Abstract
PTEN encodes a lipid phosphatase that is underexpressed in many cancers owing to deletions, mutations or gene silencing1,2,3. PTEN dephosphorylates phosphatidylinositol (3,4,5)-triphosphate, thereby opposing the activity of class I phosphatidylinositol 3-kinases that mediate growth- and survival-factor signalling through phosphatidylinositol 3-kinase effectors such as AKT and mTOR2. To determine whether continued PTEN inactivation is required to maintain malignancy, here we generate an RNA interference-based transgenic mouse model that allows tetracycline-dependent regulation of PTEN in a time- and tissue-specific manner. Postnatal Pten knockdown in the haematopoietic compartment produced highly disseminated T-cell acute lymphoblastic leukaemia. Notably, reactivation of PTEN mainly reduced T-cell leukaemia dissemination but had little effect on tumour load in haematopoietic organs. Leukaemia infiltration into the intestine was dependent on CCR9 G-protein-coupled receptor signalling, which was amplified by PTEN loss. Our results suggest that in the absence of PTEN, G-protein-coupled receptors may have an unanticipated role in driving tumour growth and invasion in an unsupportive environment. They further reveal that the role of PTEN loss in tumour maintenance is not invariant and can be influenced by the tissue microenvironment, thereby producing a form of intratumoral heterogeneity that is independent of cancer genotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Li, J. et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275, 1943–1947 (1997)
Engelman, J. A., Luo, J. & Cantley, L. C. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nature Rev. Genet. 7, 606–619 (2006)
Salmena, L., Carracedo, A. & Pandolfi, P. P. Tenets of PTEN tumor suppression. Cell 133, 403–414 (2008)
Dickins, R. A. et al. Probing tumor phenotypes using stable and regulated synthetic microRNA precursors. Nature Genet. 37, 1289–1295 (2005)
Dickins, R. A. et al. Tissue-specific and reversible RNA interference in transgenic mice. Nature Genet. 39, 914–921 (2007)
Premsrirut, P. K. et al. A rapid and scalable system for studying gene function in mice using conditional RNA interference. Cell 145, 145–158 (2011)
Di Cristofano, A., Pesce, B., Cordon-Cardo, C. & Pandolfi, P. P. Pten is essential for embryonic development and tumour suppression. Nature Genet. 19, 348–355 (1998)
Stambolic, V. et al. High incidence of breast and endometrial neoplasia resembling human Cowden syndrome in pten+/− mice. Cancer Res. 60, 3605–3611 (2000)
Gutierrez, A. et al. High frequency of PTEN, PI3K, and AKT abnormalities in T-cell acute lymphoblastic leukemia. Blood 114, 647–650 (2009)
Kim, W. I., Wiesner, S. M. & Largaespada, D. A. Vav promoter-tTA conditional transgene expression system for hematopoietic cells drives high level expression in developing B and T cells. Exp. Hematol. 35, 1231–1239 (2007)
Suzuki, A. et al. T cell-specific loss of Pten leads to defects in central and peripheral tolerance. Immunity 14, 523–534 (2001)
Aifantis, I., Raetz, E. & Buonamici, S. Molecular pathogenesis of T-cell leukaemia and lymphoma. Nature Rev. Immunol. 8, 380–390 (2008)
Guo, W. et al. Multi-genetic events collaboratively contribute to Pten-null leukaemia stem-cell formation. Nature 453, 529–533 (2008)
Erikson, J. et al. Deregulation of c-myc by translocation of the alpha-locus of the T-cell receptor in T-cell leukemias. Science 232, 884–886 (1986)
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012)
Engelman, J. A. et al. Effective use of PI3K and MEK inhibitors to treat mutant Kras G12D and PIK3CA H1047R murine lung cancers. Nature Med. 14, 1351–1356 (2008)
Nehls, M. et al. Two genetically separable steps in the differentiation of thymic epithelium. Science 272, 886–889 (1996)
Bleul, C. C. & Boehm, T. Chemokines define distinct microenvironments in the developing thymus. Eur. J. Immunol. 30, 3371–3379 (2000)
Nowell, C. S. et al. Foxn1 regulates lineage progression in cortical and medullary thymic epithelial cells but is dispensable for medullary sublineage divergence. PLoS Genet. 7, e1002348 (2011)
Wurbel, M. A. et al. The chemokine TECK is expressed by thymic and intestinal epithelial cells and attracts double- and single-positive thymocytes expressing the TECK receptor CCR9. Eur. J. Immunol. 30, 262–271 (2000)
