Abstract
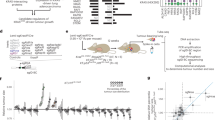
Next-generation sequencing of human tumours has refined our understanding of the mutational processes operative in cancer initiation and progression, yet major questions remain regarding the factors that induce driver mutations and the processes that shape mutation selection during tumorigenesis. Here we performed whole-exome sequencing on adenomas from three mouse models of non-small-cell lung cancer, which were induced either by exposure to carcinogens (methyl-nitrosourea (MNU) and urethane) or by genetic activation of Kras (KrasLA2). Although the MNU-induced tumours carried exactly the same initiating mutation in Kras as seen in the KrasLA2 model (G12D), MNU tumours had an average of 192 non-synonymous, somatic single-nucleotide variants, compared with only six in tumours from the KrasLA2 model. By contrast, the KrasLA2 tumours exhibited a significantly higher level of aneuploidy and copy number alterations compared with the carcinogen-induced tumours, suggesting that carcinogen-induced and genetically engineered models lead to tumour development through different routes. The wild-type allele of Kras has been shown to act as a tumour suppressor in mouse models of non-small-cell lung cancer. We demonstrate that urethane-induced tumours from wild-type mice carry mostly (94%) Kras Q61R mutations, whereas those from Kras heterozygous animals carry mostly (92%) Kras Q61L mutations, indicating a major role for germline Kras status in mutation selection during initiation. The exome-wide mutation spectra in carcinogen-induced tumours overwhelmingly display signatures of the initiating carcinogen, while adenocarcinomas acquire additional C > T mutations at CpG sites. These data provide a basis for understanding results from human tumour genome sequencing, which has identified two broad categories of tumours based on the relative frequency of single-nucleotide variations and copy number alterations1, and underline the importance of carcinogen models for understanding the complex mutation spectra seen in human cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ciriello, G. et al. Emerging landscape of oncogenic signatures across human cancers. Nature Genet. 45, 1127–1133 (2013)
Stratton, M. R., Campbell, P. J. & Futreal, P. A. The cancer genome. Nature 458, 719–724 (2009)
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013)
Johnson, L. et al. Somatic activation of the K-ras oncogene causes early onset lung cancer in mice. Nature 410, 1111–1116 (2001)
You, M., Candrian, U., Maronpot, R. R., Stoner, G. D. & Anderson, M. W. Activation of the Ki-ras protooncogene in spontaneously occurring and chemically induced lung tumours of the strain A mouse. Proc. Natl Acad. Sci. USA 86, 3070–3074 (1989)
Prior, I. A., Lewis, P. D. & Mattos, C. A comprehensive survey of Ras mutations in cancer. Cancer Res. 72, 2457–2467 (2012)
Jackson, E. L. et al. Analysis of lung tumour initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001)
Zhang, Z. et al. Wildtype Kras2 can inhibit lung carcinogenesis in mice. Nature Genet. 29, 25–33 (2001)
To, M. D. et al. Kras regulatory elements and exon 4A determine mutation specificity in lung cancer. Nature Genet. 40, 1240–1244 (2008)
Govindan, R. et al. Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 150, 1121–1134 (2012)
Forkert, P. G. Mechanisms of lung tumourigenesis by ethyl carbamate and vinyl carbamate. Drug Metab. Rev. 42, 355–378 (2010)
Kurowska, M., Labocha-Pawlowska, A., Gnizda, D., Maluszynski, M. & Szarejko, I. Molecular analysis of point mutations in a barley genome exposed to MNU and gamma rays. Mutat. Res. 738–739, 52–70 (2012)
Pfeifer, G. P. Mutagenesis at methylated CpG sequences. Curr. Top. Microbiol. Immunol. 301, 259–281 (2006)
Welch, J. S. et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 150, 264–278 (2012)
Drugan, J. K. et al. Ras interaction with two distinct binding domains in Raf-1 may be required for Ras transformation. J. Biol. Chem. 271, 233–237 (1996)
Spoerner, M., Herrmann, C., Vetter, I. R., Kalbitzer, H. R. & Wittinghofer, A. Dynamic properties of the Ras switch I region and its importance for binding to effectors. Proc. Natl Acad. Sci. USA 98, 4944–4949 (2001)
Fawdar, S. et al. Targeted genetic dependency screen facilitates identification of actionable mutations in FGFR4, MAP3K9, and PAK5 in lung cancer. Proc. Natl Acad. Sci. USA 110, 12426–12431 (2013)
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013)
Di Benedetto, M. et al. Mutation analysis of the 8p22 candidate tumour suppressor gene ATIP/MTUS1 in hepatocellular carcinoma. Mol. Cell. Endocrinol. 252, 207–215 (2006)
Frank, B. et al. Copy number variant in the candidate tumour suppressor gene MTUS1 and familial breast cancer risk. Carcinogenesis 28, 1442–1445 (2007)
Zuern, C. et al. Down-regulation of MTUS1 in human colon tumours. Oncol. Rep. 23, 183–189 (2010)
