Abstract
Many long non-coding RNAs (lncRNAs) affect gene expression1, but the mechanisms by which they act are still largely unknown2. One of the best-studied lncRNAs is Xist, which is required for transcriptional silencing of one X chromosome during development in female mammals3,4. Despite extensive efforts to define the mechanism of Xist-mediated transcriptional silencing, we still do not know any proteins required for this role3. The main challenge is that there are currently no methods to comprehensively define the proteins that directly interact with a lncRNA in the cell5. Here we develop a method to purify a lncRNA from cells and identify proteins interacting with it directly using quantitative mass spectrometry. We identify ten proteins that specifically associate with Xist, three of these proteins—SHARP, SAF-A and LBR—are required for Xist-mediated transcriptional silencing. We show that SHARP, which interacts with the SMRT co-repressor6 that activates HDAC37, is not only essential for silencing, but is also required for the exclusion of RNA polymerase II (Pol II) from the inactive X. Both SMRT and HDAC3 are also required for silencing and Pol II exclusion. In addition to silencing transcription, SHARP and HDAC3 are required for Xist-mediated recruitment of the polycomb repressive complex 2 (PRC2) across the X chromosome. Our results suggest that Xist silences transcription by directly interacting with SHARP, recruiting SMRT, activating HDAC3, and deacetylating histones to exclude Pol II across the X chromosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Guttman, M. et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477, 295–300 (2011)
Rinn, J. L. & Chang, H. Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 81, 145–166 (2012)
Wutz, A. Gene silencing in X-chromosome inactivation: advances in understanding facultative heterochromatin formation. Nature Rev. Genet. 12, 542–553 (2011)
Lee, J. T. Lessons from X-chromosome inactivation: long ncRNA as guides and tethers to the epigenome. Genes Dev. 23, 1831–1842 (2009)
McHugh, C. A., Russell, P. & Guttman, M. Methods for comprehensive experimental identification of RNA-protein interactions. Genome Biol. 15, 203 (2014)
Shi, Y. et al. Sharp, an inducible cofactor that integrates nuclear receptor repression and activation. Genes Dev. 15, 1140–1151 (2001)
You, S. H. et al. Nuclear receptor co-repressors are required for the histone-deacetylase activity of HDAC3 in vivo. Nature Struct. Mol. Biol. 20, 182–187 (2013)
Zhao, J., Sun, B. K., Erwin, J. A., Song, J. J. & Lee, J. T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, 750–756 (2008)
Plath, K. et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 300, 131–135 (2003)
Hasegawa, Y., Brockdorff, N., Kawano, S., Tsutui, K. & Nakagawa, S. The matrix protein hnRNP U is required for chromosomal localization of Xist RNA. Dev. Cell 19, 469–476 (2010)
Schoeftner, S. et al. Recruitment of PRC1 function at the initiation of X inactivation independent of PRC2 and silencing. EMBO J. 25, 3110–3122 (2006)
Kalantry, S. & Magnuson, T. The Polycomb group protein EED is dispensable for the initiation of random X-chromosome inactivation. PLoS Genet. 2, e66 (2006)
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013)
Darnell, R. B. HITS-CLIP: panoramic views of protein–RNA regulation in living cells. Wiley Interdiscipl. Rev. RNA 1, 266–286 (2010)
Ong, S. E. & Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nature Protoc. 1, 2650–2660 (2007)
Ariyoshi, M. & Schwabe, J. W. A conserved structural motif reveals the essential transcriptional repression function of Spen proteins and their role in developmental signaling. Genes Dev. 17, 1909–1920 (2003)
Raffel, G. D. et al. Ott1(Rbm15) has pleiotropic roles in hematopoietic development. Proc. Natl Acad. Sci. USA 104, 6001–6006 (2007)
Haas, S., Steplewski, A., Siracusa, L. D., Amini, S. & Khalili, K. Identification of a sequence-specific single-stranded DNA binding protein that suppresses transcription of the mouse myelin basic protein gene. J. Biol. Chem. 270, 12503–12510 (1995)
Olins, A. L., Rhodes, G., Welch, D. B., Zwerger, M. & Olins, D. E. Lamin B receptor: multi-tasking at the nuclear envelope. Nucleus 1, 53–70 (2010)
Brown, C. J. & Baldry, S. E. Evidence that heteronuclear proteins interact with XIST RNA in vitro. Somat. Cell Mol. Genet. 22, 403–417 (1996)
Sarma, K. et al. ATRX directs binding of PRC2 to Xist RNA and Polycomb targets. Cell 159, 869–883 (2014)
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011)
