, c, Fly epithelial cells were fixed and immunostained for centrosomin, tubulin and DNA. Green arrowheads indicate the presence of centrosomes in control cells. Representative images are shown (b), together with a quantification of relative centrosomin levels at the centrosome (c) for 25 cells from three control animals and for 26 cells from three AslmecD animals. A two-tailed unpaired t-test indicated that there was a significant difference in centrosomal centrosomin levels in the two cases. d, Scheme of SOP cells dividing in different orientations. A–P axis = 0° (left). Rosette plots indicate spindle axis angle measured at the onset of anaphase for 34 control cells from three animals and for 23 AslmecD cells from three animals. e, Time-lapse stills of SOP cells expressing GMA to label F-actin taken at early and mid anaphase (ana) in control (representative of 12 cells) and AslmecD (representative of 16 cells) mutant backgrounds (as shown in Fig. 1e, f), together with plot profiles (right) denoting the relative actin levels across the cell. Asterisks mark the chromosomes. f, Images show representative STLC-treated RPE-1 cells treated with or without 20 nM nocodazole (Noco) and/or RO3306 (15 cells were analysed for each condition), fixed and stained for p-ERM proteins, DNA and tubulin. g, Images in top panel show representative Mad2-depleted S2 cells treated with 25 µM colchicine and stained for F-actin (phalloidin, red in merged image), anillin (green in merged image) and Hoechst 33258 (blue in merged image), from a population of 13 cells. Similarly, the bottom panel shows images of S2 cells (representative of 13 cells) treated with colchicine, and 20 µM RO3306 to induce forced exit from mitosis, and stained for F-actin (phalloidin, red in merged image) and p-moesin (p-ERM antibody, green in merged image) and Hoechst 33258 (blue in merged image). h, i, Ratio of proximal/distal levels of cortical F-actin (h) and p-moesin (i) (refers to g). Mean is labelled in red. j, S2 cells, expressing either H2B–GFP/anillin–Cherry or Lifeact–GFP/H2B–Cherry, were imaged during mitotic exit. Representative stills and the corresponding kymographs are shown in j (equivalent to phenotype I in k). Dashed lines were used to generate the kymographs. Top panel, n = 68 cells, three experiments; bottom panel, n = 24 cells, one experiment. k, Phenotypic quantification of anillin–Cherry-expressing S2 cells treated with colchicine and forced to exit mitosis through either Mad2 depletion (as depicted in j, top panel) or through treatment with RO3306. Bar graphs depict mean and s.d. Phenotype I: DNA and cortex are polarized. Phenotype II: neither DNA nor cortex is polarized. Phenotype III: DNA is polarized but cortex is not. Mad2 RNAi, n = 68 cells, three experiments. RO3306, n = 121 cells, two experiments. In a, b, e, f, g and j, scale bars, 5 µm.
Extended Data Figure 4 DNA-induced clearance of cortical F-actin at anaphase.
a, b, Data show representative stills and corresponding quantitative data extracted from movies of 17 STLC-treated HeLa cells (from three independent experiments) expressing LifeAct–GFP and H2B–Cherry forced to flatten through Rap1* expression before and after treatment with the CDK inhibitor RO3306. a, Images show XZ cross-sections of a representative flattened HeLa cell before and after treatment with RO3306. Levels of cortical F-actin above the chromatin were normalized against the cytoplasmic fluorescence signal (ratios are shown in green on right). Graph on right shows normalized levels of cortical actin overlying the DNA before and after treatment with RO3306 (at 6 min after drug addition) for all 17 cells. b, XY cross-sections of representative cell shown in a (left). Levels of cortical actin below the chromatin (see dotted region) were normalized against the fluorescence signal in the most basal confocal section (ratios shown in green on right). Graph on right shows normalized levels of basal cortical actin lying beneath the DNA for all 17 cells. c, d, Scheme and data to test the correlation between cell elongation and anaphase chromosome movements at the anterior pole of fly SOP cells. c, Scheme depicts distances D1, D2 and D3. d, Graph shows D1, D2 and D3 plotted for anterior pole during anaphase for representative SOP cell A (shown in Fig. 2a, b, 1 of 12 analysed). e–i, Experiments to test how cortical actin is cleared from the anterior and posterior cortex of ten SOP cells during chromosome segregation. e, Scheme of cortical regions c1–c9 (as seen in Fig. 2f). f, g, Stills of the posterior and anterior poles of a representative SOP cell imaged in early anaphase. Arrowheads in f and g point to poor and strong actin clearance, respectively. h, i, Average plot of cortical actin measured over time for the posterior pole (Post) and anterior (Ant) pole (as seen in Fig. 2g, bottom left). The F-actin threshold level was set to 3.0 to facilitate the comparison between anterior and posterior poles. These data show that clearance of actin on the anterior pole occurs before clearance on the posterior pole in SOP cells. Scale bars in f and g, 5 µm. Box-and-whisker plots show median together with 10–90 percentiles. A two-tailed unpaired t-test was used to calculate statistical significance.
