Abstract
In BRAF(V600)-mutant tumours, most mechanisms of resistance to drugs that target the BRAF and/or MEK kinases rely on reactivation of the RAS–RAF–MEK–ERK mitogen-activated protein kinase (MAPK) signal transduction pathway, on activation of the alternative, PI(3)K–AKT–mTOR, pathway (which is ERK independent) or on modulation of the caspase-dependent apoptotic cascade1,2,3. All three pathways converge to regulate the formation of the eIF4F eukaryotic translation initiation complex, which binds to the 7-methylguanylate cap (m7G) at the 5′ end of messenger RNA, thereby modulating the translation of specific mRNAs4,5. Here we show that the persistent formation of the eIF4F complex, comprising the eIF4E cap-binding protein, the eIF4G scaffolding protein and the eIF4A RNA helicase, is associated with resistance to anti-BRAF, anti-MEK and anti-BRAF plus anti-MEK drug combinations in BRAF(V600)-mutant melanoma, colon and thyroid cancer cell lines. Resistance to treatment and maintenance of eIF4F complex formation is associated with one of three mechanisms: reactivation of MAPK signalling, persistent ERK-independent phosphorylation of the inhibitory eIF4E-binding protein 4EBP1 or increased pro-apoptotic BCL-2-modifying factor (BMF)-dependent degradation of eIF4G. The development of an in situ method to detect the eIF4E–eIF4G interactions shows that eIF4F complex formation is decreased in tumours that respond to anti-BRAF therapy and increased in resistant metastases compared to tumours before treatment. Strikingly, inhibiting the eIF4F complex, either by blocking the eIF4E–eIF4G interaction or by targeting eIF4A, synergizes with inhibiting BRAF(V600) to kill the cancer cells. eIF4F not only appears to be an indicator of both innate and acquired resistance but also is a promising therapeutic target. Combinations of drugs targeting BRAF (and/or MEK) and eIF4F may overcome most of the resistance mechanisms arising in BRAF(V600)-mutant cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Data deposits
Microarray data have been deposited in the ArrayExpress database under accession number E-MTAB-2607.
References
Lito, P., Rosen, N. & Solit, D. B. Tumor adaptation and resistance to RAF inhibitors. Nature Med. 19, 1401–1409 (2013)
Shi, H. et al. Acquired resistance and clonal evolution in melanoma during BRAF inhibitor therapy. Cancer Discov. 4, 80 (2014)
Tentori, L., Lacal, P. M. & Graziani, G. Challenging resistance mechanisms to therapies for metastatic melanoma. Trends Pharmacol. Sci. 34, 656–666 (2013)
Blagden, S. P. & Willis, A. E. The biological and therapeutic relevance of mRNA translation in cancer. Nature Rev. Clin. Oncol. 8, 280–291 (2011)
Silvera, D., Formenti, S. C. & Schneider, R. J. Translational control in cancer. Nature Rev. Cancer 10, 254–266 (2010)
Shao, Y. & Aplin, A. E. BH3-only protein silencing contributes to acquired resistance to PLX4720 in human melanoma. Cell Death Differ. 19, 2029–2039 (2012)
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012)
Cencic, R., Galicia-Vazquez, G. & Pelletier, J. Inhibitors of translation targeting eukaryotic translation initiation factor 4A. Methods Enzymol. 511, 437–461 (2012)
Oikonomou, E., Koc, M., Sourkova, V., Andera, L. & Pintzas, A. Selective BRAFV600E inhibitor PLX4720, requires TRAIL assistance to overcome oncogenic PIK3CA resistance. PLoS ONE 6, e21632 (2011)
Prahallad, A. et al. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature 483, 100–103 (2012)
Montero-Conde, C. et al. Relief of feedback inhibition of HER3 transcription by RAF and MEK inhibitors attenuates their antitumor effects in BRAF-mutant thyroid carcinomas. Cancer Discov. 3, 520–533 (2013)
Packer, L. M., East, P., Reis-Filho, J. S. & Marais, R. Identification of direct transcriptional targets of V600EBRAF/MEK signalling in melanoma. Pigment Cell Melanoma Res. 22, 785–798 (2009)
