Abstract
Although thousands of long non-coding RNAs (lncRNAs) are encoded in mammalian genomes, their mechanisms of action are poorly understood, in part because they are often expressed at lower levels than their proposed targets. One such lncRNA is Xist, which mediates chromosome-wide gene silencing on one of the two X chromosomes (X) to achieve gene expression balance between males and females. How a limited number of Xist molecules can mediate robust silencing of a much larger number of target genes while maintaining specificity exclusively to genes on the X within each cell is not well understood. Here, we show that Xist drives non-stoichiometric recruitment of the essential silencing protein SHARP (also known as SPEN) to amplify its abundance across the inactive X, including at regions not directly occupied by Xist. This amplification is achieved through concentration-dependent homotypic assemblies of SHARP on the X and is required for chromosome-wide silencing. Expression of Xist at higher levels leads to increased localization at autosomal regions, demonstrating that low levels of Xist are critical for ensuring its specificity to the X. We show that Xist (through SHARP) acts to suppress production of its own RNA which may act to constrain overall RNA levels and restrict its ability to spread beyond the X. Together, our results demonstrate a spatial amplification mechanism that allows Xist to achieve two essential but countervailing regulatory objectives: chromosome-wide gene silencing and specificity to the X. This suggests a more general mechanism by which other low-abundance lncRNAs could balance specificity to, and robust control of, their regulatory targets.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Previously published RAP-DNA sequencing data used in Extended Data Fig. 5b,f is available at GEO accession GSE46918. RNA-DNA SPRITE data used in Fig. 5f,g and Extended Data Fig. 5 is available at GEO accession GSE151515. CLAP sequencing data generated in this study and used in Fig. 3c,d and RAP-DNA sequencing data generated in this study and used in Fig. 5d–f is available at GEO accession GSE192574. Additional source data files are available for Figs. 1c,d,e,g, 2e,g, 3b, 4c,e,g and 5c,i,j. Source data are provided with this paper.
References
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Engreitz, J. M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Lee, J. T. Lessons from X-chromosome inactivation: long ncRNA as guides and tethers to the epigenome. Genes Dev. 23, 1831–1842 (2009).
Nagano, T. et al. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322, 1717–1720 (2008).
Dunagin, M., Cabili, M. N., Rinn, J. & Raj, A. Visualization of lncRNA by single-molecule fluorescence in situ hybridization. Methods Mol. Biol. 1262, 3–19 (2015).
Wang, K. C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 341, 789–792 (2013).
Thomson, D. W. & Dinger, M. E. Endogenous microRNA sponges: evidence and controversy. Nat. Rev. Genet. 17, 272–283 (2016).
Kopp, F. & Mendell, J. T. Functional classification and experimental dissection of long noncoding RNAs. Cell 172, 393 (2018).
Penny, G. D., Kay, G. F., Sheardown, S. A., Rastan, S. & Brockdorff, N. Requirement for Xist in X chromosome inactivation. Nature 379, 131–137 (1996).
Marahrens, Y., Loring, J. & Jaenisch, R. Role of the Xist gene in X chromosome choosing. Cell 92, 657–664 (1998).
Dyer, K. A., Canfield, T. K. & Gartler, S. M. Molecular cytological differentiation of active from inactive X domains in interphase: implications for X chromosome inactivation. Cytogenet. Genome Res. 50, 116–120 (1989).
Chaumeil, J., Le Baccon, P., Wutz, A. & Heard, E. A novel role for Xist RNA in the formation of a repressive nuclear compartment into which genes are recruited when silenced. Genes Dev. 20, 2223–2237 (2006).
Clemson, C. M., McNeil, J. A., Willard, H. F. & Lawrence, J. B. XIST RNA paints the inactive X chromosome at interphase: evidence for a novel RNA involved in nuclear/chromosome structure. J. Cell Biol. 132, 259–275 (1996).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Simon, M. D. et al. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 504, 465–469 (2013).
Sunwoo, H., Wu, J. Y. & Lee, J. T. The Xist RNA−PRC2 complex at 20-nm resolution reveals a low Xist stoichiometry and suggests a hit-and-run mechanism in mouse cells. Proc. Natl Acad. Sci. USA 112, E4216–E4225 (2015).
Markaki, Y. et al. Xist nucleates local protein gradients to propagate silencing across the X chromosome. Cell 184, 6174–6192 (2021).
Rodermund, L. et al. Time-resolved structured illumination microscopy reveals key principles of Xist RNA spreading. Science 372, eabe7500 (2021).