Extended Data Figure 5 Depletion of PP1-87B or Sds22 impairs cell elongation in SOP cells.
a–e, The correlation between cell elongation and the approach of chromatin to the cortex was analysed in control (12 cells, four animals), PP1-87B RNAi (16 cells, four animals) and Sds22 (10 cells, three animals) RNAi cells. a, Plot of the DNA-to-cortex distance during anaphase in three representative SOP cells in control, PP1-87B RNAi and Sds22 RNAi backgrounds. Anterior pole depicted. mD, mean distance during anaphase. b, Box plot of mean DNA-to-cortex distance in mid anaphase. c, Graphs show distance from cell centre to pole plotted before and after elongation onset in representative cells for each of the three conditions (in black), together with a fitted linear regression (in red). d, e, Box plot to show the slopes of linear regression analysis (as in c) before the elongation onset and after the elongation onset for control, PP1-87B RNAi and Sds22 RNAi backgrounds. f–h, Pre-furrow anaphase elongation for control (22 cells from five animals), PP1-87B RNAi (21 cells from four animals) and Sds22 RNAi (14 cells from three animals) SOP cells expressing Lifeact–GFP. f, Representative images of cells. g, Outlines of the boundary of cells shown in f at different times following the onset of anaphase. h, Graph shows pre-furrow anaphase cell elongation for cells in each background. These data show that PP1-87B or Sds22 depletion leads to a defect in anaphase elongation in SOP cells. i–k, Analysis shows anaphase elongation in control (12 cells from three animals) and AslmecD mutant (16 cells from three animals) cells. i, Images show F-actin in representative SOP cells expressing GMA in control and AslmecD mutant backgrounds. j, Outlines of boundary at different times following anaphase onset for representative cells shown in i. k, Plot of cell elongation in the backgrounds seen in i. These data show that anaphase cell elongation is not perturbed in the absence of centrosomes or astral microtubules. n, number of cells. Control, three animals; AslmecD, three animals. In f and i, scale bars, 5 µm. Box-and-whisker plots show median and 10–90 percentiles. Bar charts show mean and s.d. Significance was assessed using a two-tailed unpaired t-test. P > 0.05 was deemed not significant (n.s.).
Extended Data Figure 6 Depletion of Sds22 in human cells leads to impaired clearance of cortical actin.
a, Western blot showing depletion of Sds22 in HeLa cells through RNA interference. b, Control (representative of 18 cells) and Sds22 RNAi (representative of 11 cells) STLC-treated HeLa cells expressing Lifeact–GFP and H2B–mCherry before and after RO3306 treatment (which forces cells to exit mitosis). c, Scatter plot (median, red line) quantifying the minimal DNA-to-cortex distance after treatment with RO3306 in each case shown in b. d, Scatter plot (median, red line) showing cortical F-actin clearance below the DNA (as seen in Extended Data Fig. 4b); siRNA oligonucleotide no. 5 (seen in a) was used in experiments shown in b–d and Fig. 3c, d. In b, scale bar, 5 µm. Significance was assessed using a two-tailed unpaired t-test.
Extended Data Figure 7 Moesin is a target of PP1-87B–Sds22 and controls cortical relaxation at anaphase.
