Abstract
The mir-34/449 family consists of six homologous miRNAs at three genomic loci. Redundancy of miR-34/449 miRNAs and their dominant expression in multiciliated epithelia suggest a functional significance in ciliogenesis. Here we report that mice deficient for all miR-34/449 miRNAs exhibited postnatal mortality, infertility and strong respiratory dysfunction caused by defective mucociliary clearance. In both mouse and Xenopus, miR-34/449-deficient multiciliated cells (MCCs) exhibited a significant decrease in cilia length and number, due to defective basal body maturation and apical docking. The effect of miR-34/449 on ciliogenesis was mediated, at least in part, by post-transcriptional repression of Cp110, a centriolar protein suppressing cilia assembly. Consistent with this, cp110 knockdown in miR-34/449-deficient MCCs restored ciliogenesis by rescuing basal body maturation and docking. Altogether, our findings elucidate conserved cellular and molecular mechanisms through which miR-34/449 regulate motile ciliogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355 (2004)
He, L. & Hannon, G. J. MicroRNAs: small RNAs with a big role in gene regulation. Nature Rev. Genet. 5, 522–531 (2004)
Kim, V. N. Small RNAs: classification, biogenesis, and function. Mol. Cell 19, 1–15 (2005)
Du, T. & Zamore, P. D. microPrimer: the biogenesis and function of microRNA. Development 132, 4645–4652 (2005)
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993)
Wightman, B., Ha, I. & Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855–862 (1993)
Miska, E. A. et al. Most Caenorhabditis elegans microRNAs are individually not essential for development or viability. PLoS Genet. 3, e215 (2007)
Park, C. Y. et al. A resource for the conditional ablation of microRNAs in the mouse. Cell Rep. 1, 385–391 (2012)
Marson, A. et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134, 521–533 (2008)
Marcet, B. et al. Control of vertebrate multiciliogenesis by miR-449 through direct repression of the Delta/Notch pathway. Nature Cell Biol. 13, 693–699 (2011)
He, L. et al. A microRNA component of the p53 tumour suppressor network. Nature 447, 1130–1134 (2007)
Chang, T. C. et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol. Cell 26, 745–752 (2007)
Raver-Shapira, N. et al. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol. Cell 26, 731–743 (2007)
He, L., He, X., Lowe, S. W. & Hannon, G. J. microRNAs join the p53 network–another piece in the tumour-suppression puzzle. Nature Rev. Cancer 7, 819–822 (2007)
Hermeking, H. p53 enters the microRNA world. Cancer Cell 12, 414–418 (2007)
Satir, P. & Christensen, S. T. Overview of structure and function of mammalian cilia. Annu. Rev. Physiol. 69, 377–400 (2007)
Fliegauf, M., Benzing, T. & Omran, H. When cilia go bad: cilia defects and ciliopathies. Nature Rev. Mol. Cell Biol. 8, 880–893 (2007)
Spektor, A., Tsang, W. Y., Khoo, D. & Dynlacht, B. D. Cep97 and CP110 suppress a cilia assembly program. Cell 130, 678–690 (2007)
Tsang, W. Y. & Dynlacht, B. D. CP110 and its network of partners coordinately regulate cilia assembly. Cilia 2, 9 (2013)
Choi, Y. J. et al. miR-34 miRNAs provide a barrier for somatic cell reprogramming. Nature Cell Biol. 13, 1353–1360 (2011)
Lizé, M., Klimke, A. & Dobbelstein, M. MicroRNA-449 in cell fate determination. Cell Cycle 10, 2874–2882 (2011)
Loges, N. T. et al. DNAI2 mutations cause primary ciliary dyskinesia with defects in the outer dynein arm. Am. J. Hum. Genet. 83, 547–558 (2008)
Castleman, V. H. et al. Mutations in radial spoke head protein genes RSPH9 and RSPH4A cause primary ciliary dyskinesia with central-microtubular-pair abnormalities. Am. J. Hum. Genet. 84, 197–209 (2009)
Stubbs, J. L., Oishi, I., Izpisúa Belmonte, J. C., Kintner, C. & Izpisu, J. C. The forkhead protein Foxj1 specifies node-like cilia in Xenopus and zebrafish embryos. Nature Genet. 40, 1454–1460 (2008)
Marshall, W. F. Basal bodies: platforms for building cilia. Curr. Top. Dev. Biol. 85, 1–22 (2008)