Xiao, J. et al. Reduced expression of MTUS1 mRNA is correlated with poor prognosis in bladder cancer. Oncol Lett 4, 113–118 (2012)
Shedden, K. et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nature Med. 14, 822–827 (2008)
To, M. D. et al. Progressive genomic instability in the FVB/KrasLA2 mouse model of lung cancer. Mol. Cancer Res. 9, 1339–1345 (2011)
Ahmed, N. N., Grimes, H. L., Bellacosa, A., Chan, T. O. & Tsichlis, P. N. Transduction of interleukin-2 antiapoptotic and proliferative signals via Akt protein kinase. Proc. Natl Acad. Sci. USA 94, 3627–3632 (1997)
McFadden, D. G. et al. Genetic and clonal dissection of murine small cell lung carcinoma progression by genome sequencing. Cell 156, 1298–1311 (2014)
Nuzum, E. O., Malkinson, A. M. & Beer, D. G. Specific Ki-ras codon 61 mutations may determine the development of urethan-induced mouse lung adenomas or adenocarcinomas. Mol. Carcinog. 3, 287–295 (1990)
Berndt, A. et al. Identification of Fat4 and Tsc22d1 as novel candidate genes for spontaneous pulmonary adenomas. Cancer Res. 71, 5779–5791 (2011)
To, M. D., Rosario, R. D., Westcott, P. M., Banta, K. L. & Balmain, A. Interactions between wild-type and mutant Ras genes in lung and skin carcinogenesis. Oncogene 32, 4028–4033 (2013)
Johnson, L. et al. K-ras is an essential gene in the mouse with partial functional overlap with N-ras. Genes Dev. 11, 2468–2481 (1997)
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nature Biotechnol. 31, 213–219 (2013)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010)
Forbes, S. A. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945–D950 (2011)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012)
Boeva, V. et al. Control-free calling of copy number alterations in deep-sequencing data using GC-content normalization. Bioinformatics 27, 268–269 (2011)
Raponi, M. et al. Gene expression signatures for predicting prognosis of squamous cell and adenocarcinomas of the lung. Cancer Res. 66, 7466–7472 (2006)
Acknowledgements
This work was supported by National Cancer Institute (NCI) grants R01 CA111834, U01 CA84244, U01 CA141455 and UO1 CA176287 (to A.B.), and partly funded by the Bonnie Addario Foundation. P.M.K.W. was supported by the National Institutes of Health (NIH) training grant T32 GM007175 and a National Science Foundation GRFP award, and is currently supported by an NCI F31 NRSA award. K.D.H. was supported by the NIH training grant T32 GM007175, and is currently supported by an NCI F31 NRSA award. D.J.A. is supported by Cancer Research UK and the Wellcome Trust. We are appreciative of help and comments from our colleagues in refining this study and manuscript. We would also like to thank S. Busch for assistance with animal studies, and S. Green, T. Yuan and M. McMahon for providing the K493.1 cell line.
Author information
Authors and Affiliations
Contributions
P.M.K.W., K.D.H., M.D.T., D.J.A. and A.B. contributed to the overall study design. P.M.K.W. carried out most of the experiments, with help from M.D.T. R.D. was responsible for all of the animal studies. Sequencing and Sequenom were performed at the Sanger Institute under the supervision of D.J.A., and data processing was carried out by K.D.H., M.R., A.G.R. and T.M.K. SNV and CNA calling were carried out by K.D.H. Data analysis was carried out primarily by P.M.K.W. and K.D.H., with help from E.F. and D.A.Q. K.-Y.J. made histological assessments of all tumours. Adenomas and adenocarcinomas from the A/J mice were provided by C.J.K. and K.E.G. The manuscript was written primarily by P.M.K.W. and A.B., with contributions from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Distinct and consistent mutation spectra across tumours from carcinogen and genetic models.
a–c, Stacked heatmaps displaying the mutation spectra of all MNU-induced (a), urethane-induced (b), and KrasLA2 tumours (c), shown as normalized frequencies of all 96 possible substitutions. Substitutions are shown below each heatmap, with 5′- and 3′-flanking base context displayed on the top and right, respectively. Tumour identifier is shown to the left of each heatmap.
Extended Data Figure 2 Highly specific mutation signatures.
a, Breakdown of G > A transitions in MNU-induced tumours. 5′-flanking purine versus pyrimidine G > A substitutions, and 3′-flanking thymidine versus all other G > A substitutions, are highly significant (P < 0.0003, Wilcoxon rank-sum test). b, c, Breakdowns of A > G transitions (b) and A > T transversions (c) in urethane-induced tumours. d, e, All 96 substitutions in urethane-induced (d) and KrasLA2 tumours (e). e, The CGN > A (NCG > T) signature mutations of genomic instability are denoted. Mutation counts per tumour were normalized to total length of sequenced trinucleotide contexts in each tumour and averaged. Error bars represent s.e.m.
