Abstract

Cancer is a disease potentiated by mutations in somatic cells. Cancer mutations are not distributed uniformly along the human genome. Instead, different human genomic regions vary by up to fivefold in the local density of cancer somatic mutations1, posing a fundamental problem for statistical methods used in cancer genomics. Epigenomic organization has been proposed as a major determinant of the cancer mutational landscape1,2,3,4,5. However, both somatic mutagenesis and epigenomic features are highly cell-type-specific6,7. We investigated the distribution of mutations in multiple independent samples of diverse cancer types and compared them to cell-type-specific epigenomic features. Here we show that chromatin accessibility and modification, together with replication timing, explain up to 86% of the variance in mutation rates along cancer genomes. The best predictors of local somatic mutation density are epigenomic features derived from the most likely cell type of origin of the corresponding malignancy. Moreover, we find that cell-of-origin chromatin features are much stronger determinants of cancer mutation profiles than chromatin features of matched cancer cell lines. Furthermore, we show that the cell type of origin of a cancer can be accurately determined based on the distribution of mutations along its genome. Thus, the DNA sequence of a cancer genome encompasses a wealth of information about the identity and epigenomic features of its cell of origin.
Similar content being viewed by others
Main
Recent studies have begun to address the underlying causes of cancer mutational heterogeneity by comparing mutation rate variation to the distribution of sequence features, gene expression and epigenetic marks along the genome2,3,4,5. A major limitation of previous studies was their uniform treatment of mutations from different cancers, and their consideration of epigenetic marks from a single cell type, usually a cell type different from the cancer tissue of origin. However, cancer is far from being a disease of uniform origin, progression and cell biology. Instead, different cancer types differ in their overall mutation rates, their predominant mutation types, and the distribution of mutations along their genomes1. Substantial variation also exists in the epigenomic landscape of different tissues, specifically in patterns of chromatin accessibility, histone modifications, gene expression and DNA replication timing7,8,9,10. Full understanding of the factors contributing to mutational heterogeneity in cancer genomes thus requires the evaluation of the relationship between multiple epigenetic marks and mutation patterns in a cell-type-specific manner.
We analysed a total of 173 cancer genomes from eight different cancer types that represent a wide range of tissues of origin, carcinogenic mechanisms, and mutational signatures: melanoma11, multiple myeloma12, lung adenocarcinoma13, liver cancer14, colorectal cancer15, glioblastoma16, oesophageal adenocarcinoma17 and lung squamous cell carcinoma18. Regional variations in mutation density appeared similar, although not identical, among the different cancer types (Extended Data Fig. 1).
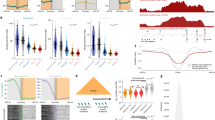
We compared the genomic distribution of mutations in these cancer genomes to 424 epigenetic features that were measured by the Epigenome Roadmap consortium9. These features were derived from 106 different cell types from 45 different tissue types, encompassing the established or likely cell types of origin of most of the cancer types that we investigated (Methods and Extended Data Fig. 2). Notably, these data derive from primary human cells and tissues rather than malignant cell lines. These epigenetic features comprised eight different types of variables, including DNase I hypersensitivity (a global measure of chromatin accessibility)7 and various histone modifications. An example of the variation in mutation density along chromosomes at a 1 Mb scale together with the density of DNase I hypersensitive sites (DHSs) is shown in Fig. 1. In this case, as in most other cases (see later), epigenomics features indicative of active chromatin and transcription were associated with low mutation density, whereas repressive chromatin features were associated with regions of high mutation density. Notably, these statistical associations do not necessarily imply causal effects of individual chromatin features, nor point to specific biological mechanisms.
a, The density of C > T mutations in melanoma alongside a 100-kb window profile of melanocyte chromatin accessibility (‘DNase I accessibility index’; shown in normalized, reverse scale; high values correspond to less accessible chromatin and vice versa). b, The number of mutations per megabase in melanoma versus DHS density, for three types of skin cells. The Spearman’s rank correlations (ρ) are shown on each panel. c, The normalized density of mutations in liver cancer and melanoma genomes as a function of density quintiles of H3K4me1 marks in liver cells and in melanocytes. For both cancer genomes, mutation density depends only on H3K4me1 marks measured in the cell of origin.