Marahrens, Y., Panning, B., Dausman, J., Strauss, W. & Jaenisch, R. Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes Dev. 11, 156–166 (1997).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Wutz, A., Rasmussen, T. P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat. Genet. 30, 167–174 (2002).
Dossin, F. et al. SPEN integrates transcriptional and epigenetic control of X-inactivation. Nature 578, 455–460 (2020).
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015).
Lu, Z. et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165, 1267–1279 (2016).
Minajigi, A. et al. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015).
Cirillo, D. et al. Quantitative predictions of protein interactions with long noncoding RNAs. Nat. Methods 14, 5–6 (2016).
Żylicz, J. J. et al. The implication of early chromatin changes in X chromosome inactivation. Cell 176, 182–197 (2019).
You, S. H. et al. Nuclear receptor co-repressors are required for the histone-deacetylase activity of HDAC3 in vivo. Nat. Struct. Mol. Biol. 20, 182–187 (2013).
Shi, Y. et al. Sharp, an inducible cofactor that integrates nuclear receptor repression and activation. Genes Dev. 15, 1140–1151 (2001).
Monfort, A. & Wutz, A. Identification of Spen as a crucial factor for Xist function through forward genetic screening in haploid embryonic stem cells. Cell Rep. 12, 554–561 (2015).
Moindrot, B. et al. A pooled shRNA screen identifies Rbm15, Spen, and Wtap as factors required for Xist RNA-mediated silencing. Cell Rep. 12, 562–572 (2015).
Nesterova, T. B. et al. Systematic allelic analysis defines the interplay of key pathways in X chromosome inactivation. Nat. Commun. 10, 3129 (2019).
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000).
Lee, J. T., Strauss, W. M., Dausman, J. A. & Jaenisch, R. A 450 kb transgene displays properties of the mammalian X-inactivation center. Cell 86, 83–94 (1996).
Hall, L. L. et al. An ectopic human XIST gene can induce chromosome inactivation in postdifferentiation human HT-1080 cells. Proc. Natl Acad. Sci. USA 99, 8677–8682 (2002).
Pacini, G. et al. Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution. Nat. Commun. 12, 3638 (2021).
Schulz, E. G. et al. The two active X chromosomes in female ESCs block exit from the pluripotent state by modulating the ESC signaling network. Cell Stem Cell 14, 203–216 (2014).
Barros de Andrade E Sousa, L. et al. Kinetics of Xist-induced gene silencing can be predicted from combinations of epigenetic and genomic features. Genome Res. 29, 1087–1099 (2019).
Arieti, F. et al. The crystal structure of the Split End protein SHARP adds a new layer of complexity to proteins containing RNA recognition motifs. Nucleic Acids Res. 42, 6742–6752 (2014).
Newberry, E. P., Latifi, T. & Towler, D. A. The RRM domain of MINT, a novel Msx2 binding protein, recognizes and regulates the rat osteocalcin promoter. Biochemistry 38, 10678–10690 (1999).
Hyman, A. A., Weber, C. A. & Jülicher, F. Liquid−liquid phase separation in biology. Annu. Rev. Cell Dev. Biol. 30, 39–58 (2014).
Lin, Y., Currie, S. L. & Rosen, M. K. Intrinsically disordered sequences enable modulation of protein phase separation through distributed tyrosine motifs. J. Biol. Chem. 292, 19110–19120 (2017).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855 (2018).
Bhat, P., Honson, D. & Guttman, M. Nuclear compartmentalization as a mechanism of quantitative control of gene expression. Nat. Rev. Mol. Cell Biol. 22, 653–670 (2021).
McSwiggen, D. T., Mir, M., Darzacq, X. & Tjian, R. Evaluating phase separation in live cells: diagnosis, caveats, and functional consequences. Genes Dev. 33, 1619–1634 (2019).
Banani, S. F. et al. Compositional control of phase-separated cellular bodies. Cell 166, 651–663 (2016).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Bracha, D. et al. Mapping local and global liquid phase behavior in living cells using photo-oligomerizable seeds. Cell 175, 1467–1480 (2018).
Lin, Y., Protter, D. S. W., Rosen, M. K. & Parker, R. Formation and maturation of phase-separated liquid droplets by RNA-binding proteins. Mol. Cell 60, 208–219 (2015).
Choi, J. et al. Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature 548, 219–223 (2017).
Yagi, M. et al. Derivation of ground-state female ES cells maintaining gamete-derived DNA methylation. Nature 548, 224–227 (2017).
