Abstract
HIV-1 immunotherapy with a combination of first generation monoclonal antibodies was largely ineffective in pre-clinical and clinical settings and was therefore abandoned1,2,3. However, recently developed single-cell-based antibody cloning methods have uncovered a new generation of far more potent broadly neutralizing antibodies to HIV-1 (refs 4, 5). These antibodies can prevent infection and suppress viraemia in humanized mice and nonhuman primates, but their potential for human HIV-1 immunotherapy has not been evaluated6,7,8,9,10. Here we report the results of a first-in-man dose escalation phase 1 clinical trial of 3BNC117, a potent human CD4 binding site antibody11, in uninfected and HIV-1-infected individuals. 3BNC117 infusion was well tolerated and demonstrated favourable pharmacokinetics. A single 30 mg kg−1 infusion of 3BNC117 reduced the viral load in HIV-1-infected individuals by 0.8–2.5 log10 and viraemia remained significantly reduced for 28 days. Emergence of resistant viral strains was variable, with some individuals remaining sensitive to 3BNC117 for a period of 28 days. We conclude that, as a single agent, 3BNC117 is safe and effective in reducing HIV-1 viraemia, and that immunotherapy should be explored as a new modality for HIV-1 prevention, therapy and cure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mehandru, S. et al. Adjunctive passive immunotherapy in human immunodeficiency virus type 1-infected individuals treated with antiviral therapy during acute and early infection. J. Virol. 81, 11016–11031 (2007)
Trkola, A. et al. Delay of HIV-1 rebound after cessation of antiretroviral therapy through passive transfer of human neutralizing antibodies. Nature Med. 11, 615–622 (2005)
Armbruster, C. et al. Passive immunization with the anti-HIV-1 human monoclonal antibody (hMAb) 4E10 and the hMAb combination 4E10/2F5/2G12. J. Antimicrob. Chemother. 54, 915–920 (2004)
Klein, F. et al. Antibodies in HIV-1 vaccine development and therapy. Science 341, 1199–1204 (2013)
West, A. P., Jr et al. Structural insights on the role of antibodies in HIV-1 vaccine and therapy. Cell 156, 633–648 (2014)
Klein, F. et al. HIV therapy by a combination of broadly neutralizing antibodies in humanized mice. Nature 492, 118–122 (2012)
Barouch, D. H. et al. Therapeutic efficacy of potent neutralizing HIV-1-specific monoclonal antibodies in SHIV-infected rhesus monkeys. Nature 503, 224–228 (2013)
Shingai, M. et al. Antibody-mediated immunotherapy of macaques chronically infected with SHIV suppresses viraemia. Nature 503, 277–280 (2013)
Horwitz, J. A. et al. HIV-1 suppression and durable control by combining single broadly neutralizing antibodies and antiretroviral drugs in humanized mice. Proc. Natl Acad. Sci. USA 110, 16538–16543 (2013)
Moldt, B. et al. Highly potent HIV-specific antibody neutralization in vitro translates into effective protection against mucosal SHIV challenge in vivo . Proc. Natl Acad. Sci. USA 109, 18921–18925 (2012)
Scheid, J. F. et al. Sequence and structural convergence of broad and potent HIV antibodies that mimic CD4 binding. Science 333, 1633–1637 (2011)
Scheid, J. F. et al. Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature 458, 636–640 (2009)
Scharf, L. et al. Antibody 8ANC195 reveals a site of broad vulnerability on the HIV-1 envelope spike. Cell Rep. 7, 785–795 (2014)
Falkowska, E. et al. Broadly neutralizing HIV antibodies define a glycan-dependent epitope on the prefusion conformation of gp41 on cleaved envelope trimers. Immunity 40, 657–668 (2014)
Klein, F. et al. Somatic mutations of the immunoglobulin framework are generally required for broad and potent HIV-1 neutralization. Cell 153, 126–138 (2013)
Balazs, A. B. et al. Antibody-based protection against HIV infection by vectored immunoprophylaxis. Nature 481, 81–84 (2012)
Glassman, P. M. & Balthasar, J. P. Mechanistic considerations for the use of monoclonal antibodies for cancer therapy. Cancer Biol. Med. 11, 20–33 (2014)
Nettles, R. E. et al. Pharmacodynamics, safety, and pharmacokinetics of BMS-663068, an oral HIV-1 attachment inhibitor in HIV-1-infected subjects. J. Infect. Dis. 206, 1002–1011 (2012)
Nimmerjahn, F. & Ravetch, J. V. Antibody-mediated modulation of immune responses. Immunol. Rev. 236, 265–275 (2010)
Euler, Z. et al. Cross-reactive neutralizing humoral immunity does not protect from HIV type 1 disease progression. J. Infect. Dis. 201, 1045–1053 (2010)
Matsushita, S., Yoshimura, K., Ramirez, K. P., Pisupati, J. & Murakami, T. Passive transfer of neutralizing mAb KD-247 reduces plasma viral load in patients chronically infected with HIV-1. AIDS 29, 453–462 (2015)