The comparison of individual epigenomic features with local mutation density revealed that the genomic distribution of chromatin features corresponding to the tumour’s cell type of origin is more strongly associated with local mutation density than the distribution of features found in unrelated cell types. For example, DHSs from melanocytes explained a substantially larger fraction of the variance in melanoma mutation density than DHSs from other cell types, even from the same tissue (skin) (Fig. 1b). As another example, even though H3K4me1 marks in melanocytes and hepatocytes are highly correlated (r = 0.8), the distribution of mutations in liver cancer followed the levels of H3K4me1 in hepatocytes but not in melanocytes, whereas melanoma mutations correlated with the levels of H3K4me1 in melanocytes but not in hepatocytes (Fig. 1c).
This initial observation suggested that the impact of chromatin structure on local mutation density is highly cell-type specific. The comprehensive representation of different cell types in the Epigenome Roadmap could thus enable improved prediction accuracy of mutations compared to previous studies. To rigorously quantify the contribution of different chromatin marks and gene expression to regional mutation density, and the extent of cell type specificity, we used Random Forest regression (Methods).
Remarkably, epigenetic features, together with replication timing measured in ENCODE consortium cell lines19, collectively accounted for 74–86% of the variance in mutation density in seven cancer types (Fig. 2a). In glioblastoma, for which fewer mutations were available for the analysis, 55% of the variance in mutation density could be explained. This is substantially higher than in earlier studies4 and indicates that, at least for these cancer types, we have identified a set of epigenetic variables and cell types that almost fully predict the mutational variability along the genome. This enhanced prediction accuracy was not simply due to the larger size of the training data relative to previous studies, as the predictive ability dropped by only ∼2–6% when only 10% of the data was used (Extended Data Fig. 3).
Pearson correlation between observed and predicted mutation densities along chromosomes is shown. a, Actual versus predicted mutation densities in eight cancers. b, c, Prediction accuracy represented as mean ± s.e.m. (estimated using tenfold cross-validation). Panels show prediction accuracy for all mutations and for nucleotide changes predominant in the corresponding cancer (b), and prediction accuracy in lung adenocarcinoma genomes stratified by smoking history and predominant nucleotide changes (G>T or C>T) (c).
Prediction accuracy in individual samples is expected to be lower than in samples pooled by cancer type due to tumour heterogeneity, sampling variance, and a lower number of mutations available for the analysis (Extended Data Fig. 4). To evaluate the influences of these variables on the prediction ability of the Random Forest model, we simulated mutation data sets of variable sizes generated by the model itself, and compared the prediction accuracy of simulated and real data as a function of the number of mutations. For most samples, epigenomic features explained most (on average 70%) of the maximally predicted variance (Extended Data Fig. 5), and more than was explained by earlier studies2,4 when matching data set sizes. As a point of direct comparison with an earlier study2 that did not use cell-type-specific chromatin marks, our model explained 50% of the variance in mutation density in the melanoma cell line COLO829 (ref. 20), compared with 29% in the earlier study.
The prediction accuracy was similar whether testing for all mutations or only the mutations of the predominant type (Extended Data Fig. 6) in each cancer type1,21 (Fig. 2b). A notable exception was lung adenocarcinoma, where a larger fraction of the variance could be explained for G > T mutations associated with smoking1,13,22 than for C > T mutations. This difference was observed for both samples with G > T transversions13 and C > T transitions as the leading mutational sources (Fig. 2c).
Prediction accuracy was fully explained by chromatin features, with gene expression and nucleotide content not providing any further improvement to the accuracy of the model. Even though gene expression has been unequivocally demonstrated to influence mutation density, chromatin features appear to be statistically stronger predictors (Extended Data Fig. 7).