Avasthi, P. & Marshall, W. F. Stages of ciliogenesis and regulation of ciliary length. Differentiation 83, S30–S42 (2012)
Werner, M. E. & Mitchell, B. J. Understanding ciliated epithelia: the power of Xenopus. Genesis 50, 176–185 (2012)
Gomperts, B. N., Gong-Cooper, X. & Hackett, B. P. Foxj1 regulates basal body anchoring to the cytoskeleton of ciliated pulmonary epithelial cells. J. Cell Sci. 117, 1329–1337 (2004)
Klos Dehring, D. A. et al. Deuterosome-mediated centriole biogenesis. Dev. Cell 27, 103–112 (2013)
Martinez-Anton, A. et al. Changes in microRNA and mRNA expression with differentiation of human bronchial epithelial cells. Am. J. Respir. Cell Mol. Biol. 49, 384–395 (2013)
Lewis, B. P., Burge, C. B. & Bartel, D. P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005)
Miranda, K. C. et al. A pattern-based method for the identification of microRNA binding sites and their corresponding heteroduplexes. Cell 126, 1203–1217 (2006)
Hoh, R. A., Stowe, T. R., Turk, E. & Stearns, T. Transcriptional program of ciliated epithelial cells reveals new cilium and centrosome components and links to human disease. PLoS ONE 7, e52166 (2012)
Tanos, B. E., Yang, H. & Soni, R. Centriole distal appendages promote membrane docking, leading to cilia initiation. Genes Dev. 27, 163–168 (2013)
Lai, Y. et al. Inflammation-mediated upregulation of centrosomal protein 110, a negative modulator of ciliogenesis, in patients with chronic rhinosinusitis. J. Allergy Clin. Immunol. 128, 1207–1215.e1 (2011)
Cao, J. et al. miR-129–3p controls cilia assembly by regulating CP110 and actin dynamics. Nature Cell Biol. 14, 697–706 (2012)
Park, T. J., Mitchell, B. J., Abitua, P. B., Kintner, C. & Wallingford, J. B. Dishevelled controls apical docking and planar polarization of basal bodies in ciliated epithelial cells. Nature Genet. 40, 871–879 (2008)
Delaval, B., Covassin, L., Lawson, N. D. & Doxsey, S. Centrin depletion causes cyst formation and other ciliopathy-related phenotypes in zebrafish. Cell Cycle 10, 3964–3972 (2011)
Okada, N. et al. A positive feedback between p53 and miR-34 miRNAs mediates tumor suppression. Genes Dev. (2014)
Kott, E. et al. Loss-of-function mutations in RSPH1 cause primary ciliary dyskinesia with central-complex and radial-spoke defects. Am. J. Hum. Genet. 93, 561–570 (2013)
Kunimoto, K. et al. Coordinated ciliary beating requires Odf2-mediated polarization of basal bodies via basal feet. Cell 148, 189–200 (2012)
Deblandre, G. A., Wettstein, D. A., Koyano-Nakagawa, N. & Kintner, C. A two-step mechanism generates the spacing pattern of the ciliated cells in the skin of Xenopus embryos. Development 126, 4715–4728 (1999)
Tsao, P. N. et al. Notch signaling controls the balance of ciliated and secretory cell fates in developing airways. Development 136, 2297–2307 (2009)
D’Angiolella, V. et al. SCFCyclin Fcontrols centrosome homeostasis and mitotic fidelity through CP110 degradation. Nature 466, 138–142 (2010)
Li, J. et al. USP33 regulates centrosome biogenesis via deubiquitination of the centriolar protein CP110. Nature 495, 255–259 (2013)
Sive, H. L., Grainger, R. M. & Harland, R. M. Early Development of Xenopus laevis (Cold Spring Harbor Laboratory Press, 2000)
Walentek, P., Beyer, T., Thumberger, T., Schweickert, A. & Blum, M. ATP4a is required for Wnt-dependent Foxj1 expression and leftward flow in Xenopus left-right development. Cell Rep. 1, 516–527 (2012)
Walentek, P. et al. A novel serotonin-secreting cell type regulates ciliary motility in the mucociliary epidermis of Xenopus tadpoles. Development 141, 1526–1533 (2014)
Vladar, E. K. & Brody, S. L. Analysis of ciliogenesis in primary culture mouse tracheal epithelial cells. Methods Enzymol. 525, 285–309 (Elsevier, 2013)
Hagenlocher, C., Walentek, P., Müller, C., Thumberger, T. & Feistel, K. Ciliogenesis and cerebrospinal fluid flow in the developing Xenopus brain are regulated by foxj1. Cilia 2, 12 (2013)
Acknowledgements