Chaumeil, J., Le Baccon, P., Wutz, A. & Heard, E. A novel role for Xist RNA in the formation of a repressive nuclear compartment into which genes are recruited when silenced. Genes Dev. 20, 2223–2237 (2006)
Arieti, F. et al. The crystal structure of the Split End protein SHARP adds a new layer of complexity to proteins containing RNA recognition motifs. Nucleic Acids Res. 42, 6742–6752 (2014)
Li, J., Lin, Q., Wang, W., Wade, P. & Wong, J. Specific targeting and constitutive association of histone deacetylase complexes during transcriptional repression. Genes Dev. 16, 687–692 (2002)
Keohane, A. M., O’Neill, L. P., Belyaev, N. D., Lavender, J. S. & Turner, B. M. X-inactivation and histone H4 acetylation in embryonic stem cells. Dev. Biol. 180, 618–630 (1996)
Riising, E. M. et al. Gene silencing triggers polycomb repressive complex 2 recruitment to CpG islands genome wide. Mol. Cell 55, 347–360 (2014)
van der Vlag, J. & Otte, A. P. Transcriptional repression mediated by the human polycomb-group protein EED involves histone deacetylation. Nature Genet. 23, 474–478 (1999)
Davidovich, C., Zheng, L., Goodrich, K. J. & Cech, T. R. Promiscuous RNA binding by Polycomb repressive complex 2. Nature Struct. Mol. Biol. 20, 1250–1257 (2013)
Fackelmayer, F. O., Dahm, K., Renz, A., Ramsperger, U. & Richter, A. Nucleic-acid-binding properties of hnRNP-U/SAF-A, a nuclear-matrix protein which binds DNA and RNA in vivo and in vitro. Eur. J. Biochem. 221, 749–757 (1994)
Wang, Z. et al. Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell 138, 1019–1031 (2009)
Kuo, M. H. & Allis, C. D. Roles of histone acetyltransferases and deacetylases in gene regulation. Bioessays 20, 615–626 (1998)
Wutz, A., Rasmussen, T. P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nature Genet. 30, 167–174 (2002)
Engreitz, J. M. et al. RNA-RNA interactions enable specific targeting of noncoding RNAs to nascent pre-mRNAs and chromatin sites. Cell 159, 188–199 (2014)
Kalli, A. & Hess, S. Effect of mass spectrometric parameters on peptide and protein identification rates for shotgun proteomic experiments on an LTQ-orbitrap mass analyzer. Proteomics 12, 21–31 (2012)
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nature Biotechnol. 26, 1367–1372 (2008)
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011)
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for mass spectrometry-based proteomics. Methods Mol. Biol. 604, 55–71 (2010)
Geiger, T. et al. Use of stable isotope labeling by amino acids in cell culture as a spike-in standard in quantitative proteomics. Nature Protoc. 6, 147–157 (2011)
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014)
Jeon, Y. & Lee, J. T. YY1 tethers Xist RNA to the inactive X nucleation center. Cell 146, 119–133 (2011)
Agrelo, R. et al. SATB1 defines the developmental context for gene silencing by Xist in lymphoma and embryonic cells. Dev. Cell 16, 507–516 (2009)
Royce-Tolland, M. E. et al. The A-repeat links ASF/SF2-dependent Xist RNA processing with random choice during X inactivation. Nature Struct. Mol. Biol. 17, 948–954 (2010)
Theodosiou, Z. et al. Automated analysis of FISH and immunohistochemistry images: a review. Cytometry A 71, 439–450 (2007)
Fumagalli, M. et al. Telomeric DNA damage is irreparable and causes persistent DNA-damage-response activation. Nature Cell Biol. 14, 355–365 (2012)
Acknowledgements
We thank J. Engreitz for extensive discussions, help in adapting the RAP method, and critical comments on the manuscript; A. Gnirke, S. Carr, J. Jaffe and M. Schenone for initial discussions about the RAP-MS method; A. Collazo, E. Lubek, and L. Cai for microscopy help; A. Wutz for providing transgenic cell lines; R. Eggleston-Rangel for assistance with mass spectrometry; S. Grossman, I. Amit, M. Garber and J. Rinn for comments on the manuscript and helpful suggestions; and S. Knemeyer for illustrations. C.A.M. is supported by a post-doctoral fellowship from Caltech. C.-K.C. is supported by an NIH NRSA training grant (T32GM07616). Imaging was performed in the Biological Imaging Facility, with the support of the Caltech Beckman Institute and the Arnold and Mabel Beckman Foundation. This work was funded by the Gordon and Betty Moore Foundation (GBMF775), the Beckman Institute, and NIH (1S10RR029591-01A1 to S.H.), an NIH Director’s Early Independence Award (DP5OD012190), the Rose Hills Foundation, Edward Mallinckrodt Foundation, Sontag Foundation, Searle Scholars Program, and funds from the California Institute of Technology.