Campbell, D. J. & Butcher, E. C. Intestinal attraction: CCL25 functions in effector lymphocyte recruitment to the small intestine. J. Clin. Invest. 110, 1079–1081 (2002)
Youn, B. S., Kim, C. H., Smith, F. O. & Broxmeyer, H. E. TECK, an efficacious chemoattractant for human thymocytes, uses GPR-9–6/CCR9 as a specific receptor. Blood 94, 2533–2536 (1999)
Uehara, S., Grinberg, A., Farber, J. M. & Love, P. E. A role for CCR9 in T lymphocyte development and migration. J. Immunol. 168, 2811–2819 (2002)
Buonamici, S. et al. CCR7 signalling as an essential regulator of CNS infiltration in T-cell leukaemia. Nature 459, 1000–1004 (2009)
Walters, M. J. et al. Characterization of CCX282-B, an orally bioavailable antagonist of the CCR9 chemokine receptor, for treatment of inflammatory bowel disease. J. Pharmacol. Exp. Ther. 335, 61–69 (2010)
Eksteen, B. & Adams, D. H. GSK-1605786, a selective small-molecule antagonist of the CCR9 chemokine receptor for the treatment of Crohn’s disease. IDrugs 13, 472–781 (2010)
Hollander, M. C., Blumenthal, G. M. & Dennis, P. A. PTEN loss in the continuum of common cancers, rare syndromes and mouse models. Nature Rev. Cancer 11, 289–301 (2011)
Jotta, P. Y. et al. Negative prognostic impact of PTEN mutation in pediatric T-cell acute lymphoblastic leukemia. Leukemia 24, 239–242 (2010)
Muellner, M. K. et al. A chemical-genetic screen reveals a mechanism of resistance to PI3K inhibitors in cancer. Nature Chem. Biol. 7, 787–793 (2011)
Ilic, N., Utermark, T., Widlund, H. R. & Roberts, T. M. PI3K-targeted therapy can be evaded by gene amplification along the MYC-eukaryotic translation initiation factor 4E (eIF4E) axis. Proc. Natl Acad. Sci. USA 108, E699–E708 (2011)
Huesken, D. et al. Design of a genome-wide siRNA library using an artificial neural network. Nature Biotechnol. 23, 995–1001 (2005)
Fellmann, C. et al. Functional identification of optimized RNAi triggers using a massively parallel sensor assay. Mol. Cell 41, 733–746 (2011)
Dow, L. E. et al. A pipeline for the generation of shRNA transgenic mice. Nature Protocols 7, 374–393 (2012)
Beard, C., Hochedlinger, K., Plath, K., Wutz, A. & Jaenisch, R. Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis 44, 23–28 (2006)
Kistner, A. et al. Doxycycline-mediated quantitative and tissue-specific control of gene expression in transgenic mice. Proc. Natl Acad. Sci. USA 93, 10933–10938 (1996)
Suzuki, A. et al. High cancer susceptibility and embryonic lethality associated with mutation of the PTEN tumor suppressor gene in mice. Curr. Biol. 8, 1169–1178 (1998)
Berger, R. L. & Boos, D. D. P-values maximized over a confidence set for the nuisance parameter. J. Am. Stat. Assoc. 89, 1012–1016 (1994).
Scuoppo, C. et al. A tumour suppressor network relying on the polyamine–hypusine axis. Nature 487, 244–248 (2012)
Lakshmi, B. et al. Mouse genomic representational oligonucleotide microarray analysis: detection of copy number variations in normal and tumor specimens. Proc. Natl Acad. Sci. USA 103, 11234–11239 (2006)
Zhao, Z. et al. p53 loss promotes acute myeloid leukemia by enabling aberrant self-renewal. Genes Dev. 24, 1389–1402 (2010)
Tubo, N. J. et al. A systemically-administered small molecule antagonist of CCR9 acts as a tissue-selective inhibitor of lymphocyte trafficking. PLoS ONE 7, e50498 (2012)
Gärtner, F. et al. Immature thymocytes employ distinct signaling pathways for allelic exclusion versus differentiation and expansion. Immunity 10, 537–546 (1999)
O'Neil et al. Activating Notch1 mutations in mouse models of T-ALL. Blood 107, 781–785 (2006)
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nature Biotechnol. 31, 46–53 (2013)
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004)
Hochberg, Y. & Benjamini, Y. More powerful procedures for multiple significance testing. Stat. Med. 9, 811–818 (1990)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Subramanian, A., Kuehn, H., Gould, J., Tamayo, P. & Mesirov, J. P. GSEA-P: a desktop application for Gene Set Enrichment Analysis. Bioinformatics 23, 3251–3253 (2007)
Huang da. W, Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57 (2009)
Acknowledgements