We thank M. J. Bennett, M. Butler, B. Dynlacht, W. Finkbeiner, P. Kysar, B. Lee, T. Machen, B. Mitchell and J. Wallingford for constructs, technical assistance, stimulating discussions and helpful input. We also thank P. Margolis for careful reading of our manuscript. L.H. acknowledges an R01 and an R21 grant from NCI (R01 CA139067, 1R21CA175560-01), a CIRM new faculty award (RN2-00923-1), a TRDRP research grant (21RT-0133), and a research scholar award from American Cancer Society (ACS, 123339-RSG-12-265-01-RMC). R.S. acknowledges the support of Siebel postdoctoral fellowship and CIRM postdoctoral fellowship. P.W. was funded by the Deutsche Forschungsgemeinschaft (DFG, Wa 3365/1-1), and frog work in the Harland laboratory was funded by NIH grant GM42341. M.L. would like to thank M. Dobbelstein for support and discussions, and was financed by a Dorothea Schloezer Fellowship.
Author information
Authors and Affiliations
Contributions
R.S. identified and characterized PCD-like motile cilia defects in mir-34/449 TKO mice, defined miR-34/449 expression in ciliated epithelia, identified and validated cp110 as a key miR-34/449 target in mice. P.W. contributed all Xenopus data, in particular providing functional data validating cp110 as a key miR-34/449 target. N.S. performed all immunofluorescence experiments in mice, and contributed to target validation experiments. R.S. and P.W. both made significant contribution to experimental planning and result interpretation. M.L., A.K. and M.K. generated and characterized mir-449 KO mice, defined miR-449 expression patterns in mouse embryos and contributed to the revision of the manuscript. J.S.L. and G.D. contributed to histology analyses and qRT–PCR analyses. P.L. contributed to the high-speed imaging experiments. Y.W. characterized miR-34/449 expression in human respiratory epithelia. R.H. contributed to the interpretation of data and to manuscript preparation. L.H. generated mir-34a and mir-34b/34c KO mice, interpreted the data and coordinated with different groups to complete this study. R.S., P.W., N.S. and L.H. were the major contributors to the preparation of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 The generation and phenotypic characterization of mir-34/449 TKO mice.
a, miR-34/449 miRNAs are evolutionarily conserved with extensive sequence homology across many species. miR-34a has a more ancient evolutionary history compared to the rest of miR-34/449 miRNAs. miR-34a is conserved in Deuterostome, Ecdysozoa and Lophotrochozoa, yet the rest of miR-34/449 miRNAs have only vertebrate homologues. b, Diagrams of the targeted deletion strategy to generate mir-449 knockout mice. Since all mir-449 miRNAs are within intron 2 of their host gene, cdc20b, we deleted mir-449 with a minimally predicted impact on cdc20b. c, miR-449 expression is absent in mir-449−/− knockout animals, as demonstrated by real-time PCR analyses in lung tissues from littermate-controlled wild-type and mir-449−/− mice at postnatal day 35 (n = 3). miR-449a real-time PCR primers exhibit a modest cross-reaction with miR-34 miRNAs. d, mir-34/449 TKO mice have a significant postnatal attenuation in body weight. Littermate-controlled mir-34a−/−; mir-34b/34c−/−, mir-34a−/−; mir-34b/34c−/−; mir-449+/− and TKO mice were monitored for their body weight every other day for 30 days after birth. Paired t-test, ***P < 0.001. e, Surviving mir-34/449 TKO mice exhibit coughing/sneezing-like phenotype. The respiratory noise of littermate-controlled mir-34a−/−; mir-449−/− DKO and TKO mice was shown by sound wave analysis at postnatal day 30 (n = 14). f, Pulmonary inflammation occurs in a subset of mir-34/449 TKO mice. A representative H&E analysis of lung tissues from an adult TKO mouse indicates an increased infiltration by inflammatory cells. A total of 3 out of 15 TKO mice examined exhibit lung infection. g, mir-34a−/−; mir-34b/34c−/− DKO mice resemble wild-type mice, exhibiting no obvious respiratory defects in paranasal sinus or lung (n = 3). All error bars represent s.e.m.