Pratilas, C. A. et al. V600EBRAF is associated with disabled feedback inhibition of RAF-MEK signaling and elevated transcriptional output of the pathway. Proc. Natl Acad. Sci. USA 106, 4519–4524 (2009)
Moerke, N. J. et al. Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G. Cell 128, 257–267 (2007)
Bordeleau, M. E. et al. Therapeutic suppression of translation initiation modulates chemosensitivity in a mouse lymphoma model. J. Clin. Invest. 118, 2651–2660 (2008)
Cencic, R. et al. Synergistic effect of inhibiting translation initiation in combination with cytotoxic agents in acute myelogenous leukemia cells. Leuk. Res. 34, 535–541 (2010)
Sadlish, H. et al. Evidence for a functionally relevant rocaglamide binding site on the eIF4A–RNA complex. ACS Chem. Biol. 8, 1519–1527 (2013)
Gupta, S. V. et al. Resistance to the translation initiation inhibitor silvestrol is mediated by ABCB1/P-glycoprotein overexpression in acute lymphoblastic leukemia cells. AAPS J. 13, 357–364 (2011)
Liu, T. et al. Synthetic silvestrol analogues as potent and selective protein synthesis inhibitors. J. Med. Chem. 55, 8859–8878 (2012)
Thuaud, F. et al. Synthetic analogue of rocaglaol displays a potent and selective cytotoxicity in cancer cells: involvement of apoptosis inducing factor and caspase-12. J. Med. Chem. 52, 5176–5187 (2009)
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nature Protocols 7, 1534–1550 (2012)
Acknowledgements
The authors thank J. Tanaka for providing hippuristanol. We thank the following Gustave Roussy platforms: Imaging and Cytometry Platform IRCIV (S. Salome-Desmoulez), Module de Développement en Pathologie, SIRIC SOCRATE (J. Adam), Translational Research Laboratory and Biobank (M. Breckler and L. Lacroix), Plateforme d’évaluation Préclinique (P. Gonin and K. Ser-le Roux), Genomic Core Facility (N. Pata-Merci) and Bioinformatic Core Facility (G. Meurice). We also thank V. Camara-Clayette for help with 35S experiments, S. Roy for patient data collection and L. Saint Ange for text editing. C.R. and S.V.’s team was supported by Institut National du CAncer (INCA), Association pour la Recherche sur le Cancer (ARC) and Ligue contre le Cancer via an Integrated Research Action Program Melanoma (PAIR Melanome), Cancéropôle Ile de France and Ensemble Contre le Mélanome. L.D. was supported by the Association pour la Recherche sur le Cancer (ARC). We also thank the ARC and AAREC Filia Research for fellowships to N.R. and C.B. O.H. was supported by the Wenner-Gren Foundation and the Swedish Society of Medicine.
Author information
Authors and Affiliations
Contributions
L.B., H.M.-M., I.G., D.A., O.H., G.T., C.B., N.R., F.T., N.K.-K., S.A. and L.D. designed and performed experiments and analysed the data. C.M., E.R., M.T. and A.M.E. provided clinical samples and gave advice. S.V. and C.R. supervised all research, wrote the manuscript and are joint senior authors.
Corresponding authors
Ethics declarations
Competing interests
C.R. is a consultant to GlaxoSmithKline, Roche, Bristol-Myers Squibb and Merck.
Extended data figures and tables
Extended Data Figure 1 Sensitivity of melanoma cell lines to anti-BRAF, anti-MEK or anti-MNK inhibitors.
a, Short-term growth-inhibition assay of the indicated cell lines (SK-MEL-28, A375, Mel888, SK-MEL-5, A2058, Mel624 and Mel10) treated with increasing concentrations of vemurafenib, dabrafenib, PD0325901, trametinib and CGP 57380 for 48 h. The Mel10 cell line does not have a mutated BRAF gene and was used as a control vemurafenib-insensitive cell line. Cell viability was determined using the WST-1 cell proliferation assay. The data are presented as the mean ± s.d. (n = 3). b, c, Long-term colony formation assay of the indicated cell lines. Cells were grown in the absence or presence of vemurafenib (b) or dabrafenib and trametinib (c) at the indicated concentrations for 2 weeks. For each cell line, all dishes were fixed at the same time, stained and photographed.