Liao, H. X. et al. Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 496, 469–476 (2013)
Wei, X. et al. Antibody neutralization and escape by HIV-1. Nature 422, 307–312 (2003)
Doria-Rose, N. A. et al. Developmental pathway for potent V1V2-directed HIV-neutralizing antibodies. Nature 509, 55–62 (2014)
Klein, F. et al. Enhanced HIV-1 immunotherapy by commonly arising antibodies that target virus escape variants. J. Exp. Med. 211, 2361–2372 (2014)
Deeks, S. G., Lewin, S. R. & Havlir, D. V. The end of AIDS: HIV infection as a chronic disease. Lancet 382, 1525–1533 (2013)
Bournazos, S. et al. Broadly neutralizing anti-HIV-1 antibodies require Fc effector functions for in vivo activity. Cell 158, 1243–1253 (2014)
Diskin, R. et al. Increasing the potency and breadth of an HIV antibody by using structure-based rational design. Science 334, 1289–1293 (2011)
Halper-Stromberg, A. et al. Broadly neutralizing antibodies and viral inducers decrease rebound from HIV-1 latent reservoirs in humanized mice. Cell 158, 989–999 (2014)
Zhou, T. et al. Multidonor analysis reveals structural elements, genetic determinants, and maturation pathway for HIV-1 neutralization by VRC01-class antibodies. Immunity 39, 245–258 (2013)
Somsouk, M. et al. The immunologic effects of mesalamine in treated HIV-infected individuals with incomplete CD4+ T cell recovery: a randomized crossover trial. PLoS ONE 9, e116306 (2014)
Pereyra, F. et al. The major genetic determinants of HIV-1 control affect HLA class I peptide presentation. Science 330, 1551–1557 (2010)
Montefiori, D. C. Evaluating neutralizing antibodies against HIV, SIV, and SHIV in luciferase reporter gene assays. Curr. Protoc. Immunol. 12, Unit 12.11. (2005)
Li, M. et al. Human immunodeficiency virus type 1 env clones from acute and early subtype B infections for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 79, 10108–10125 (2005)
van 't Wout, A. B., Schuitemaker, H. & Kootstra, N. A. Isolation and propagation of HIV-1 on peripheral blood mononuclear cells. Nature Protocols 3, 363–370 (2008)
Laird, G. M. et al. Rapid quantification of the latent reservoir for HIV-1 using a viral outgrowth assay. PLoS Pathog. 9, e1003398 (2013)
Salazar-Gonzalez, J. F. et al. Deciphering human immunodeficiency virus type 1 transmission and early envelope diversification by single-genome amplification and sequencing. J. Virol. 82, 3952–3970 (2008)
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010)
West, A. P., Jr Computational analysis of anti-HIV-1 antibody neutralization panel data to identify potential functional epitope residues. Proc. Natl Acad. Sci. USA 110, 10598–10603 (2013)
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Acknowledgements
We thank all individuals who participated in this study. We thank the Rockefeller University Hospital Clinical Research Support Office and nursing staff, as well as T. Kümmerle, C. Wyen, and L. Siegel for help with recruitment; S. Kiss for ophthalmologic assessments; F. Maldarelli and J. Lifson for single-copy analysis and B. Freemire for technical assistance; J. Pring, A. Almaktari, C. Unsen, S. Hadrigan, E. Thomas, H. Gruell, D. Gillor, and U. Sandaradura de Silva for sample processing and study coordination, and N. Rodziewicz for help with HIV-1 culturing. We thank A. Louie and C. Conrad for help with regulatory submissions; L. Thomas for IND-enabling studies, L. Vitale for cell line and anti-idiotype antibody development, B. Riordan, A. Rayo and J. Andreozzi for anti-idiotype ELISA method development and sample analysis, R. Hammond for process development, and S. DiSciullo for 3BNC117 manufacturing; J. Perry for performing neutralization assays. We thank M. Suarez-Farinas for support with statistical analysis; P. Fast and H. Park for clinical monitoring; and E. Gotschlich and B. Coller for input on study design. J.C.C.L. is supported by an award from CNPq “Ciencia sem Fronteiras” Brazil (248676/2013-0). This work was supported in part by the Bill and Melinda Gates Foundation Collaboration for AIDS Vaccine Discovery (CAVD) Grants OPP1033115 (M.C.N.), OPP1092074 (M.C.N.), OPP1040753 (A.P.W.) and OPP1032144 (M.S.S.), by grant #UL1 TR000043 from the National Center for Advancing Translational Sciences (NCATS), by a grant from the Robertson Foundation to M.C.N., in part with Federal funds from the NCI/NIH, under Contract No. HHSN261200800001E, and a grant from the German Center for Infection Research (DZIF) to G.F. 3BNC117 was generated from a subject in the International HIV Controller Study, supported by the Mark and Lisa Schwartz Foundation and CAVD Grant 43307. M.C.N. and B.D.W. are Howard Hughes Medical Institute Investigators. The authors declare no competing financial interests.