Extended Data Figure 2 Phenotypic characterization of reproductive organs and brain in mir-34/449 TKO mice.
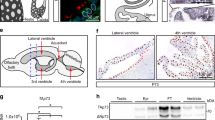
a, Adult male and female mir-34/449 TKO mice are infertile. Male (left) and female (right) reproductive organs from littermate-controlled DKO and TKO mice were subjected to H&E staining (n = 3). Boxes indicate areas depicted in Fig. 1e. b, Adult mir-34a−/−; mir-449−/− DKO male mice exhibit no defects in spermatogenesis. c, Adult mir-34a−/−; mir-34b/34c−/− DKO female mice display no defects in reproductive organs. d, The adult mir-34/449 TKO brains do not exhibit hydrocephalus, yet they are smaller in size than wild-type and DKO controls. The a/b markings on the images indicate the coronal to horizontal ratios. n = 3 for b, c and d.
Extended Data Figure 3 miR-34/449 miRNAs are enriched in airway MCCs.
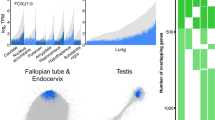
a, Most miR-34/449 miRNAs are enriched in tissues with motile cilia. Using real-time PCR, the expression of miR-34a, miR-34c and miR-449c was measured in multiple tissues from newborn, P10, P20 and adult wild-type mice. Both miR-34c and miR-449c are exclusively expressed in tissues with motile cilia, whereas miR-34a exhibits a broader expression pattern (n = 3). b, The real-time PCR assay for each miR-34/449 miRNA specifically detects the corresponding miRNA. The specificity of each miRNA real-time PCR assay was validated using testis RNA from wild-type (WT), mir-34a−/−, mir-34b/34c−/−, mir-449−/−, and TKO mice at postnatal day 35. The miR-449a assay shows a slight cross reaction with homologous miRNAs (n = 3). c, In situ hybridization of each miR-34/449 miRNA exhibits specific detection. No measurable miR-34/449 in situ signal is detected in TKO lung sections at postnatal day 25 (n = 2). d, miR-34/449 miRNAs are enriched in tracheal MCCs. In situ hybridization analyses demonstrate that miR-34c and miR-449c are specifically expressed in the tracheal MCCs, whereas miR-34a is expressed in both tracheal MCCs and the surrounding cell types (n = 2). e, mir-34/449 TKO mice do not exhibit significant alterations in Foxj1 expression. Quantification of Foxj1 positive cells (left, n = 3) and Foxj1 mRNA (right, n = 4) was performed for well-controlled wild-type, DKO and TKO tracheas, using immunofluorescence and real-time PCR, respectively. Paired t-test; ns, P > 0.05. f, mir-34a−/−; mir-449−/− DKO tracheal epithelia are morphologically indistinguishable from wild-type controls in scanning electron microscopy (SEM) analyses (n = 3). All error bars represent s.e.m.
Extended Data Figure 4 miR-34/449 deficiency causes defective ciliation and basal body docking in mouse airway MCCs.