Quinodoz, S. A. et al. Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell 174, 744–757 (2018).
Wollerton, M. C., Gooding, C., Wagner, E. J., Garcia-Blanco, M. A. & Smith, C. W. J. Autoregulation of polypyrimidine tract binding protein by alternative splicing leading to nonsense-mediated decay. Mol. Cell 13, 91–100 (2004).
Ottens, F. & Gehring, N. H. Physiological and pathophysiological role of nonsense-mediated mRNA decay. Eur. J. Physiol. 468, 1013–1028 (2016).
Müller-McNicoll, M., Rossbach, O., Hui, J. & Medenbach, J. Auto-regulatory feedback by RNA-binding proteins. J. Mol. Cell Biol. 11, 930–939 (2019).
Rossbach, O. et al. Auto- and cross-regulation of the hnRNP L proteins by alternative splicing. Mol. Cell. Biol. 29, 1442–1451 (2009).
Quinodoz, S. A. et al. RNA promotes the formation of spatial compartments in the nucleus. Cell 184, 5775–5790 (2021).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Ran, F. A. et al. Genome engineering using the CRISPR−Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Klock, H. E., Koesema, E. J., Knuth, M. W. & Lesley, S. A. Combining the polymerase incomplete primer extension method for cloning and mutagenesis with microscreening to accelerate structural genomics efforts. Proteins Struct. Funct. Genet. 71, 982–994 (2008).
Wu, S. C. Y. et al. piggyBac is a flexible and highly active transposon as compared to Sleeping Beauty, Tol2, and Mos1 in mammalian cells. Proc. Natl Acad. Sci. USA 103, 15008–15013 (2006).
Alberti, S. et al. A user’s guide for phase separation assays with purified proteins. J. Mol. Biol. 430, 4806–4820 (2018).
Wachsmuth, M. Molecular diffusion and binding analyzed with FRAP. Protoplasma 251, 373–382 (2014).
Koulouras, G. et al. EasyFRAP-web: a web-based tool for the analysis of fluorescence recovery after photobleaching data. Nucleic Acids Res. 46, W467–W472 (2018).
Banerjee, A. K. et al. SARS-CoV-2 disrupts splicing, translation, and protein trafficking to suppress host defenses. Cell 183, 1325–1339 (2020).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Acknowledgements
We thank M. Elowitz, S. Chong, I. Goronzy and D. Honson (Caltech) for their critical comments on the manuscript, A. Pandya-Jones, Y. Markaki and K. Plath (UCLA) for initial discussions and guidance on SHARP visualization, and F. Dossin, A. Loda and E. Heard (EMBL) for sharing their cell lines and cell culture protocols. We would like to thank A. Chow (Caltech) for helpful comments and support with the cell culture work done at the Guttman laboratory, the Biological Imaging Facility at Caltech for their help with microscopy, the Flow Cytometry Facility at Caltech for their help with cell sorting, and the Millard and Muriel Jacobs Genetics and Genomics Laboratory at Caltech for their help with sequencing. We also thank S. Hiley for contributions to writing and editing this manuscript, G. Riddihough and M. Bao (Life Science Editors) for editorial assistance, and I.M. Strazhnik (Caltech) for helping with illustrations. This work was supported by the National Institutes of Health (NIH) 4DN program (U01 DA040612 and U01 HL130007), NIH Directors’ Transformative Research Award (R01 DA053178), the New York Stem Cell Foundation and funds from the California Institute of Technology. M.G. is a NYSCF-Robertson Investigator. J.W.J. was supported by a Biology and Biological Engineering postdoctoral fellowship from Caltech. A.K.B. was funded by National Heart, Lung, and Blood Institute F30-HL136080 and the University of Southern California MD/PhD Program.
Author information
Authors and Affiliations
Contributions
J.W.J. conceived of this project with M.G. J.W.J. and M.S. performed experiments, analyzed and interpreted data, generated figures and wrote the paper. A.K.B. performed all CLAP sequencing experiments and provided comments and edits for the manuscript. J.T. created the SHARP rescue constructs with A.K.B. and assisted with cell culture. M.R.B. worked with A.K.B. on CLAP sequencing experiments, worked with J.W.J. on RAP sequencing experiments, analyzed sequencing data and provided comments and edits for the manuscript. M.G. oversaw all experiments and analysis, performed analyses and generated figures, and wrote the paper with J.W.J. and M.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Anke Sparmann, Beth Moorefield and Carolina Perdigoto were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
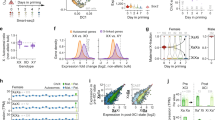
Extended Data Fig. 1 SHARP enrichment over the Xi increases in a non-stoichiometric manner relative to Xist.