Author information
Authors and Affiliations
Contributions
C.A.M. developed the RAP-MS method, designed, performed, and analysed RAP-MS experiments and data, C.-K.C. designed, performed, and analysed Xist functional experiments, A.C. designed, performed, and oversaw experiments, C.F.S. helped develop RAP-MS and performed experiments, C.T., P.M., A.P.-J., A.M., A.A.S., J.S. performed experiments, M.J.S., M.B., C.B. analysed data, E.S.L. helped develop initial ideas for adapting RAP for protein detection, S.H. oversaw mass spectrometry development and data analysis, K.P. helped design Xist RAP-MS and functional experiments and analysed data, M.G. conceived, designed and oversaw the entire project and integrated the data, C.A.M., C.-K.C. and M.G. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 RAP-MS recovers and enriches the majority of Xist RNA from mouse ES cells, and these cells can be efficiently labelled with SILAC.
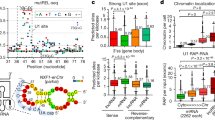
a, RT-qPCR measuring the percentage of the total cellular Xist or 18S recovered after RAP-MS of Xist. Values are computed as the amount of each RNA in the elution divided by the amount of RNA in the starting (‘input’) lysate material. Error bars represent the standard error of the mean from 5 biological replicates. b, Enrichment of Xist after RAP-MS captures from pSM33 cells as measured by qPCR. Bars indicate RNA levels of Xist, 18S, and Oct4 after purification of Xist, normalized to RNA in input sample. Each bar represents the RNA levels of Xist, 18S, and Oct4 after purification of Xist, normalized to RNA in input sample, from 3 biological replicates. c, SILAC labelling efficiency of a representative culture of pSM33 mouse ES cells after 10 days of growth (3 cell passages) in SILAC medium. Peptides were analysed by mass spectrometry, and values indicate the fraction of identified peptides with heavy-label incorporation with different levels of peptide labelling (shown in bins).
Extended Data Figure 2 RAP-MS identifies proteins that are known to directly interact with specific ncRNAs, and separates specific RNA interacting proteins from background proteins.
a, SILAC ratios of top proteins enriched in the RAP-MS U1 snRNA, 18S rRNA, and 45S pre-rRNA experiments. b, SILAC ratio plot of replicate captures of U1 snRNA versus 18S rRNA from one of two biologically independent label-swap experiments. Proteins associated with U1 are consistently found in U1 samples, both light and heavy labelled (top right quadrant), and proteins specifically associated with 18S are consistently identified in 18S, both light and heavy (lower left quadrant). Background contaminant proteins have low enrichments (centre of panel) or are consistently found in the light channel and do not replicate between experiments (that is, keratin, streptavidin). c, SILAC ratio plot of replicate captures of U1 snRNA versus 45S pre-rRNA from one label-swap experiment. Proteins that are known to associate with 45S pre-rRNA are consistently identified in 45S captures.
Extended Data Figure 3 Immunoprecipitation of the identified Xist-interacting proteins confirms Xist RNA interaction.