a, mir-34/449 TKO trachea exhibit reduced MCC ciliation. Quantification of fully ciliated MCCs (γ-tub and Ac-α-tub double-positive) and partially/non-ciliated MCCs, Ac-α-tub weak/negative) was performed in littermate controlled DKO and TKO mouse tracheas, using data from all three experiments in Fig. 3a. The number of cells with MCC identity (γ-tub positive) is unaffected in TKO tracheas, yet one-third of the TKO MCCs display aberrant Ac-α-tub staining, indicating ciliation defects. Paired t-test; ns, P > 0.05, ***P < 0.001. b, The mir-34a−/−; mir-34b/34c−/− DKO mice exhibit normal ciliogenesis in tracheal MCCs. Whole tracheas from age-matched adult wild-type and mir-34a−/−; mir-34b/34c−/− DKO mice were analysed by immunofluorescence staining for Ac-α-tub (cilia) and γ-tubulin (basal bodies) (n = 3). c, mir-34/449 TKO primary tracheal epithelial cells exhibit ciliation defects in air liquid interface (ALI) culture. ALI culture of MCCs were derived from tracheas of littermate-controlled mir-34a−/−; mir-34b/34c+/−; mir-449−/− and TKO mice, and subjected to immunofluorescence staining for Ac-α-tub (cilia) and γ-tub (basal bodies). In TKO and control ALI culture, comparable levels of γ-tub positive cells are observed, however a large portion of TKO γ-tub positive cells displayed a partial or complete loss of Ac-α-tub staining. The letters on the image indicate fully (a), partially (b) or non-ciliated (c) MCCs (n = 2). d, Basal bodies fail to dock to the apical membrane of mir-34/449 TKO MCCs in ALI culture. Lateral projections of confocal micrographs described in (c) show impaired apical localization of γ-tub staining in TKO MCCs from ALI cultures, suggesting a defective basal body docking to the apical membrane. e, mir-34a−/−; mir-449−/− DKO trachea exhibit no defects in basal body docking using transmission electron microscopy (TEM) analyses (n = 3). f, TKO tracheal MCCs exhibit a defective subapical actin network. Whole tracheas from adult wild-type, mir-34a−/−; mir-34b/34c−/− DKO and TKO mice were analysed by immunofluorescence staining for Ac-α-tub (cilia) and phalloidin-488 (actin) (n = 2). All error bars represent s.e.m.
Extended Data Figure 5 Major basal body structural components are intact in mir-34/449 TKO MCCs revealed by transmission electron microscopy.
a, b, Apically docked (a) and undocked (b) basal bodies in mir-34/449 TKO MCCs have intact structural components. Basal body transition fibres (top), basal feet (middle) and striated rootlets (bottom) have comparable morphology among WT, DKO and TKO MCCs. For the top panel an arrow indicates a representative transverse view of transition fibres. For the middle panel an arrow indicates a representative transverse view of nine microtubule triplets with basal feet and arrowhead indicates a representative transverse view collected from a different height of a basal body, containing nine microtubule triplets without basal feet. For the bottom panel, an arrow indicates the longitudinal view of basal feet and arrowhead indicates the striated rootlet structure. c, Directionality of basal bodies (top) and axonemes (middle) is moderately affected in mir-34/449 TKO MCCs. In the top panel arrows point to the directions indicated by basal foot. In the middle panel red lines connecting the central pair of axonemes indicate the rotational polarity of each ciliary axoneme. In the bottom panel the angles of the axoneme directionality were statistically analysed as bidirectional circular data. The average angel was set from 0° to 180° axis. mir-34/449 TKO ciliary axonemes have moderately un-coordinated directionality compared to WT and DKO controls. d, mir-34/449 TKO axonemes exhibit intact structures, including nine outer microtubule doublets, two central microtubule singlets and dynein arms (n = 3).
Extended Data Figure 6 miR-34/449 deficiency in frog MCCs causes defective ciliogenesis without affecting cell fate specification.
a, Injection of Ctrl or miR-34/449 MOs does not affect general embryonic development or neural tube closure. Xenopus laevis embryos were injected unilaterally with MOs at the 2–4-cell stage and analysed at neurula stages (18–20). Targeting of the skin ectoderm was confirmed by co injection of fluorescent rhodamine dextran. b, Frog miR-34/449 morphants do not exhibit hydrocephalus. Embryos were injected animally with control or miR-34/449 MOs into both dorsal blastomeres at the 4-cell stage to target the neural tube and brain regions. Subsequently, the whole brains were dissected and analysed at stage 45/46. The lack of hydrocephalus in miR-34/449 morphants argues against a role of miR-34/449 in ependymal ciliation. c, Quantification of fully ciliated, partially ciliated or non-ciliated MCCs reveals no significant change in total number of MCC-fated cells in miR-34/449 morphants. Error bars represent s.d. Wilcoxon two-sample test, ns, P > 0.05. d, e, foxj1 expression and specification of MCC fate is unaltered in miR-34/449 deficient embryos. d, Embryos were unilaterally injected with Ctrl or miR-34/449 MOs to the right side at the 2–4-cell stage, cultured until stage 21 or 32 and processed for in situ hybridization to monitor foxj1 expression in the mucociliary epithelium of the skin. No change in foxj1 expression can be detected. e, Real-time PCR analysis in Ctrl or miR-34/449 MOs injected skin explants at stage 26 (onset of ciliation) does not indicate reduced expression levels of foxj1. Error bars represent s.e.m. f, miR-34/449 deficient frog embryos exhibit normal development of mucociliary cell types. Detailed analysis of the embryonic skin at stage 30–32 reveals the presence (specification and intercalation) of all cell types in miR-34/449 morphants, including large goblet cells, small secretory cells (SSC), Ac-α-tub positive ciliated cells (MCC) and non-tubulin enriched ion secreting cells (ISC). g, miR-34/449 morphant MCCs exhibit an uneven distribution of basal bodies. Sas6-gfp mRNA was injected at the 2–4-cell stage to visualize basal bodies at stage 30–32. In control embryos Sas6–GFP foci are evenly distributed in fully ciliated MCCs, whereas miR-34/449 morphant MCCs are characterized by an uneven distribution and aggregation of basal bodies, which frequently fail to grow cilia (Ac-α-tub staining). Such phenotype is characteristic for basal body docking defects. Total numbers of embryos/cells analysed were Ctrl MO (4/7) and miR-34/449 MOs (6/10).