a) Schematic of dox-inducible Xist expression system. The endogenous Xist promoter is replaced with a TetO element that can be activated upon the addition of doxycycline. b) Percent of cells expressing zero, one, or two alleles of the silenced Atrx gene as measured by RNA-FISH at various timepoints after Xist induction. c) Illustration of SHARP enrichment analysis over the Xi in TX-SHARP-HALO female mESCs. The Xi region was demarcated based on Xist RNA-FISH; SHARP was demarcated by either direct HALO labelling or immunofluorescence (anti-HALO). Fluorescent intensities of RNA-FISH probes, HALO tag, or anti-HALO immunofluorescence were then quantified within the defined Xi region and plotted. d) Representative images of Xist and SHARP localization in TX-SHARP-HALO female mESCs across 72 hours of Xist expression (n > 15 cells per timepoint from two independent experiments); Xist visualized by RNA-FISH (magenta); SHARP visualized by immunofluorescence labelling with anti-HALO antibody (green). Images shown as max. projections; scale bars 10 μm. e) Quantification of total fluorescence intensity of Xist (RNA-FISH) in multiple individual cells over 48 hours of Xist expression (Fig. 1b). f) Quantification of Xist and SHARP intensities in multiple individual cells over 72 hours of Xist expression (Extended Data Fig. 1d). Top panel: area of the territory coated by Xist RNA (µm2); middle panel: average fluorescent intensity of Xist (RNA-FISH) per unit within the Xist territory; bottom panel: average fluorescent intensity of SHARP (anti-Halo antibody) per unit within the Xist territory. g) Representative images of Xist and SHARP localization in TX-SHARP-HALO female mESCs after 48 and 72 hours of RA-induced differentiation (n > 10 cells for each timepoint); Xist visualized by RNA-FISH (magenta), SHARP visualized by direct Halo labelling (green). Images shown as max. projections; scale bars 10 μm. h) Quantification of Xist and SHARP in individual differentiated cells (Extended Data Fig. 1g). Left panel: area of the territory coated by Xist RNA (µm2); top right panel: average fluorescent intensity of Xist (RNA-FISH) per unit within the Xist territory; bottom right panel: average fluorescent intensity of SHARP (direct Halo labelling) per unit within the Xist territory.
Extended Data Fig. 2 SHARP forms multivalent, concentration-dependent assemblies in the nucleus.
a) Schematic of the domains included in the eGFP-tagged FL-SHARP and ΔIDR-SHARP, and the mCherry-tagged FUS-ΔIDR-SHARP rescue constructs used in Fig. 2 and Extended Data Fig. 2. b) FRAP recovery curve of eGFP-tagged FL-SHARP (red), positive control PTBP1 (forms assemblies; light blue), and negative control EED (does not form assemblies; dark blue). Points represent mean value; error bars represent standard deviation of at least five replicates. c) Schematic depicting physical characteristics of concentration-dependent assemblies, including foci formation, fission and fusion, and rapid diffusion of proteins within an assembly (inset). d) Images across nine time-points from a live-cell movie of eGFP-tagged FL-SHARP in transiently transfected HEK293T cells (Supplementary Video 1,2) showing non-diffused, focal organization of SHARP molecules. Top panel: 3D reconstructions of the fluorescent intensity signal; middle panel: 3D volume reconstructions color-coded based on the volume of the focus; bottom panel: zoom-in representing one region of the nucleus that changes volume; Fluorescent Intensity (FI) e) Comparison of diffused or non-diffused localization patterns of FL-SHARP at different dox concentrations. Left: images representing FL-SHARP expressed with either 0.1x dox (diffused) or 1x dox (non-diffused) in transiently transfected HEK293T cells; images shown as max projections; scale bars 10 μm. Right: histograms representing fluorescent intensities for two cells showing diffused and non-diffused localization patterns. The intensity at the 99th percentile of each distribution is shown with the dashed lines. f) Images representing nuclear localization pattern of eGFP-tagged proteins in transiently transfected HEK293T cells. Left: proteins that have not been reported to form assemblies (HALO and EED); Right: an eGFP tagged protein that has been reported to form assemblies (Ptbp1) and ΔSPOC-SHARP that also forms assemblies. Images shown as max projections; scale bars 10 μm.