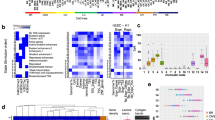
When considering individual feature contributions to mutation rate prediction, between six and nineteen variables passed the significance threshold in any individual cancer type. There was a sweeping association between cancer mutations and chromatin marks measured in the cell type of origin of each cancer (Fig. 3a). For instance, six out of the top ten features explaining variation in melanoma mutation density were derived from melanocytes (Figs 1 and 3b). Similarly, seven out of the nine top features explaining mutational profiles in liver cancer were measured in liver cells. Comparable results were obtained for multiple myeloma, colorectal adenocarcinoma and glioblastoma, where most of the significant features were measured in haematopoietic, intestine mucosa and brain tissues, respectively. For oesophageal adenocarcinoma, the top predictors were chromatin features derived from stomach mucosa rather than from oesophageal tissues; this is expected given that the analysed oesophageal adenocarcinomas were triggered by Barrett’s oesophagus cells that resemble stomach epithelial cells23. Lung adenocarcinoma and lung squamous cell carcinoma were the only exceptions in that the top predictors were scattered among different tissue groups; the lack of tissue specificity in these cases likely results from the absence of epigenomic marks from normal lung epithelial cells in our data set.
a, Features (blue rectangles) that significantly contributed to the predictions in at least one cancer type (see Methods). b, Melanoma mutation density versus the density of chromatin modifications in melanocytes. c, Prediction accuracy (mean ± s.e.m. estimated using tenfold cross-validation) of models separately trained on features from different tissues for each cancer type. Red bars, tissues with the highest prediction accuracy. Red line, prediction accuracy when using all 424 epigenetic features. d, Comparison of predictions accuracies of liver cancer mutation density from features of normal liver cells versus cancer cells (HepG2). e, Mutation density in COLO829 melanoma cell line versus DHS density in COLO829, melanocytes, DHSs specific to COLO829 (not observed in melanocytes) and DHSs specific to melanocytes (not observed in COLO829). Spearman’s rank correlation coefficient is given for each comparison.
The results of the Random Forest regression were confirmed using backward feature selection to identify the minimal set of epigenetic predictors of mutations in each cancer type (Methods). As few as three to five features were sufficient to capture the variance explained by the full set of 424 different features (Extended Data Fig. 8), and in all cancers besides lung (as above), most of these features were derived from the corresponding cell types of origin. As a more direct test, we grouped all epigenomic data by cell or tissue type and compared the collective explanatory power of chromatin features derived from the cell types of origin versus unmatched cell types. The results of this analysis confirmed the cell type specificity of the association between chromatin features and mutation density (Fig. 3c).
The above results pose a key question on whether epigenomic features derived from the cell type of origin are the strongest determinants of cancer mutations or whether they simply serve as the best available proxies to the chromatin organization of the corresponding malignant cells. The availability of epigenomic data for the liver cancer cell line HepG2 (ref. 8) and for melanoma cell lines made it possible to directly address this question. Surprisingly, in both cases, epigenomic features from the cell type of origin resulted in a higher prediction accuracy than those from the cancer cell lines. The Random Forest predictor trained on chromatin features of HepG2 was less accurate in predicting the liver cancer mutation density than the analogous predictor trained on features of hepatocytes (Fig. 3d). Similarly, chromatin accessibility in melanocytes was a much better predictor of mutation density in the COLO829 melanoma cell line (Fig. 3e and Extended Data Fig. 9). Thus, chromatin features associated with carcinogenesis do not determine cancer mutations to the same extent as chromatin features of the cells of origin. We envision two potential explanations for this observation. First, most of the somatic mutations observed in cancers may arise before the epigenetic changes linked to neoplastic progression. Second, advanced tumours may undergo specific epigenetic changes that distinguish them from other tumours of the same type.
Taken together, the above results strongly suggest that the cell of origin of an individual tumour sample could be predicted from its mutation pattern alone. Mutation profiles of individual samples cluster according to cancer type, and, consequently cell of origin (Fig. 4a). We developed a straightforward predictor based on enrichment of epigenomic variables from a single cell type among the top 20 variables selected by the Random Forest analysis. This approach classified 88% of melanoma, colorectal, liver, multiple myeloma, oesophageal and glioblastoma cancer genomes to melanocytes, colonic mucosa, liver, haematopoietic, stomach mucosa and brain tissues, respectively (Fig. 4b). Thus, mutational patterns contain sufficient information for identifying the cell type of origin of a tumour. We propose that sequencing the DNA of a tumour of unknown primary origin can allow the precise identification or categorization of the cell type of origin of that tumour.
a, Principal coordinate analysis (PCOA) of the distribution of mutations in individual cancer genomes. Filled circles represent cancers for which the correct cell type of origin was identified. b, The accuracy of cell type of origin prediction for individual cancer genomes: the number of cancer samples that were assigned to the correct (solid colours) or incorrect (textures) cell types of origin based on their mutation profile.