Extended Data Figure 7 cp110 is a direct target of miR-34/449 miRNAs.
a, A schematic representation of two predicted miR-34/449 binding sites in the mouse Cp110 3′ UTR and in the luciferase reporter construct that contains Cp110 3′ UTR. b, Cp110 protein levels at postnatal day 23 are elevated in mir-34/449 TKO tracheal epithelia. c, The expression of Luc-Cp110-3′UTR exhibits miR-34b-dependent repression in NIH/3T3 cells. Error bars represent s.e.m., n = 3. Paired t-test, * P < 0.05. d, A schematic representation of one predicted miR-34/449 binding site in the frog cp110 3′UTR. A truncated cp110 construct, cp110Δ3′UTR, was made to generate a cp110 cDNA without the miR-34/449 target site. e, Real-time PCR monitoring cp110 reveals elevated mRNA expression levels of cp110 in miR-34/449 morphant frog skin explants as compared to Ctrl MO injected specimens. **P<0.01. Paired t-test. Error bars, s.e.m. f, Timeline of MCC ciliation and recapitulation of ciliation defects in skin explants (animal caps). Representative confocal images from staged whole embryos and skin explants injected with either Ctrl MO or miR-34/449 MOs show the onset of ciliation at stage 26 and fully ciliated skin ectoderm at stage 32 in whole embryos and Ctrl MO injected skin explants. miR-34/449 MOs injected skin explants develop MCC ciliogenesis defects comparable to whole embryo treatment. Cilia: Ac-α-Tub (red), Actin: Phalloidin-488 (green) and nuclei: DAPI (blue). g, Expression of cp110, foxj1 and miR-34/449 RNAs during time course of ciliation in skin explants. Explants at stage 10 represent unciliated MCC precursors, explants at stage 26 represent MCCs at the onset of ciliation, and stage 32 explants represent fully ciliated ectodermal epithelium. cp110 mRNA levels decrease over the time course of ciliation, with the strongest decrease between stage 10 and 26, while foxj1 mRNA levels rapidly increase during this time. miR-34a, -34b and -449c levels strongly increase between stage 10 and stage 26; and only a moderate increase or even decrease can be observed between stage 26 and 32, similar to foxj1 expression levels. Error bars represent s.e.m. n = 2, technical replicates on pools of 30 skin explants for each time point.
Extended Data Figure 8 miR-34/449 miRNAs promote ciliogenesis by repressing cp110.
