Abstract
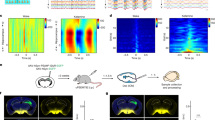
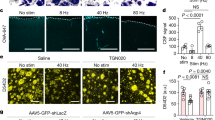
The mammalian taste system is responsible for sensing and responding to the five basic taste qualities: sweet, sour, bitter, salty and umami. Previously, we showed that each taste is detected by dedicated taste receptor cells (TRCs) on the tongue and palate epithelium1. To understand how TRCs transmit information to higher neural centres, we examined the tuning properties of large ensembles of neurons in the first neural station of the gustatory system. Here, we generated and characterized a collection of transgenic mice expressing a genetically encoded calcium indicator2 in central and peripheral neurons, and used a gradient refractive index microendoscope3 combined with high-resolution two-photon microscopy to image taste responses from ganglion neurons buried deep at the base of the brain. Our results reveal fine selectivity in the taste preference of ganglion neurons; demonstrate a strong match between TRCs in the tongue and the principal neural afferents relaying taste information to the brain; and expose the highly specific transfer of taste information between taste cells and the central nervous system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yarmolinsky, D. A., Zuker, C. S. & Ryba, N. J. P. Common sense about taste: from mammals to insects. Cell 139, 234–244 (2009)
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nature Methods 6, 875–881 (2009)
Jung, J. C., Mehta, A. D., Aksay, E., Stepnoski, R. & Schnitzer, M. J. In vivo mammalian brain imaging using one- and two-photon fluorescence microendoscopy. J. Neurophysiol. 92, 3121–3133 (2004)
Simon, S. A., de Araujo, I. E., Gutierrez, R. & Nicolelis, M. A. The neural mechanisms of gustation: a distributed processing code. Nature Rev. Neurosci. 7, 890–901 (2006)
Chandrashekar, J. et al. The cells and peripheral representation of sodium taste in mice. Nature 464, 297–301 (2010)
Huang, A. L. et al. The cells and logic for mammalian sour taste detection. Nature 442, 934–938 (2006)
Mueller, K. L. et al. The receptors and coding logic for bitter taste. Nature 434, 225–229 (2005)
Zhao, G. Q. et al. The receptors for mammalian sweet and umami taste. Cell 115, 255–266 (2003)
Chaudhari, N. & Roper, S. D. The cell biology of taste. J. Cell Biol. 190, 285–296 (2010)
Frank, M. & Pfaffmann, C. Taste nerve fibers: a random distribution of sensitivities to four tastes. Science 164, 1183–1185 (1969)
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000)
Akerboom, J. et al. Optimization of a GCaMP calcium indicator for neural activity imaging. J. Neurosci. 32, 13819–13840 (2012)
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013)
Li, X. et al. Human receptors for sweet and umami taste. Proc. Natl Acad. Sci. USA 99, 4692–4696 (2002)
Nelson, G. et al. An amino-acid taste receptor. Nature 416, 199–202 (2002)
Adler, E. et al. A novel family of mammalian taste receptors. Cell 100, 693–702 (2000)
Oka, Y., Butnaru, M., von Buchholtz, L., Ryba, N. J. P. & Zuker, C. S. High salt recruits aversive taste pathways. Nature 494, 472–475 (2013)
Tomchik, S. M., Berg, S., Kim, J. W., Chaudhari, N. & Roper, S. D. Breadth of tuning and taste coding in mammalian taste buds. J. Neurosci. 27, 10840–10848 (2007)
Ghanouni, P. et al. The effect of pH on β2 adrenoceptor function. Evidence for protonation-dependent activation. J. Biol. Chem. 275, 3121–3127 (2000)
Heyer, B. R., Taylor-Burds, C. C., Tran, L. H. & Delay, E. R. Monosodium glutamate and sweet taste: generalization of conditioned taste aversion between glutamate and sweet stimuli in rats. Chem. Senses 28, 631–641 (2003)
Chen, X., Gabitto, M., Peng, Y., Ryba, N. J. P. & Zuker, C. S. A gustotopic map of taste qualities in the mammalian brain. Science 333, 1262–1266 (2011)