a–c, The effect of constitutively active moesin on anaphase polar relaxation. a, An SOP cell (1 of 13 cells) expressing constitutively active moesin (moesin(T559D)) imaged in metaphase and anaphase (top panel) and a kymograph of a cross-section over time (yellow box). Only the anterior pole is indicated. b, Plot of the DNA-to-cortex distance over time for a representative moesin–GFP-expressing cell (cell X) and a moesin(T559D)–GFP-expressing cell (cell Y in a). mD, mean distance during anaphase. c, Box-and-whisker plot (median and 10–90 percentiles) of mD in moesin–GFP and moesin(T559D)–GFP-expressing cells. This shows that the DNA comes into close apposition to the cortex in cells expressing constitutively active moesin as the result of failure to trigger efficient polar relaxation, as it does in cells depleted for PP1-87B or Sds22. d–f, The effect of moesin(T559D) expression on pre-furrow elongation in the same experiment as a–c. d, Images show representative SOP cells expressing moesin–GFP or moesin(T559D)–GFP transgenes at metaphase and anaphase (out of 13 cells in each case). e, Outlines of the boundary of cells shown in d at different times during anaphase. f, Bar graph of cell elongation in these two backgrounds showing mean and s.d. As observed in PP1-87B- or Sds22-depleted cells, moesin(T559D)–GFP-expressing cells show aberrant cell elongation at anaphase. g, Immunoprecipitation (IP) assays showing moesin dephosphorylation by PP1-87B–Sds22. Calyculin A (CalA) is an inhibitor of PP1 activity (left panel). Upon addition of CalA, PP1-87B activity is suppressed, leading to higher levels of phosphorylated moesin than in the absence of the compound (see p-Flag–moesin immunoblotting). PP1-87B acts with Sds22 to dephosphorylate active moesin (right panel). Upon Sds22 depletion, PP1-87B is less efficient in inactivating moesin. Red arrows indicate PP1-87B–GFP band. Results in g were replicated three times. h, Scheme of PP1–Sds22-dependent inactivation of moesin. In a and d, scale bar, 5 µm. Significance was assessed using a two-tailed unpaired t-test. PM, plasma membrane.
Extended Data Figure 8 Depletion of PP1-87B and Sds22 or expression of moesin(T559D)–GFP lead to severe shape defects in telophase cells.
a–d, Data show the impact of PP1-87B or Sds22 silencing and moesin(T559D) overexpression on telophase cell shape. a, Stills show representative telophase cells in control (1 from 32), PP1-87B RNAi (1 from 31) and Sds22 RNAi (1 from 27) backgrounds. Circularity of cells, C, is indicated. F-actin is labelled by Lifeact–GFP. b, Images show representative stills of telophase moesin–GFP (1 from 19) or moesin(T559D)–GFP (1 from 15) cells. Circularity of cells, C,is indicated. PIIa and PIIb are the cells that result from an asymmetric SOP division (in a and b; in each image, PIIa is the daughter cell on the left and PIIb is the daughter cell on the right.) c, Box plot of circularity of nascent cells at telophase in control, PP1-87B RNAi and Sds22 RNAi tissues. d, Box plot of circularity of nascent cells at telophase in moesin–GFP and moesin(T559D)–GFP-expressing tissues. In a and b, scale bar, 5 µm. Box-and-whisker plots show median and 10–90 percentiles. Significance was assessed using a two-tailed unpaired t-test.
Extended Data Figure 9 Polarization of cortical myosin II in anaphase does not depend on PP1 phosphatase.
a–c, Data show the impact of PP1-87B silencing (16 cells from four animals) on myosin repolarization during anaphase onset relative to a control (12 cells from four animals). a, b, Stills of representative control (a) and PP1-87B-depleted (b) SOP cells in anaphase labelled for myosin II (Sqh–mCherry) and F-actin (GMA) (top), together with the corresponding kymographs showing the equator–pole (E–P) perimeter section during anaphase progression. c, Schematic and graph show length of actin and myosin-II domains along the E–P perimeter in control and PP1-87B RNAi SOP cells. Mean and s.d. are shown. Significance was assessed using a two-tailed unpaired t-test. P > 0.05 was deemed not significant (n.s.). Scale bar, 5 µm. n, number of cells. These data show that although PP1-87B controls the polarization of cortical actin in anaphase, it does not affect the timely accumulation of myosin II at the equator.
Extended Data Figure 10 Local accumulation of Sds22 triggers polar blebbing in anaphase.