Extended Data Figure 2 Xenograft study.
a, Growth of A375 and Mel624 cells as tumour xenografts in nude mice treated with vehicle or increasing concentrations of PLX4720 (Plx). Mean tumour volumes + s.e.m. are shown (vehicle and 200 mg Plx per kg body weight groups of A375- and Mel624-xenografted mice comprised 4 mice; 90 mg and 417 mg Plx per kg body weight groups of A375-xenografted mice comprised 3 mice; 90 mg and 417 mg Plx per kg body weight groups of Mel624-xenografted mice comprised 5 mice). Significance was determined by one-sided Mann–Whitney U test (*, P < 0.05; **P < 0.01). b, Interactions between eIF4E and eIF4G (eIF4E–eIF4G) or eIF4E and 4EBP1 (eIF4E–4EBP1) detected by in situ proximity ligation assay (PLA) in paraffin-embedded tissue sections of A375 and Mel624 xenografts from Plx-treated mice. The interactions were visualized as brown spots. c, Quantification of the PLA showing the ratio between the number of purple spots corresponding to the eIF4E–eIF4G interaction per 100 nuclei and the number of purple spots corresponding to the eIF4E–4EBP1 interaction per 100 nuclei, normalized to the same ratio before treatment. Error bars, s.d. d, Growth of Mel624 cells as tumour xenografts in nude mice treated with vehicle, a low dose of Plx alone, FL3 alone or a combination of Plx and FL3. FL3 treatment was stopped after 13 days. The data are presented as the mean + s.e.m. (Vehicle, 15 mg FL3 per kg body weight, and 90 mg Plx per kg body weight plus 15 mg FL3 per kg body weight groups comprised 6 mice; 90 mg Plx per kg body weight group comprised 3 mice.) Significance was determined by one-sided Mann–Whitney U test (*, P < 0.05).
Extended Data Figure 3 Protein synthesis rates and polysome profiles.
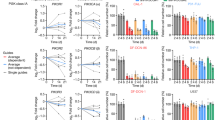
a, Protein synthesis rates were determined in A375 cells treated for the indicated times (3 or 24 h) with vehicle (dimethylsulphoxide (DMSO)) or 0.7 or 6 µM vemurafenib or for 15 min with 10 µM cycloheximide. Cells were then pulsed for 30 min with [35S]Cys/Met, and the incorporation of 35S into proteins was quantified and normalized to the total protein amount. The data are presented as the mean (n = 2). b, Polysome profiles of A375 and Mel624 cells treated with DMSO or 6 µM vemurafenib for 3 h. The area under the curve was measured with ImageJ software. OD, optical density; RNP, ribonucleoparticle. c, Volcano plot showing a subset of 251 mRNAs that were differentially translated between vemurafenib-treated and untreated A375 cells. Of these 251 mRNAs, 73 were overexpressed (red), and 178 were underexpressed (green). d, A table showing some of the genes that encode mRNAs whose translation is inhibited by vemurafenib in A375 cells (see also Supplementary Table 2). These mRNAs belong to the TOP mRNA class, the members of which contain a TOP motif and/or encode translation factors and ribosomal proteins.
Extended Data Figure 4 Translation of selected mRNAs in vemurafenib-treated melanoma cell lines.
The abundance of EEF1G, HNRNPA1, EIF3L, FOSB, TBP (control) and HPRT (control) transcripts in fractions from Extended Data Fig. 3b were quantified by quantitative reverse transcription PCR (qRT–PCR). The percentage of each mRNA in each fraction was calculated.