a, Representative examples of confocal images used for quantification of MCC ciliation in (b). Embryos were stained for Ac-α-tub (cilia) and phalloidin-488 (actin). White boxes indicate areas depicted in Fig. 5c. b, Quantification of MCC ciliation in a, d and Fig. 5c. χ2-test, ns P > 0.05, ***P < 0.001. c, Centrin4–GFP incorporation into basal bodies is affected in miR-34/449 deficient embryos. The centrin4-gfp mRNA was injected at the 2–4-cell stage to visualize basal bodies in MCCs at stage 32, and centrosomes in neighbouring epithelial cells. In Ctrl morphant embryos, Centrin4–GFP staining in basal bodies (smaller foci in ciliated cells) and centrosomes (bigger foci in non-ciliated cells, green arrowheads) are equally strong. In contrast, Centrin4–GFP staining in basal bodies is greatly reduced in miR-34/449 morphants, without alteration of fluorescent intensity in centrosomes of neighbouring cells. Total numbers of embryos/cells analysed were Ctrl MO (6/17), miR-34/449 MOs (7/23). d, Representative examples of confocal images from cp110 overexpression experiments used for quantification of MCC ciliation in b. White boxes indicate areas depicted in Fig. 5c. e, The number of MCC-fated cells in miR-34/449 or cp110 morphants, and embryos injected with cp110 DNA constructs is not reduced. Quantification of total MCC numbers (fully ciliated, partially ciliated or non-ciliated MCCs) is shown for frog embryos injected with various MOs/DNAs (corresponding to a, b, d and Fig. 5c). Error bars represent s.d.
Extended Data Figure 9 Gain and loss of cp110 affects MCC basal bodies, but not the apical actin meshwork.
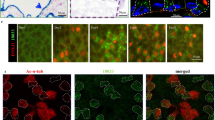
a, cp110 overexpression phenocopies miR-34/449 knockdown. Centrin4–RFP enrichment is strongly reduced in cp110Δ3′UTR overexpressing MCCs. It is noteworthy, that whereas ciliation and incorporation of Centrin4 are strongly affected in cp110Δ3′UTR injected embryos, formation of the apical actin meshwork appears largely unaffected. Together with the lack of cp110 knockdown to rescue the apical actin meshwork in miR-34/449 morphants, these data indicate an additional effect of miR-34/449 miRNAs on Actin formation/organization, which is cp110 independent. Cilia: Ac-α-tub, basal bodies: Centrin4–RFP, actin: phalloidin-488. Embryos/cells analysed: uninjected (4/7), cp110Δ3′UTR (6/10). b, The cellular basis for ciliation defects in cp110 morphants is probably due to the atypical failure of basal bodies to separate from each other, thus they appear to be aggregated in clusters of cp110-deficient MCCs. Nevertheless, Centrin4–RFP incorporation or apical localization of basal bodies is not affected in cp110 morphants. Basal bodies, Centrin4–RFP; actin: phalloidin-488. Total numbers of embryos/cells analysed: un-injected (2/3), cp110 MO (5/8). Embryos were derived from at least two females and independent fertilizations per Xenopus experiment.
Supplementary information
Postnatal mortality and coughing/sneezing-like phenotype in miR-34/449 TKO mice.
Postnatal mortality and coughing/sneezing-like phenotype in miR-34/449 TKO mice. (WMV 21479 kb)
Live imaging of multiciliary beating and mucociliary transport in the tracheas of adult littermate-controlled mir-34a-/-; mir-449-/- DKO and TKO mice.
Live imaging of multiciliary beating and mucociliary transport in the tracheas of adult littermate-controlled mir-34a-/-; mir-449-/- DKO and TKO mice. (WMV 10158 kb)
Live imaging of multiciliary beating and mucocliliary transport in the tracheas of adult wild-type and mir-34a-/-; mir34b/34c-/- DKO mice.
Live imaging of multiciliary beating and mucocliliary transport in the tracheas of adult wild-type and mir-34a-/-; mir34b/34c-/- DKO mice. (WMV 7885 kb)
Rights and permissions
About this article
Cite this article
Song, R., Walentek, P., Sponer, N. et al. miR-34/449 miRNAs are required for motile ciliogenesis by repressing cp110. Nature 510, 115–120 (2014). https://doi.org/10.1038/nature13413
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13413
This article is cited by
-
Targeted therapy using nanocomposite delivery systems in cancer treatment: highlighting miR34a regulation for clinical applications
Cancer Cell International (2023)
-
A local translation program regulates centriole amplification in the airway epithelium
Scientific Reports (2023)
-
Non-coding RNAs and chromatin: key epigenetic factors from spermatogenesis to transgenerational inheritance
Biological Research (2021)
-
Bicc1 and Dicer regulate left-right patterning through post-transcriptional control of the Nodal inhibitor Dand5
Nature Communications (2021)
-
The roles of microRNAs in mouse development
Nature Reviews Genetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.