Traditionally, statistical prediction in cancer has made use of gene expression data. We therefore constructed an analogous predictor of cell of origin using RNA sequencing data from 167 glioblastoma multiforme and 370 skin cutaneous melanoma samples24. This predictor achieved accuracies of 78% and 57% on these cancer types, slightly lower than the mutation-based predictor. Although these two classifiers are not directly comparable, it is clear that genome sequence carries at least the same amount of information about the cell of origin as gene expression data does.
In conclusion, our observations suggest that cancer mutation density is linked to the epigenomic profile in a highly cell-type-specific manner. Thus, DNA sequence is informative about the origin of an individual tumour. The accumulating epigenomic data on human cell types opens the perspective for accurate prediction of the cell of origin of a cancer from its genome sequence.
Methods
Data
We divided the human genome into 1 Mb regions, excluding regions overlapping centromeres and telomeres, as well as regions where the fraction of uniquely mappable base pairs was lower than 0.92. We calculated the mean signal for different histone modifications, DNase I hypersensitivity and replication timing in different cell types, and used these 424 features to predict mutation density along the genome in eight different cancer types (see below).
We calculated mutation density by obtaining data for 173 individual cancer genomes, belonging to eight cancer types: melanoma (25 genomes)11, lung adenocarcinoma (24 genomes)13, lung squamous cell carcinoma (12 genomes)18, oesophageal adenocarcinoma (9 genomes)17, liver (64 genomes)14, multiple myeloma (23 genomes)12, colorectal cancer15 (CRC, 9 genomes) and glioblastoma (7 genomes)16. The whole genome of the COLO-829 cell line has been sequenced by the Sanger Institute. The COLO-829 cell line was derived from metastatic tissue. The liver cancers were sequenced by the National Cancer Center Research Institute in Japan. The mutation lists for the COLO829 cell line and liver cancer that we used in this study can be found at (http://dcc.icgc.org/repository/legacy_data_releases/version_07/) under the folders Malignant_Melanoma-WTSI-UK (COLO-829) and Liver_Cancer-NCC-JP. The rest of the genomes were sequenced and analysed by the Broad Institute and called using MuTect25 (http://www.broadinstitute.org/cancer/cga/mutect).
For each cancer type we counted the overall number of mutations in all individual cancer genomes belonging to that cancer type. We also determined the mutation densities for all possible types of mutations in each cancer types by counting different types of mutations in 1-Mb windows and normalizing for the sequence composition of each window.
We downloaded data for 7 different histone modifications and DNase I hypersensitivity from Epigenomics Roadmap9and ENCODE8 (Extended Data Fig. 1). Epigenomic data are available from the NCBI via the GEO series GSE18927 for University of Washington Human Reference Epigenome Mapping Project at (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE18927). Data used in this study can also be viewed via multiple browsers outlined at the (http://roadmapepigenomics.org/) website.
Fetal tissues were obtained from morphologically normal fetuses by the Birth Defects Research Laboratory in the Department of Paediatrics at the University of Washington, collected under an IRB-approved protocol. Blood cell subsets were collected from fully consenting, normal donors at the Cellular Therapy Laboratory and cGMP Cell Processing Facility under the direction of Shelly Heimfeld at the Fred Hutchinson Cancer Research Center with IRB-approval.
For histone modifications we combined reads for all samples belonging to one cell type and calculated RPKM values for 1-Mb windows along the genome. We also calculated the average number of DNase I hypersensitivity peaks overlapping 1-Mb windows across all samples belonging to a certain cell type. We used BEDOPS26 to map reads and DHS peaks to intervals.
We obtained data for four different Repli-seq experiments from the ENCODE project (Extended Data Fig. 1) and determined replication timing as the average value of wavelet-smoothed signal in each 1-Mb window. Lymphoblastoid cell line replication time was obtained from ref. 27 and averaged over 1-Mb windows along the genome.