Extended Data Figure 5 Formation of the eIF4F complex using a PLA procedure.
a, The specificity of the PLA was evaluated by omitting one of the two primary antibodies or the two primary antibodies and by omitting the minus and plus PLA probes. b, The interactions between eIF4E and eIF4G (eIF4E–eIF4G) and eIF4E and 4EBP1 (eIF4E–4EBP1) detected by in situ PLA in the indicated cell lines treated for 24 h with the small molecule inhibitor 4EGI-1, which disrupts eIF4E–eIF4G interaction14. The interactions were visualized as red fluorescent spots. Cell nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI) (blue). c, The interactions between eIF4E and eIF4G (eIF4E–eIF4G) or eIF4E and 4EBP1 (eIF4E–4EBP1) detected by in situ PLA in siRNA-mediated eIF4E-depleted A375 cells. The interactions were visualized as red fluorescent spots. The cell nuclei were stained with DAPI (blue). d, Western blot analysis with antibodies specific for eIF4E and β-actin, to evaluate siRNA-mediated depletion of eIF4E.
Extended Data Figure 6 Formation of the eIF4F complex in vemurafenib-treated melanoma cell lines and analysis of eIF4G cleavage.
a, Quantification of the PLA showing the number of red spots (corresponding to eIF4E–eIF4G or eIF4E–4EBP1 interactions) per 100 nuclei (quantified with Volocity software). The data are presented as the mean ± s.d. (n = 3). Significance was determined by Mann–Whitney U test (*, P < 0.05; **, P < 0.01). b, The interactions between eIF4E and eIF4G (eIF4E–eIF4G) or eIF4E and 4EBP1 (eIF4E–4EBP1) detected by in situ PLA in A375 cells treated for 24 h with vemurafenib at the indicated concentrations. The interactions were visualized as red fluorescent spots. The cells were counterstained with DAPI (blue). White numbers correspond to means, and error bars are derived from three replicates. c, Time course of the interactions between eIF4E and eIF4G (eIF4E–eIF4G) detected by in situ PLA in response to the treatment of A375 cells with 6 µM vemurafenib. The interactions were visualized as red fluorescent spots at the indicated times. d, A375 cells were treated (or not treated) with 6 µM vemurafenib and collected at the indicated times for western blot analysis with antibodies specific for eIF4G, eIF4E and GAPDH. The cleaved eIF4G:full-length eIF4G density ratio and the eIF4E:GAPDH density ratio (indicated in blue for each time point) were quantified by densitometric scanning of the blots followed by ImageJ analysis.
Extended Data Figure 7 Formation of the eIF4F complex in anti-BRAF and anti-MEK treated melanoma cell lines and analysis of the synergistic effect of FL3 and anti-MEK compounds.
a, The interactions between eIF4E and eIF4G (eIF4E–eIF4G) detected by in situ PLA in A375 and A2058 cell lines treated for 24 h with dabrafenib, trametinib or a combination of dabrafenib and trametinib. The interactions were visualized as red fluorescent spots with confocal microscopy (Leica SPE). Cell nuclei were stained with DAPI (blue). b, Quantification of the PLA results showing the number of red spots (corresponding to the eIF4E–eIF4G or eIF4E–4EBP1 interactions) per 100 nuclei calculated with Volocity software. The data are presented as the mean ± s.d. (n = 3). Significance was determined by the Mann–Whitney U test (*, P < 0.05; **, P < 0.01). c, Isobolograms showing the correlation between the observed and expected effects of the combination of MEK inhibitors (such as PD0325901 or trametinib) and FL3 on the Mel624 cell line. The upper left region of the figure represents increasing degrees of synergy. The minimum and maximum Bliss index values are shown as an interval within square brackets under each isobologram. The Bliss index was calculated as the ratio observed:expected, where 1, <0 or >1 indicate additive, antagonistic or synergistic effects, respectively.