Author information
Authors and Affiliations
Contributions
M.C. and F.K. planned and implemented the study, analysed the data, and wrote the manuscript; J.C.C.L. performed sequence analyses, and contributed to writing the manuscript; M.S.S. performed TZM.bl neutralization assays; A.P.W. assisted with sequence analyses; N.B., G.K., S.B.-A., M.W.-P., M.P., L.A.B. implemented the study; L.N. and M.B. performed cloning and sequencing; I.S., C.L. coordinated sample processing; T.H. performed ELISA assays; and R.J.G. performed single copy assays. T.K. was responsible for 3BNC117 manufacture and provided regulatory guidance; J.F.S., B.D.W., J.A.H. contributed to study design and helped with the manuscript; R.M.G. contributed to study design; and G.F. and S.J.S. contributed to study design and implementation. M.C.N. planned and implemented the study, analysed the data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 HIV-1 neutralizing activity of 3BNC117.
a, Summary of 3BNC117 neutralizing in vitro activity based on 237 HIV-1 isolates comprising 6 different clades. Data were retrieved from the ‘AntibodyDatabase’ by A.P.W. (ref. 39). b, Illustration of the fraction (that is, % coverage; y axis) of HIV-1 isolates that are neutralized at a given IC50 (μg ml−1; x axis) using the same data set.
Extended Data Figure 2 CD4+ and CD8+ T-cell counts before and after 3BNC117 infusion.
a, Absolute numbers (cells μl−1) of CD4+ and CD8+ T-cell counts of all enrolled HIV-1-infected participants at screen, on the day of 3BNC117 infusion (day 0), and at day 28 after infusion. b, Percentage of CD4+ and CD8+ T cells for the same subjects and time points. Mean and standard deviation are indicated in red and black, respectively. No significant differences between pre-infusion and post infusion levels were detected by using one-way ANOVA.
Extended Data Figure 3 3BNC117 serum concentration and activity in single subjects.
a, b, Serum levels of 3BNC117 in all uninfected (a) and HIV-1-infected (b) individuals that received 1, 3, 10, or 30 mg kg−1 3BNC117 at day 0. Antibody levels were measured by a sandwich ELISA using an anti-3BNC117 specific antibody (green) or by measuring the 3BNC117 serum activity in a TZM.bl neutralization assay (blue).
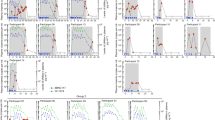
Extended Data Figure 4 3BNC117 sensitivity and changes in viraemia in 2 ART-treated subjects.
Both subjects were on ART when enrolled in the study and received a single dose (2B2, 3 mg kg−1; 2C2, 10 mg kg−1) of 3BNC117 at day 0. The left y axis shows log10 change in viraemia from baseline, and right y axis shows antibody level measured in ELISA. Blue line reflects change in VL and dotted grey line antibody level. Numbers indicate IC50 values for 3BNC117 of autologous viral isolates measured by TZM.bl assay, colour-coded as indicated on the right.
Extended Data Figure 5 Correlating viral decay with 3BNC117 sensitivity and starting viral load.
a, Maximum decline in viral load in ART-untreated HIV-1-infected participants with baseline 3BNC117-sensitive viruses (IC50 < 1 μg ml−1) versus pre-treatment (day 0) viral load (Pearson coefficient r = 0.72 P = 0.03; Spearman coefficient rho = 0.78, P = 0.02). b, Maximum drop in viral load in HIV-1-infected and viremic individuals receiving a 10 or 30 mg kg−1 dose of 3BNC117 (y axis) versus baseline autologous virus sensitivity to 3BNC117 (x axis; Pearson coefficient r = 0.69 P = 0.03; Spearman coefficient rho = 0.41, P = 0.23).
Supplementary information
Supplementary Figures
This file contains Supplementary Figure 1, which shows the HIV-1 envelope sequence analysis. (PDF 382 kb)
Rights and permissions
About this article
Cite this article
Caskey, M., Klein, F., Lorenzi, J. et al. Viraemia suppressed in HIV-1-infected humans by broadly neutralizing antibody 3BNC117. Nature 522, 487–491 (2015). https://doi.org/10.1038/nature14411
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14411
This article is cited by
-
A classification algorithm based on dynamic ensemble selection to predict mutational patterns of the envelope protein in HIV-infected patients
Algorithms for Molecular Biology (2023)
-
Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Nature Communications (2023)
-
Mucosal application of the broadly neutralizing antibody 10-1074 protects macaques from cell-associated SHIV vaginal exposure
Nature Communications (2023)
-
Chemical inhibition of DPP9 sensitizes the CARD8 inflammasome in HIV-1-infected cells
Nature Chemical Biology (2023)
-
Computer-aided Affinity Enhancement of a Cross-reactive Antibody against Dengue Virus Envelope Domain III
Cell Biochemistry and Biophysics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.