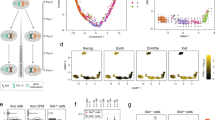
Extended Data Fig. 3 Formation of SHARP assemblies is required for SHARP enrichment on the Xi and is dispensable for Xist binding.
a) Generation of SHARP-KO cell line in TX mESCs. Top: schematic of CRISPR cut sites used to generate SHARP-KO mESCs and PCR primers used to screen for KO clones; bottom: agarose gel confirming homozygous deletion of SHARP in SHARP-KO clone H8 mESCs. b) Schematics of constructs used to generate rescue cell lines in TX SHARP-KO or TX SHARP-HALO-AID backgrounds. Grey arrow represents dox-inducible promoter; blue box represents HALO (or eGFP) tags used; light green boxes represent RNA Recognition Motifs (RRM); wavy green line represents the Intrinsically Disordered Regions (IDRs); dark green box represents the Spen Paralog and Ortholog C-terminal (SPOC) domain. Full-length SHARP (FL-SHARP), deletion of RRM domain (ΔRRM-SHARP), deletion of IDR domain (ΔIDR-SHARP), deletion of IDR domain and insertion of alternative IDR domain from FUS protein (FUS-ΔIDR-SHARP). c) Schematic showing experimental workflow for generating and enriching stable SHARP rescue mESCs (FL-SHARP, ΔRRM-SHARP, ΔIDR-SHARP, FUS-ΔIDR-SHARP) using constructs from Extended Data Fig. 3b. d) Representative images of SHARP enrichment (HALO, green) over the Xi (anti-Ezh2 immunofluorescence, magenta) in female mESCs containing dox-inducible Xist, genetic deletion of SHARP, and stable integrations of HALO-tagged FL-SHARP, ΔRRM-SHARP, or ΔIDR-SHARP (n > 10 cells per condition). Xist and SHARP rescue constructs induced with doxycycline for 72 hours; images shown as Z-sections; scale bars 10 μm e) Diagram of image analysis workflow for quantifying SHARP enrichment over the Xi (Fig. 3b).
Extended Data Fig. 4 SHARP binding to RNA (via RRMs) and formation of assemblies (via IDRs) are both required for chromosome-wide gene silencing.
a) Schematic of mouse X chromosome showing the locations of the various genes probed in RNA-FISH experiments. b) Frequency of Xist induction (left) and X chromosome ploidy (right) in wildtype and SHARP-KO mESCs based on quantification of RNA-FISH images. c) Quantification of RNA-FISH images (Fig. 4b) representing the frequency of cells containing two, one, or zero actively transcribed alleles. Left to right: wildtype (-dox); wildtype (+dox); dox-induced SHARP-KO; dox-induced, auxin-treated SHARP-AID female mESCs. d) RNA-FISH images from SHARP-KO female mESCs containing stable integrations of (left to right): FL-SHARP, ΔRRM-SHARP, ΔIDR-SHARP, or FUS-ΔIDR-SHARP after >72 hours of dox induction. Cells were stained for DAPI (blue) and probed for Xist (magenta), escape gene Kdm5c (yellow), and silenced genes Gpc4 or MeCP2 (green). Images shown as max projections; scale bars 10 μm; total cell numbers (Extended Data Fig. 4e) from three independent experiments. e) Quantification of RNA-FISH images (Extended Data Fig. 4d) representing the frequency of cells containing two, one, or zero actively transcribed alleles for the various SHARP rescue constructs in SHARP-KO female mESCs. f) RNA-FISH images from SHARP-AID female mESCs containing stable integrations of (left to right): FL-SHARP, ΔRRM-SHARP, ΔIDR-SHARP, or FUS-ΔIDR-SHARP after >72 hours of dox induction. Cells were stained for DAPI (blue) and probed for Xist (magenta), escape gene Kdm5c (yellow), and silenced genes Gpc4 or MeCP2 (green). Images shown as max projections; scale bars 10 μm; total cell numbers (Extended Data Fig. 4g) from three independent experiments. g) Quantification of RNA-FISH images (Extended Data Fig. 4f) representing the frequency of cells containing two, one, or zero actively transcribed alleles for the various SHARP rescue constructs in SHARP-KO female mESCs.