To control for the effect of sequence features on mutation density, for each 1-Mb window we also calculated GC content, the number of CpG, GpC, and ApT dinucleotides, and fraction of the window overlapping coding regions, known genes and CpG islands.
To control for the effect of expression on mutation density we downloaded mRNA-seq data from the Epigenomics Roadmap9, for 38 different cell types for which expression data was available (Extended Data Table 1). We combined reads for all samples belonging to one cell type and calculated reads per kilobase per million mapped reads (RPKM)values for the set of all protein coding exons in 1-Mb windows, the set of all protein coding and lncRNA exons in 1-Mb windows, the maximally expressed gene in a 1-Mb window or non-genic regions in 1-Mb windows.
Random Forest regression
Random Forest is a non-parametric machine learning method that combines the output of an ensemble of regression trees to predict the value of a continuous response variable28. The use of multiple regression trees reduces the risk of over-fitting and makes the method robust to outliers and noise in the input data. For each regression tree, a training set of n observations are drawn, with replacement, from the data set. The remaining data (out-of-bag data) constitutes the test set for this tree, and is used to compute the mean squared prediction error of the tree. The prediction for each observation is made by taking the average of predictions over all trees for which the observation was part of the out-of-bag data.
Random Forest provides an internal measure of the importance of different predictor variables, based on out-of-bag data. The mean squared error calculated on the out-of-bag data is recorded in every tree grown in the forest. The values of all the predictor variables are then randomly permuted in all the out-of-bag observations and the mean squared error is computed again. The difference between the two errors is averaged over all the trees, and normalized by the standard error, representing the raw importance score for each variable.
We used Random Forest with 1,000 trees to predict mutation densities in 1Mb non-overlapping windows in the eight different cancer types using 424 predictor variables (epigenetic features and replication timing; Extended Data Fig. 1). We divided the data into ten non-overlapping sets and predicted the number of mutations in each cancer type using tenfold cross-validation. For each sample, the predicted value corresponded to the predicted mutation density when this observation was part of the test set. We used Pearson product-moment correlation to interpret the prediction accuracy. The fraction of variance explained by each model was calculated as the Pearson correlation coefficient squared.
Controlling for the effect of sequence features and expression on prediction accuracy
We created different subsets of features corresponding to chromatin (histone modifications and DNase hypersensitivity, 419 features), replication timing (5 features), sequence (7 features) and expression (38 features). We then used Random Forest regression with tenfold cross-validation to predict mutation density in different cancers, where for each cancer type we trained different models: on each subset of features separately and on combinations of different subsets of features.
Variable importance analysis
Variable importance was calculated for each predictor variable in each cancer type by permuting the variable, that is, randomly shuffling the data values so that the relationship between the response and predictor variables was destroyed. The percent of increase in mean squared error of prediction was then calculated. Since the variable importance can be influenced by both the correlation and the scale of the variables, we calculated the empirical P value of variable importance measures by repeatedly permuting the response variable in Random Forest models, in order to determine the distribution of measured importance values for each predictor variable29. This procedure was repeated 1,000 times, and the number of times in which the importance measure in the original data set was lower or equal to the permuted importance measure was counted; this count represented the P value, with a count of one corresponding to a significance level of P < 0.001.
Feature selection
We applied backward elimination to identify a minimal set of predictors for each cancer type. Backward elimination is a ‘greedy’ algorithm which finds the locally optimal subset of features, but does not guarantee finding the global optimum. However, it is less computationally intensive than searching all possible feature subsets when the number of features (p) is large (in our case p = 424). Initially, we trained a Random Forest with tenfold cross-validation on the complete set of variables and determined the importance of all the variables in the model (the importance was calculated as the mean importance of the variable across 10 rounds of cross-validation). We then ranked the variables according to their importance and determined the top 20 variables. We then sequentially trained 20 models, removing the least important variable at each step, until only one predictor variable was left for training.
Principal coordinate analysis
We used principal coordinate analysis to visualize the dissimilarities in mutation density distributions between individual cancer genomes. Dissimilarity was calculated as 1 – Pearson correlation coefficient, for all possible combinations of individual cancer genomes.