a, Confocal cross-sections of a representative anaphase epithelial cell showing co-localization of Sds22 and the kinetochore protein Spc25 (1 of 10 cells). Insets are of regions shown by arrowheads. b–e, Data to assess the impact of KNL1 silencing on Sds22–GFP localization and polar relaxation. b, Representative epithelial cell expressing Sds22–GFP imaged during anaphase, together with a corresponding kymograph of anaphase progression in c. Black arrowheads point to polar blebbing. Inverted lookup table in b, c. Darker tone indicates a stronger GFP signal. In a and b, scale bar, 5 µm. d, Line scans across kinetochore regions denoted by the green arrowheads in representative images shown in Fig. 4a. e, Levels of Sds22–GFP at kinetochores in control (17 cells from four animals) and KNL1 RNAi cells (nine cells from three animals) normalized against the cytoplasmic GFP signal. Graphs show mean and s.d. Significance was assessed using a two-tailed unpaired t-test. f, Graphic representation of the blue-light-induced cryptochrome-based protein–protein interaction system underlying the data shown in Fig. 4d. This scheme shows how the CRY2-tagged Sds22 subunit interacts with membrane-tethered CIBN upon blue-light irradiation, promoting fast translocation of the phosphatase to the plasma membrane and inactivation of cortical moesin and, consequently, abrogation of F-actin linkage to the membrane.
Supplementary information
Live imaging of an SOP cell labeled for F-actin (grey) and DNA (red) from anaphase onset (0 sec) until mid-anaphase.
Live imaging of an SOP cell labeled for F-actin (grey) and DNA (red) from anaphase onset (0 sec) until mid-anaphase. Polar relaxation occurs when the DNA masses come into close apposition with the cortex. Scale bar = 5 μm. (This video refers to Fig.2a-b) (AVI 166 kb)
Live imaging of a PP1-87B-depleted SOP cell labeled for F-actin (grey) and DNA (red) from anaphase onset (0 sec) until mid-anaphase.
Live imaging of a PP1-87B-depleted SOP cell labeled for F-actin (grey) and DNA (red) from anaphase onset (0 sec) until mid-anaphase. Scale bar = 5 μm. (This video refers to Fig.3a-b) (AVI 197 kb)
Live imaging of an Sds22-depleted SOP cell labeled for F-actin (grey) and DNA (red) from anaphase onset (0 sec) until mid-anaphase.
Live imaging of an Sds22-depleted SOP cell labeled for F-actin (grey) and DNA (red) from anaphase onset (0 sec) until mid-anaphase. Scale bar = 5 μm. (This video refers to Fig.3a-b) (AVI 148 kb)
Live imaging of an SOP cell labeled for F-actin (green) and DNA (red) from anaphase onset (0 sec) until late telophase.
Live imaging of an SOP cell labeled for F-actin (green) and DNA (red) from anaphase onset (0 sec) until late telophase. Scale bar = 5 μm. (This video refers to Extended Data Fig.8a)Scale bar = 5 μm. (This video refers to Extended Data Fig.8a) (AVI 451 kb)
Live imaging of a PP1-87B-depleted SOP cell labeled for F-actin (green) and DNA (red) from anaphase onset (0 sec) until late telophase.
Live imaging of a PP1-87B-depleted SOP cell labeled for F-actin (green) and DNA (red) from anaphase onset (0 sec) until late telophase. Scale bar = 5 μm. (This video refers to Extended Data Fig.8a) (AVI 1379 kb)
Live imaging of an Sds22-depleted SOP cell labeled for F-actin (green) and DNA (red) from anaphase onset (0 sec) until late telophase.
Live imaging of an Sds22-depleted SOP cell labeled for F-actin (green) and DNA (red) from anaphase onset (0 sec) until late telophase. Scale bar = 5 μm. (This video refers to Extended Data Fig.8a) (AVI 1578 kb)
Live imaging of an SOP cell expressing MoesinT559D-GFP (green) from anaphase onset (0 sec) until late telophase.
Live imaging of an SOP cell expressing MoesinT559D-GFP (green) from anaphase onset (0 sec) until late telophase. Scale bar = 5 μm. (This video refers to Extended Data Fig.8b). (AVI 209 kb)
Rights and permissions
About this article
Cite this article
Rodrigues, N., Lekomtsev, S., Jananji, S. et al. Kinetochore-localized PP1–Sds22 couples chromosome segregation to polar relaxation. Nature 524, 489–492 (2015). https://doi.org/10.1038/nature14496
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14496
This article is cited by
-
Spindle positioning and its impact on vertebrate tissue architecture and cell fate
Nature Reviews Molecular Cell Biology (2021)
-
Optogenetic relaxation of actomyosin contractility uncovers mechanistic roles of cortical tension during cytokinesis
Nature Communications (2021)
-
Inhibition of polar actin assembly by astral microtubules is required for cytokinesis
Nature Communications (2021)
-
Protein phosphatase 1 regulates atypical mitotic and meiotic division in Plasmodium sexual stages
Communications Biology (2021)
-
Co-option of Plasmodium falciparum PP1 for egress from host erythrocytes
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.