Beidler, L. M. & Smallman, R. L. Renewal of cells within taste buds. J. Cell Biol. 27, 263–272 (1965)
Grill, H. J. & Norgren, R. The taste reactivity test. II. Mimetic responses to gustatory stimuli in chronic thalamic and chronic decerebrate rats. Brain Res. 143, 281–297 (1978)
Zaidi, F. N. & Whitehead, M. C. Discrete innervation of murine taste buds by peripheral taste neurons. J. Neurosci. 26, 8243–8253 (2006)
Barretto, R. P. et al. Time-lapse imaging of disease progression in deep brain areas using fluorescence microendoscopy. Nature Med. 17, 223–228 (2011)
Sollars, S. I. & Hill, D. L. In vivo recordings from rat geniculate ganglia: taste response properties of individual greater superficial petrosal and chorda tympani neurones. J. Physiol. (Lond.) 564, 877–893 (2005)
Thévenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998)
Mukamel, E. A., Nimmerjahn, A. & Schnitzer, M. J. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63, 747–760 (2009)
Rousseeuw, P. J. & Croux, C. Alternatives to the median absolute deviation. J. Am. Stat. Assoc. 88, 1273–1283 (1993)
Hyman, A. M. & Frank, M. E. Effects of binary taste stimuli on the neural activity of the hamster chorda tympani. J. Gen. Physiol. 76, 125–142 (1980)
Acknowledgements
We thank the National Institute of Dental and Craniofacial Research (NIDCR) transgenic-core and C. Guo at Janelia Farms for help in generating the Thy1-GCaMP3 mouse lines, B. Shields for histology support, and Y. Oka and M. Butnaru for nerve recording and pharmacological advice. We also thank members of the Zuker laboratory for helpful comments. This research was supported in part by the intramural research program of NIDCR (N.J.P.R.). C.S.Z. is an investigator of the Howard Hughes Medical Institute and a Senior Fellow at Janelia Farms.
Author information
Authors and Affiliations
Contributions
R.P.J.B. designed the study, carried out the imaging experiments, analysed data and wrote the paper; S.G.-S. developed viral gene delivery to ganglion neurons and characterized the transgenic lines; J.C. characterized the transgenic lines, carried out initial imaging experiments and analysed data; D.A.Y. collected and analysed data; M.J.S. provided microendoscopy expertise; N.J.P.R. and C.S.Z. designed the study, analysed data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Thy1–GCaMP3 mice show normal physiological responses to tastants.
a, Representative nerve recording traces from control and Thy1-GCaMP3 mice in response to various tastants (see Methods for details). b, Quantification of neural responses (mean + s.e.m.) show that Thy1-GCaMP3 mice (n = 4) are indistinguishable from wild-type mice (n = 3; Student’s t-test; NaCl, P = 0.85; bitter, P = 0.46; sour, P = 0.94; sweet, P = 0.69; umami, P = 0.77). Recordings were normalized to responses to KCl (500 mM). Horizontal bars below the traces mark the time and duration of the stimulus.
Extended Data Figure 2 Reproducibility of tastant-evoked responses in geniculate ganglion neurons.
a, Representative images of calcium-evoked GCaMP3 activity in response to sweet (left) and bitter (right) stimulation. Four relative fluorescence images are shown from separate trials. In each trial, the identical cell populations were activated. b, We tested 105 sweet responding cells and 168 bitter responding cells for their reproducibility in our automated scoring algorithm for four trials. The histograms show the number of times the cells respond all four times, three out of four, two out of four, and one out of four. c, Sample traces of four representative neurons challenged with 50 trials of the same tastant over a time window of 10 min. Note the high reliability in the activation of the neurons. This experiment also illustrates the desensitization of bitter neurons (bottom traces) over time. Horizontal bars below the traces mark the time and duration of the stimulus.