Prediction of tissue of origin for individual cancer genomes
For each individual cancer genome we predicted the density of mutations using Random Forest regression with tenfold cross-validation. We used the full set of features and determined the top 20 features according to the variable importance measure. We then calculated the enrichment of each tissue type among the top 20 features using the hypergeometric test and chose the tissue showing the most significant enrichment as the most likely tissue of origin for the individual cancer genome. We then calculated the percentage of individual cancers where the assigned tissue of origin matched the predicted tissue of origin.
Prediction of cell of origin using gene expression
For each individual cancer we downloaded gene expression data from The Cancer Genome Atlas16 and calculated the expression of the same genes in the 38 cell types for which mRNA-seq data was available from the Epigenomics Roadmap (Extended Data Table 1). For each cancer we trained a Random Forest regression model in which the gene expression values in cancer were used as the response variable and the gene expression in normal cells as the predictors. We identified the predictor variable, which showed the highest value of variable importance in the model and assigned the corresponding cell type as cell of origin of the cancer.
Sample size
No statistical methods were used to predetermine sample size.
Accession codes
References
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013)
Hodgkinson, A., Chen, Y. & Eyre-Walker, A. The large-scale distribution of somatic mutations in cancer genomes. Hum. Mutat. 33, 136–143 (2012)
Liu, L., De, S. & Michor, F. DNA replication timing and higher-order nuclear organization determine single-nucleotide substitution patterns in cancer genomes. Nature Commun. 4, 1502 (2013)
Schuster-Böckler, B. & Lehner, B. Chromatin organization is a major influence on regional mutation rates in human cancer cells. Nature 488, 504–507 (2012)
Woo, Y. H. & Li, W. H. DNA replication timing and selection shape the landscape of nucleotide variation in cancer genomes. Nature Commun. 3, 1004 (2012)
Zhu, J. et al. Genome-wide chromatin state transitions associated with developmental and environmental cues. Cell 152, 642–654 (2013)
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012)
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012)
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature http://dx.doi.org/10.1038/nature14248 (this issue)
Ryba, T. et al. Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res. 20, 761–770 (2010)
Berger, M. F. et al. Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 485, 502–506 (2012)
Chapman, M. A. et al. Initial genome sequencing and analysis of multiple myeloma. Nature 471, 467–472 (2011)
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012)
Totoki, Y. et al. High-resolution characterization of a hepatocellular carcinoma genome. Nature Genet. 43, 464–469 (2011)
Bass, A. J. et al. Genomic sequencing of colorectal adenocarcinomas identifies a recurrent VTI1A–TCF7L2 fusion. Nature Genet. 43 964–968 10.1038/ng.936 (2011)
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013)
Dulak, A. M. et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nature Genet. 45, 478–486 (2013)
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012)
Hansen, R. S. et al. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc. Natl Acad. Sci. USA 107, 139–144 (2010)
Pleasance, E. D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, 191–196 (2010)
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Pleasance, E. D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, 184–190 (2010)
Reid, B. J., Li, X., Galipeau, P. C. & Vaughan, T. L. Barrett’s oesophagus and oesophageal adenocarcinoma: time for a new synthesis. Nature Rev. Cancer 10, 87–101 (2010)
Hudson, T. J. et al. International network of cancer genome projects. Nature 464, 993–998 (2010)
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nature Biotechnol. 31, 213–219 (2013)
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012)
Koren, A. et al. Differential Relationship of DNA replication timing to different forms of human mutation and variation. Am. J. Hum. Genet. 91, 1033–1040 (2012)
Breiman, L. Random Forests. Mach. Learn. 45, 5–32 (2001)
Altmann, A., Tolosi, L., Sander, O. & Lengauer, T. Permutation importance: a corrected feature importance measure. Bioinformatics 26, 1340–1347 (2010)
Acknowledgements
This work was supported by NIH grants R01 MH101244, U54 CA143874 to S.R.S. and U01 ES017156, P01 HL53750, U54 HG007010 to J.A.S. R.K and K.V. acknowledge the Integra-Life Seventh Framework Program (grant number 315997) and the EMBO Young Investigator Program (Installation grant 1431/2006 to K.V.).
Author information
Authors and Affiliations
Contributions
S.R.S., J.A.S., P.P. and R.K. conceived the project and provided leadership. P.P., R.K., A.K., M.S.L., R.S., A.K., R.T., E.R., A.R and K.V. analyzed the data and contributed to scientific discussions. S.R.S, P.P., R.K., A.K. and J.A.S. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Epigenomic data are available from the NCBI via the GEO series GSE18927 for University of Washington Human Reference Epigenome Mapping Project.