Extended Data Figure 8 Formation of the eIF4F complex in various vemurafenib-treated cell lines.
a, The eIF4E–eIF4G and eIF4E–4EBP1 interactions detected by in situ PLA in the Malme-3M melanoma cell line and its vemurafenib-resistant counterpart (R-Malme-3M) after treatment with 6 μM vemurafenib (24 h) or no treatment. The interactions were visualized as red fluorescent spots. Cell nuclei were stained with DAPI (blue). White numbers correspond to the mean ± s.d. derived from three replicates quantified with ImageJ. b, The eIF4E–eIF4G and eIF4E–4EBP1 interactions detected by in situ PLA in the HT-29 colon cancer cell line and the BCPAP thyroid cancer cell line treated with 6 μM vemurafenib (24 h) or untreated. The interactions were visualized as red fluorescent spots. Cell nuclei were stained with DAPI (blue).
Extended Data Figure 9 Involvement of BMF in relative resistance to vemurafenib and analysis of rapamycin effects on AKT, S6 and 4EBP1 phosphorylation.
a, Relative proliferation of A375 cells after siRNA-mediated depletion of 27 mRNAs associated with the result in Fig. 2d. The data are presented as the mean ± s.d. (n = 3). Significance was determined by Student’s t- test (*, P < 0.05). b, Relative expression of BMF mRNA (as determined by qRT–PCR) after siRNA-mediated depletion of BMF (si-BMF) in A375 cells. Data are presented as the mean ± s.d. (n = 3). Significance was determined by Student’s t-test (**, P < 0.01). c, Western blot analysis with antibodies specific for phosphorylated (P-) and/or total AKT, S6, 4EBP1 and ERK1/2 (ERK) on cell extracts of the MDA-MB-468 breast cancer and Mel624 melanoma cell lines treated for 24 h with the indicated concentrations of rapamycin.
Extended Data Figure 10 Formation of the eIF4F complex in each of the tumour samples from seven patients.
The eIF4E–eIF4G and eIF4E–4EBP1 interactions detected by in situ PLA in tumour samples collected from each patient (patient 1 to 7), before and during vemurafenib or dabrafenib treatment. Each brown spot represents an interaction.
Supplementary information
Supplementary Information
This file contains Supplementary Text and the Chemical formula of flavaglines used in this study. (PDF 264 kb)
Supplementary Table 1
Determination of half-maximum inhibitory concentration (IC50) for the indicated compounds and mutational screening in each of the melanoma cell lines used in this study. (XLS 38 kb)
Supplementary Table 2
List of genes encoding mRNAs that are translationnally regulated by vemurafenib. (XLSX 81 kb)
Supplementary Table 3
Lists of genes transcriptionally regulated by vemurafenib with a >2 fold change (FC) in either A375 (tab 1) or Mel624 (tab 2). (XLSX 29 kb)
Supplementary Table 4
Patient profile table. (XLS 23 kb)
Supplementary Table 5
Mutational screening in the tumours of the 7 patients. (XLS 29 kb)
Supplementary Table 6
List of ON-TARGETplus siRNA sequences provided from ThermoScientific used in Extended Data Fig. 8. (XLS 37 kb)
Supplementary Table 7
Oligonucleotides used for qPCR experiments. (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Boussemart, L., Malka-Mahieu, H., Girault, I. et al. eIF4F is a nexus of resistance to anti-BRAF and anti-MEK cancer therapies. Nature 513, 105–109 (2014). https://doi.org/10.1038/nature13572
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13572
This article is cited by
-
BRAF — a tumour-agnostic drug target with lineage-specific dependencies
Nature Reviews Clinical Oncology (2024)
-
Candidate biomarkers for treatment benefit from sunitinib in patients with advanced renal cell carcinoma using mass spectrometry-based (phospho)proteomics
Clinical Proteomics (2023)
-
Engineering an autonomous VH domain to modulate intracellular pathways and to interrogate the eIF4F complex
Nature Communications (2022)
-
Targeted intervention of eIF4A1 inhibits EMT and metastasis of pancreatic cancer cells via c-MYC/miR-9 signaling
Cancer Cell International (2021)
-
The plasticity of mRNA translation during cancer progression and therapy resistance
Nature Reviews Cancer (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.