Extended Data Fig. 5 Low Xist expression levels limit its ability to spread to autosomes.
a) Top: schematic depicting expected ratios of SHARP to Xist based on increasing concentration of Xist RNA; bottom: diagrams illustrating non-stoichiometric and stoichiometric gene silencing by Xist. b) Randomized Xist localization in simulated single cells (permutations) compared to experimental data. Top: Xist localization after 48 hours of RA-differentiation in female mESCs from bulk RAP-DNA experiments16; middle: gene density across X, only genes that undergo XCI are plotted; bottom: randomized localization of 200 Xist molecules in 1000 random “cells”; Xist represented by grey squares. c) Simulation visualizing localization of Xist molecules over genes that undergo transcriptional silencing during XCI across 20 cells (zoom-in from Extended Data Fig. 5b). Percent reflects proportion of Xist molecules overlapping the gene in all 1000 permutations. d) Expression levels of Xist in female mESCs treated with increasing dox concentrations as measured by RT-qPCR. e) Comparison of Xist occupancy (black lines; 3x dox RAP data) and DNA contact frequency with Xist locus (red lines; SPRITE data61) across 1 Mb DNA regions of chr8. Xist enrichment and 3D distance are normalized to their median coverage across chromosome 8 to place them on the same relative scale. f) Scatterplot representing the frequency of 3D contacts between each 1 Mb autosomal bin with the Xist locus (SPRITE data61; y axis) and DNA sites enriched by Xist when expressed at high concentration (3x dox; RAP-DNA data; X-axis). g) Relative Xist expression upon RA-induced differentiation (left) and dox-induction (right) of SHARP-AID mESCs in the absence or presence of auxin as measured by RT-qPCR (primer pair Xist 2). Dots represent individual replicates; n = 6 for differentiation conditions; n = 3 for dox induction conditions; data represented as mean ± s.d. h) Relative Xist expression in RA differentiated wildtype cells that do not contain dox-inducible Xist (left) and TX cells with dox-inducible Xist (right) as measured by RT-qPCR (primer pairs Xist 1, 2). i) Left: images representing Xist territories in wildtype cells without dox-inducible Xist and TX cells with dox-inducible Xist; total cell numbers (right) from two independent experiments. Right: quantification of percent of nucleus occupied by Xist in the same cell lines as measured by RNA-FISH (Xist) and DAPI staining (nucleus); scale bars 10 μm. Dots represent individual replicates; red bars represent median.
Supplementary information
Supplementary Information
Supplementary Note, Tables 1–3 and legends for Videos 1–3
Supplementary Video 1
Live imaging of HEK293T cell expressing FL-SHARP labeled with eGFP representing fluorescent intensity over time.
Supplementary Video 2
Live imaging of HEK293T cell expressing FL-SHARP labeled with eGFP representing 3D surface reconstructions based on fluorescent intensities over time, volume color-coded.
Supplementary Video 3
Live imaging of HEK293T cell expressing ΔIDR-SHARP labeled with eGFP representing fluorescent intensity over time.
Source data
Source Data Fig. 1
Intensity quantification for Xist, SHARP over Xi territory.
Source Data Fig. 2
Intensity per pixel for FL-SHARP titration and for FL-SHARP versus ∆IDR-SHARP.
Source Data Fig. 3
Normalized enrichment scores for SHARP over Xi territory.
Source Data Fig. 4
Spot scoring for RNA-FISH.
Source Data Fig. 5
Xist percent nucleus data; RT-qPCR fold change calculations.
Source Data Extended Data Fig. 3
Unprocessed gel image for SHARP-KO mESC line.
Rights and permissions
About this article
Cite this article
Jachowicz, J.W., Strehle, M., Banerjee, A.K. et al. Xist spatially amplifies SHARP/SPEN recruitment to balance chromosome-wide silencing and specificity to the X chromosome. Nat Struct Mol Biol 29, 239–249 (2022). https://doi.org/10.1038/s41594-022-00739-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00739-1
This article is cited by
-
Transcription regulation by long non-coding RNAs: mechanisms and disease relevance
Nature Reviews Molecular Cell Biology (2024)
-
Looking back at 30 years of Nature Structural & Molecular Biology
Nature Structural & Molecular Biology (2024)
-
RNA binding by the glucocorticoid receptor attenuates dexamethasone-induced gene activation
Scientific Reports (2023)
-
RNA polymerase II depletion from the inactive X chromosome territory is not mediated by physical compartmentalization
Nature Structural & Molecular Biology (2023)
-
The SPOC domain is a phosphoserine binding module that bridges transcription machinery with co- and post-transcriptional regulators
Nature Communications (2023)