Extended data figures and tables
Extended Data Figure 3 Scatter plots of the measured number of somatic mutations per Mb in different cancer genomes versus the number of mutations predicted by the Random Forest algorithm.
The training set consisted of 10% of the data, the remaining 90% was used to test the predictions.
Extended Data Figure 4 Prediction accuracy of the models trained on individual cancers as a function of the number of mutations.
The red line represents the prediction accuracy of the model used to predict the mutation density of samples pooled by cancer type (sum of all mutations in individual cancers of a certain cancer type). N – number of individual cancers per cancer type.
Extended Data Figure 5 Sampling variance.
Red, the squared correlation coefficient (R2) between the observed mutational profile and the profiles predicted by Random Forest. Blue, the maximal attainable variance explained, calculated as the average correlation coefficient squared (R2) between the mutational profiles predicted by Random Forest and 100 simulated mutational profiles modelled as a Poisson distribution with the mean predicted by epigenomic features. N – number of individual cancers per cancer type.
Extended Data Figure 7 Prediction accuracy of models obtained using different subsets of predictor variables.
a, Comparison of the prediction accuracy obtained using the full set of chromatin features, 38 chromatin features measured in cell types for which expression data was available, and expression data. Expression in 1-Mb windows was calculated using mRNA-seq reads mapping to either protein coding exons, protein coding and lncRNA exons, maximally expressed gene or non-genic regions, and normalized by the cumulative length of each of these regions, respectively. Bars represent the mean prediction accuracy; error bars represent standard errors of the mean prediction accuracy estimated using tenfold cross-validation. b, Distribution of the percent of variance explained in 10 folds of cross-validation (n = 10) for models trained on chromatin, replication, expression (non-genic mRNA-seq) or sequence features, or a combination of these subsets of features. Models trained on chromatin features were compared to all other models for a certain cancer type (Wilcoxon rank-sum test). Significant differences, Benjamini–Hochberg-corrected: P < 0.01, P < 0.001. Box plots, band inside the box, median; box, first and third quartiles; whiskers, most extreme values within 1.5 × inter-quartile range from the box; points, outliers.
Extended Data Figure 8 Feature selection by using the backward elimination procedure.
For each cancer type, variables are ordered from top to bottom by decreasing importance. Each bar represents the fraction of variance explained by the model using the corresponding bar and all bars above it. Error bars represent the standard error of the mean variance explained by the model, estimated using tenfold cross-validation. The red line indicates the cutoff needed to achieve the prediction accuracy of the full model – 1 s.e.m. For each cancer type, features measured in related cell lines are shown in red.
Extended Data Figure 9 The number of mutations per megabase in COLO829 cell line versus DHS density in melanoma cell lines, melanocytes, DHSs specific to melanomas and DHSs specific to melanocytes.
Correlation is calculated using the Spearman’s rank correlation coefficient. DHS density in melanoma cell lines corresponds to DHSs measured in 11 melanoma cell lines. DHSs specific to melanomas correspond to DHSs observed in melanomas but not observed in melanocytes. DHSs specific to melanocytes correspond to DHSs observed in melanocytes but not observed in melanomas.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Polak, P., Karlić, R., Koren, A. et al. Cell-of-origin chromatin organization shapes the mutational landscape of cancer. Nature 518, 360–364 (2015). https://doi.org/10.1038/nature14221
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14221
This article is cited by
-
Cell cycle gene alterations associate with a redistribution of mutation risk across chromosomal domains in human cancers
Nature Cancer (2024)
-
Structure-guided design of a selective inhibitor of the methyltransferase KMT9 with cellular activity
Nature Communications (2024)
-
Integrative analysis reveals early epigenetic alterations in high-grade serous ovarian carcinomas
Experimental & Molecular Medicine (2023)
-
Investigating the tissue specificity and prognostic impact of cis-regulatory cancer risk variants
Human Genetics (2023)
-
Carcinoma of unknown primary (CUP): an update for histopathologists
Cancer and Metastasis Reviews (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.