Extended Data Figure 3 Kras G12D mutation induces tumours with different histologies compared with codon 61 mutants.
a, Representative papillary, solid, and mixed tumour histologies (×200 magnification). b, Breakdown of different histologies in each treatment group. Histologies from KrasLA2 and MNU groups were significantly different compared with those from urethane, but there was no significant difference between KrasLA2 and MNU (Fisher exact test, Holm’s correction for multiple comparisons).
Extended Data Figure 4 Germline Kras genotype influences mutation specificity in urethane-induced tumours.
a, Kras mutant alleles for urethane tumours are plotted as coloured squares for all three oncogenic alleles detected in these tumours. Kras genotype is indicated as either white (wild type (WT)) or black (heterozygous) squares. b, Highly significant switch in Kras codon 61 mutations between tumours from wild-type mice and Kras+/− mice (Fisher exact test). c, No significant difference was seen between the exome-wide rates of the substitutions underlying Kras Q61R (CAA > G) and Q61L (CAA > T) mutations between tumours from wild-type and Kras+/− mice (Wilcoxon rank-sum test). NS, not significant.
Extended Data Figure 5 MTUS1 is a tumour suppressor in mouse and human lung cancer.
a, Polymerase chain reaction with quantitative reverse transcription (qRT–PCR) quantification of short interfering RNA (siRNA) knockdown of Mtus1 in a Kras G12D mouse lung cancer cell line (K493.1) (Wilcoxon rank-sum test). CTL, control. b, MTT assay shows increased growth after Mtus1 knockdown (Wilcoxon rank-sum test). Four independent trials were performed and growth was significantly increased by day 3 after knockdown in each experiment. One representative trial is shown. c, MTUS1 expression is significantly associated with overall survival in human lung adenocarcinoma (P = 0.00097, χ2 = 10.9). Analysis was performed using the clinical covariates gender, age, pack years smoked, and stage.
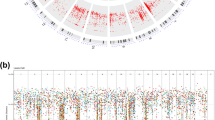
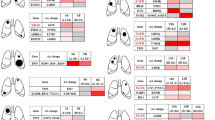
Extended Data Figure 6 Proportion of tumours with CNAs in each treatment group.
Amplifications and deletions were defined as regions with a log2 ratio greater than 0.5 or less than −0.5, respectively. Chromosomes are arranged on the x axis in a head-to-tail formation.
Extended Data Figure 7 Histological confirmation of lung adenocarcinomas.
a, b, Representative histologies (×400 magnification) of A/J (a) and FVB/N (b) adenocarcinomas. Zoom insets show tumour cell crowding and scattered mitotic figures (black arrowheads), nuclear atypia including enlargement and moderate pleomorphism, nuclear membrane irregularity, and prominent nucleoli. Scale bars, 20 μm.
Extended Data Figure 8 Comparison of urethane signature mutations in adenomas and adenocarcinomas.
Urethane A > G transitions (left) and A > T transversions (right) are shown in A/J adenocarcinomas, FVB/N adenocarcinomas and FVB/N adenomas. Mutation counts per tumour were normalized to total length of sequenced trinucleotide contexts in each tumour and averaged. Error bars represent s.e.m.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 and 3-8. (PDF 321 kb)
Supplementary Data
This file contains Supplementary Table 2. (XLSX 17 kb)
Supplementary Data
VCF file of all SNVs called in the 82 lung adenomas. (TXT 5724 kb)
Supplementary Data
VCF file of all SNVs called in the 22 lung adenocarcinomas. (TXT 658 kb)
Supplementary Data
Sample to ID key file. (TXT 8 kb)
Rights and permissions
About this article
Cite this article
Westcott, P., Halliwill, K., To, M. et al. The mutational landscapes of genetic and chemical models of Kras-driven lung cancer. Nature 517, 489–492 (2015). https://doi.org/10.1038/nature13898
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13898
This article is cited by
-
Inhibition of ADAM9 promotes the selective degradation of KRAS and sensitizes pancreatic cancers to chemotherapy
Nature Cancer (2024)
-
KRAS allelic imbalance drives tumour initiation yet suppresses metastasis in colorectal cancer in vivo
Nature Communications (2024)
-
The role of APOBEC3B in lung tumor evolution and targeted cancer therapy resistance
Nature Genetics (2024)
-
Tumor heterogeneity impairs immunogenicity in mismatch repair deficient tumors
Nature Genetics (2023)
-
Micro-CT acquisition and image processing to track and characterize pulmonary nodules in mice
Nature Protocols (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.