RNA immunoprecipitation experiments were performed for seven Xist-interacting proteins (black bars), two control RNA binding proteins that were not identified by RAP-MS and IgG (grey bars) in UV-crosslinked cell lysate after 6 h of Xist induction by doxycycline addition (Methods). The RNA associated with each protein was measured and enrichment levels were computed relative to the level of the RNA in total cellular input and normalized to the total efficiency of capture in each sample to allow for direct comparison across all immunoprecipitation experiments (Methods). a, Enrichment of the Xist lncRNA after immunoprecipitation from a sample of pSM33 male cells. b, Immunoprecipitation of SHARP was performed from a sample of UV-crosslinked females ES cells that were treated with retinoic acid for 24 h. The levels of recovered Xist lncRNA (black bars), Neat1 lncRNA (white bars), and 45S pre-ribosomal RNA (grey bars) were measured by RT-qPCR. Enrichment of each RNA after capture with anti-SHARP antibody was calculated relative to the level of RNA captured with IgG control antibody. c, The enrichment of various lncRNAs after immunoprecipitation in pSM33 male cells—including Neat1, Malat1, Firre, and Tug1—are shown. d, The enrichment of various mRNA controls after immunoprecipitation in pSM33 male cells—including Oct4, Nanog, Stat3, and Suz12—are shown.
Extended Data Figure 4 Previously identified proteins associated with XCI are not required for Xist-mediated transcriptional silencing.
a, To confirm the specificity of our assay, we tested the function of several proteins that were previously identified to associate with Xist, but not to silence transcription, for their role in transcriptional silencing in our inducible male ES cells before Xist induction (−Dox; left) or after Xist induction for 16 h (+Dox; middle and right). Representative images are shown after knockdown of each protein. DAPI (blue), Xist (red), and Gpc4 (green). b, Quantification of the copy number of Gpc4 before and after Xist induction upon treatment with different siRNAs. Error bars represent the standard error of the mean across 50 individual cells from one experiment. ****P value < 0.001 between +Dox and –Dox cells based on an unpaired two-sample t-test. Scale bars on the images represent 5 μm. Importantly, while these proteins do not have a role in the initiation of transcriptional silencing, we do not mean to imply that they do not have other roles in XCI.
Extended Data Figure 5 SHARP, LBR, SAF-A, SMRT, and HDAC3 are required for Xist-mediated transcriptional silencing.
a, Representative images showing staining of DAPI (blue), Xist (red), and Gpc4 (green) for different siRNA knockdown in male ES cells before Xist induction (−Dox; left) or after Xist induction for 16 h (+Dox; middle and right). b, Quantification of the copy number of Gpc4 in –Dox and +Dox cells after knockdown with siRNAs targeting different mRNAs. Error bars represent the standard error of the mean across 50 individual cells from one experiment. NS, not significantly different between +Dox and –Dox cells; ****P value < 0.001 between +Dox and –Dox cells based on an unpaired two-sample t-test. Scale bars on the images represent 5 μm. c, Knockdown of SHARP, LBR, or SAF-A abrogates Xist-mediated gene silencing without causing pluripotency defects. Representative images showing staining of Nanog (cyan), Xist (red), and Gpc4 (green) upon knockdown of SHARP, LBR or SAF-A after 16 h of Xist induction with doxycycline. Scale bars on the images represent 5 μm.
Extended Data Figure 6 SHARP is required for silencing many genes across the X chromosome.
a, A diagram showing the locations of Xist (red), X-linked silenced genes (black), and X-linked escaped genes (green) along the X chromosome. b, Representative images showing staining of DAPI (blue), Xist (red), X-linked silenced genes (green), and X-linked escaped genes (yellow) upon knockdown of SHARP or control male ES cells before Xist induction (−Dox) or after Xist induction for 16 h (+Dox). Knock of SHARP abolishes the silencing of Atrx, Gpc4, Rbmx, Smc1a and Mecp2, which are normally silenced upon Xist expression, but has no effect on Mid1 and Pir, which normally escape Xist-mediated silencing. The bar graphs show the quantification of the copy number of the mRNA for each gene for –Dox and +Dox cells upon transfection with SHARP siRNA or control siRNA; error bars represent the standard error of the mean across 50 individual cells from one experiment. NS, not significantly different, ****P value < 0.001, and **P value < 0.01 between +Dox and –Dox cells based on an unpaired two-sample t-test. Scale bars on the images represent 5 μm.