We are grateful to J. Cappellani, D. Grace, J. Simon and M. Taylor for technical assistance, S.-Y. Kim for performing tetraploid embryo complementation, J. Cheng for RNA-seq analysis, M. Riggs for CGH array analysis, M. Lupu and C. Le for MRI imaging, V. Longo and P. Zanzonico for PET analysis, J. Pichardo for data management, L. Lopez and A. Giri for tissue microarray construction, M. Saborowski for help with IHC staining, A. Roselló-Díez and M. Asher for expertise in antibody staining characterization and C. Sherr for constructive comments and editorial advice. We thank J. Jaen, M. Penfold and M. Walters from ChemoCentryx for providing the CCR9 small molecule inhibitor. C.M. was supported by a fellowship from the DFG (Mi1210/1-1). I.A. received support by a fellowship from the Deutsche Krebshilfe (DKH no. 109902). C.S. was supported by the Angel Foundation with a Curt Engelhorn fellowship. This work was also supported by a program project grant from the National Cancer Institute, a Leukemia and Lymphoma Society Specialized Center of Research, and philanthropic funds from the Don Monti Foundation. S.W.L. is supported by the Geoffrey Beene Foundation and is an investigator at the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.M. and S.W.L. designed the study. C.M., C.S., P.K.P. and L.L. performed shRNA design and testing, targeting vector construction and E.S. cell targeting. C.M., B.B., I.A., C.S. and L.L. performed mouse breeding, transplantation experiments and analysed data. C.M. and I.A. performed in vitro migration assays and analysed data. C.M. and H.B. ran the mouse MRI experiments and analysed data. C.M. and B.B. performed the 18F-FDG-PET experiments and analysed data. J.H. performed CGH analysis, and J.H. and C.M. analysed data. J.T.-F. performed the histopathological analysis of mouse and human tumours, and J.T-F. and C.M. analysed data. B.B. and C.M. performed paraffin embedding, sectioning and IHC staining of mouse tissues and analysed data. C.M. performed flow cytometry, immunoblotting and analysed data. J.N. performed the RNA-seq sample processing and J.N., J.M., J.R.D., C.S. and C.M. analysed data. C.S. ran the GSEA analysis and comparison with human expression data. M.A. and M.R.S. performed the SKY analysis of mouse tumours, and M.A., M.R.S. and C.M. analysed data. S.W.L. supervised the project. C.M., C.S., B.B. and S.W.L. wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Pten shRNA-transgenic mice enable conditional expression of PTEN and develop tumours after prolonged Pten knockdown.
a, Western blot analysis of PTEN protein knockdown in NIH 3T3 cells infected with different Pten shRNAs at low multiplicity of infection. l.e., long exposure; s.e., short exposure. b, PTEN protein knockdown assessed by western blot in ES cell clones targeted with two different Pten shRNAs, either treated with Dox or left untreated. c, MEFs from Rosa26-rtTA;shPten.1522 transgenic mice, wild-type control mice, or Pten+/− mice were treated with Dox for the indicated times and analysed for PTEN, GFP and ACTB expression by western blot. d, Overall survival of mice receiving bone marrow cells from tTA-transgenic mice infected with an inducible TRE-GFP-miR-30 (TGM) retroviral vector expressing shPten.1522, shPten.2049 or control after irradiation with 600 rad. e, Fluorescence image of a CAGGS-rtTA;shPten.1522 mouse on Dox for 5 days and a CAGGS-rtTA-only control mouse. f, Flow cytometric analysis of the peripheral blood of a CAGGS-rtTA;shPten.1522 mouse on Dox and an off Dox control mouse for myeloid (CD11b) and GFP marker expression 10 days after initiating Dox food. g, Overall survival curve of CMV-rtTA;shPten.1522 double-transgenic and control mice (single-transgenic shPten.1522 or CMV-rtTA). Dox treatment for shRNA induction was started after weaning (at ∼4 weeks of age). h, Situs of a tumour-bearing CAGGS-rtTA;shPten.1522 double-transgenic mouse. A large thymic tumour (full arrow), as well as enlarged lymph nodes (dashed arrows) and spleen (arrowhead), are visible. i, Immunohistochemical staining of kidney sections from a CAGGS-rtTA;shPten.1522 mouse for the indicated antigens. Arrows highlight a tumour infiltrate around a kidney venule. Scale bars, 100 μm.