Extended Data Figure 3 Quantification of taste ganglion responses.
a–d, Rank-ordered plot of calcium transient amplitudes for various singly tuned ganglion neurons (see text and Fig. 3). For each cell, the mean response amplitudes for preferred stimulus (red) and the mean amplitude of its next-strongest tastant response (grey) are shown; minor dots indicate individual trial amplitudes. e, Quantification of mean response amplitudes in singly tuned salt ganglion neurons before and after amiloride treatment (10 μM, n = 23 cells; paired t-test, P < 0.001). f, Quantification of mean response amplitudes in singly tuned bitter cells before and after AITC treatment17 (3 mM, n = 63 cells; paired t-test, P < 0.001).
Extended Data Figure 4 Bitter–sour ganglion cells receive taste information from bitter T2R-expressing cells.
Distribution of bitter, sour, and bitter–sour ganglion cells in a sample of control animals (n = 4) and in animals expressing tetanus toxin in PKD2L1-expressing TRCs (PKD2L1-TeNT; n = 3). As expected, no cells responsive to citric acid (50 mM) are detected in PKD2L1-TeNT mice17. However, bitter–sour cells are unaffected (see Fig. 4), suggesting that activation of T2R-expressing TRCs mediates these acidic responses. As predicted, subsequent application of the bitter TRC inactivator AITC17 abolishes bitter responses of the bitter ganglion neurons, as well as the bitter and sour responses in the bitter–sour cells. Note that the solid bars showing less than 1 cell are used to illustrate the lack of responding cells.
Extended Data Figure 5 Representation of taste quality does not cluster within the geniculate ganglion.
a, Two-photon endoscopic image (left) of a geniculate ganglion expressing GCaMP3. Highlighted are the locations of the facial (VII) and greater superficial petrosal (GSP) cranial nerves. The right panel shows approximately 50 neurons colour-coded according to their taste preference in this field. Sour, yellow; sweet, magenta; NaCl, green; bitter, cyan. Scale bar, 200 μm. b, Representative fields of the geniculate ganglion from 6 different mice. The random distribution of neurons representing the various taste qualities is independent of sensor, or method of sensor delivery/expression (AAV-GCaMP6 or Thy1-GCaMP3); colour scheme same as for a.
Extended Data Figure 6 Representation of taste mixtures.
a, Imaging fields of three representative geniculate ganglia illustrating the ensembles of neurons recruited by two different single taste stimuli presented separately (left panels) versus the ensemble of neurons activated by a mixture of the two compounds presented together (right panels). See text for details; as expected there are no mixture-specific responders, and very few cells responded to each tastant in the mix: only 3 out of 113 cells examined with bitter + sweet responded to both tastants, 5 out of 301 cells examined with sour + salty responded to both, and 0 of 39 examined with salty + sweet responded to both tastants. We note that sour stimuli are known to suppress sweet responses30, but such suppression is sweet-cell autonomous and not due to interactions between sweet and sour TRCs (data not shown). b, To quantitatively examine the impact of taste mixes on the responses of individual ganglion neurons, we analysed their response amplitudes in the presence of the single tastant versus the binary mix. Shown are plots of response amplitudes of a representative set of bitter, sweet, salty and sour geniculate neurons stimulated with their selective tastant (x axis) versus their response amplitude when in the presence of an additional tastant (as indicated in the y axis; shown are average ΔF/F over 4 trials). 95% confidence intervals were determined using a ratio t-test: bitter + sweet/bitter, 0.73–0.91; sweet + bitter/sweet, 1.15–1.34; NaCl + sour/NaCl 0.74–1.00; sour + NaCl/sour, 0.95–1.16.
Rights and permissions
About this article
Cite this article
Barretto, R., Gillis-Smith, S., Chandrashekar, J. et al. The neural representation of taste quality at the periphery. Nature 517, 373–376 (2015). https://doi.org/10.1038/nature13873
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13873
This article is cited by
-
Experience-dependent changes in affective valence of taste in male mice
Molecular Brain (2023)
-
A chemo-mechanical cochleostomy preserves hearing for the in vivo functional imaging of cochlear cells
Nature Protocols (2023)
-
Dual-sensory fusion self-powered triboelectric taste-sensing system towards effective and low-cost liquid identification
Nature Food (2023)
-
A bottom-up reward pathway mediated by somatostatin neurons in the medial septum complex underlying appetitive learning
Nature Communications (2022)
-
Gut–brain circuits for fat preference
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.