Extended Data Figure 7 Multiple independent siRNAs targeting SHARP, LBR, SAF-A, HDAC3, or SMRT demonstrate the same silencing defect.
a, Representative images showing staining of DAPI (blue), Xist (red), and Gpc4 (green) after knockdown of proteins using independent, non-overlapping, siRNA pools, or individual siRNA deconvoluted from the pool before Xist induction (−Dox; left) or after Xist induction for 16 h (+Dox; middle and right). Cells were either transfected with the siRNA pool from Dharmacon (siRNA-D), Qiagen (siRNA-Q) or Ambion/Life Technologies (siRNA-A), or each individual siRNA deconvoluted from the pool from Dharmacon (siRNA-D1, 2, 3, 4) or Qiagen (siRNA-Q1, 2, 3, 4). b, Quantification of the copy number of Gpc4 in –Dox and +Dox cells after knockdown with siRNAs targeting different mRNAs. Error bars represent the standard error of the mean across 50 individual cells from one experiment. NS, not significantly different between +Dox and –Dox cells based on an unpaired two-sample t-test. Scale bars on the images represent 5 μm. We excluded all siRNAs that did not reduce the targeted mRNA level by >70% (Methods). The sequences of deconvoluted siRNAs are shown in Supplementary Table 2.
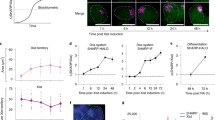
Extended Data Figure 8 SHARP, LBR, SAF-A, SMRT, and HDAC3 are required for transcriptional silencing in differentiating female ES cells.
a, Representative images showing staining of DAPI (blue), Xist (red), and Gpc4 (green) upon knockdown of specific proteins using different siRNAs in female ES cells before differentiation (−RA; left) or after differentiation for 24 h (+RA; middle and right). RA, retinoic acid. b, Quantification of the copy number of Gpc4 for –RA and +RA cells upon transfection with different siRNAs. Error bars represent the standard error of the mean across 50 individual cells from one experiment. NS, not significantly different between +RA and –RA cells; ****P value < 0.001, **P value < 0.01, and *P value < 0.05 between +RA and –RA cells based on an unpaired two-sample t-test. Scale bars on the images represent 5 μm.
Extended Data Figure 9 SHARP is required for exclusion of RNA polymerase II from the Xist-coated territory in differentiating female ES cells.
Images of individual cells that are labelled with Xist (red), RNA Polymerase II (green), and DAPI (blue) across different siRNA conditions (rows) in female ES cells after 24 h of retinoic acid treatment. The dashed white region represents the outlined Xist-coated territory.
Extended Data Figure 10 SHARP is required for PRC2 recruitment across the Xist-coated territory in differentiating female ES cells.
Images of individual cells that are labelled with Xist (red), Ezh2 (green) and DAPI (blue) across different siRNA conditions (rows) in female ES cells after 24 h of differentiation. The dashed white region represents the outlined Xist-coated territory.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-5 and Supplementary Tables 1-2. (PDF 252 kb)
Rights and permissions
About this article
Cite this article
McHugh, C., Chen, CK., Chow, A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015). https://doi.org/10.1038/nature14443
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14443
This article is cited by
-
Regulatory function and mechanism research for m6A modification WTAP via SUCLG2-AS1- miR-17-5p-JAK1 axis in AML
BMC Cancer (2024)
-
An estrogen-regulated long non-coding RNA NCALD promotes luminal breast cancer proliferation by activating GRHL2
Cancer Cell International (2024)
-
Prediction of on-target and off-target activity of CRISPR–Cas13d guide RNAs using deep learning
Nature Biotechnology (2024)
-
Non-coding RNAs in disease: from mechanisms to therapeutics
Nature Reviews Genetics (2024)
-
Pan-cancer analysis identifies SPEN mutation as a predictive biomarker with the efficacy of immunotherapy
BMC Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.