Extended Data Figure 2 Vav-tTA;shPten transgenic mice with targeted shPten expression in the haematopoietic lineage display thymic hyperplasia by 6 weeks, and a subset develops thymic tumours infiltrating multiple peripheral organs.
a, Brightfield (left) and fluorescence (right) images of spleen and thymus from Vav-tTA;shLuc (control) and Vav-tTA;shPten double-transgenic mice at 5 weeks. b, Florescence-activated cell sorting analysis of spleen and thymus single-cell suspensions from Vav-tTA;shLuc/shPten mice for CD4/CD8 expression. c, IHC analysis of thymic tissue from 6-week-old Vav-tTA;shLuc and Vav-tTA;shPten mice. Sections were stained with haematoxylin and eosin (H&E), anti-GFP or anti-PTEN antibodies, showing heterogeneous GFP staining and correspondingly variable PTEN knockdown. Scale bars, 200 μm for H&E and GFP, 100 μm for PTEN. The insets are ×2 magnifications. d, Thymus weight of 6-week-old Vav-tTA;shLuc and Vav-tTA;shPten mice (n = 5 for both groups, P < 0.006 by t-test). e, Brightfield and GFP-fluorescence images of a Vav-tTA;shPten mouse with tumours. f, IHC staining of spleen, liver and kidney tissues from a Vav-tTA;shPten mouse with primary T-cell disease. Sections were stained for H&E, GFP, PTEN and phospho-AKT(S473) as indicated, showing heterogeneous staining owing to variable tumour infiltration. Scale bars, 100 μm, insets 25 μm.
Extended Data Figure 3 Immunophenotype, chromosomal aberrations and Notch1 mutations observed in murine shPten tumours.
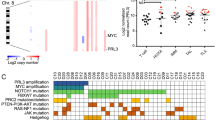
a, Flow cytometric analysis of organ infiltration by primary tumours in Vav-tTA;shPten.1522 transgenic mice. Single-cell suspensions of indicated tissues were analysed for eGFP, Thy1.2, CD4 and CD8 expression. BM, bone marrow; LN, lymph node; PB, peripheral blood. b, Spectral karyotyping analysis of a T-cell tumour arising in Vav-tTA;shPten.1522 mice, showing a t(14;15) translocation. c, CGH analysis of a Vav-tTA;shPten.1522 leukaemia. Genomic tumour DNA was analysed on Affymetrix CGH SNP arrays and compared to normal skin tissue. x axis indicates genomic coordinate and y axis represents log2(tumour/germline). d, Schematic of the murine NOTCH1 protein generated using protein paint (http://explore.pediatriccancergenomeproject.org/proteinPainter), highlighting the different NOTCH1 protein domains and the mutations detected in the murine shPten and Pten−/− T-ALL tumours.
Extended Data Figure 4 Summary of karyotyping and Notch1 sequencing of shPten T-ALL tumours.
a, Results from a multiplex FISH analysis of three different primary shPten-induced T-ALL tumours. At least ten cells were analysed for each sample, and chromosomal gains, deletions or translocations are highlighted. b, Summary of Notch1 mutations identified in shPten and Pten knockout tumours. c, Sequence of all shRNAs targeting murine Pten that were tested in the study. Sense and guide strand are highlighted in red.
Extended Data Figure 5 GSEA shows similar gene expression patterns in human and mouse T-ALL lacking Pten.
a, GSEA of a mouse shPten signature in PTEN-altered human T-ALL was tested after establishing the shPten-dependent signature using the 100 most upregulated genes in shPten T-ALL samples (untreated, n = 3) against PTEN-restored samples (Dox-treated, n = 4) as determined by RNA-seq analysis (data not shown). Publicly available human T-ALL gene expression profiles (GSE28703, n = 47) were processed using RMA (quantile normalization) and supervised for PTEN status (PTEN altered including PTEN deletion, mutation or both, n = 10; PTEN wild-type (WT), n = 37) according to the published sample annotation18. Statistical significance of GSEA results was assessed using 1,000 samples permutations. b, For enrichment of human PTEN T-ALL signature in mouse shPten knockdown (kd) T-ALL (Dox-off) profiles against Pten-restored (Dox-on) profiles, a human PTEN-disrupted signature was generated by including the 100 most upregulated genes in PTEN-disrupted versus PTEN-wild type T-ALL samples. Mouse genes were ranked by supervising untreated to Dox-treated shPten T-ALLs. Statistical significance of human PTEN-disrupted signature enrichment was assessed using 1,000 gene set permutations.
Extended Data Figure 6 Secondary recipients of shPten T-ALL cells display extensive intestinal tumour infiltration similar to a subset of human patients characterized by peripheral T-cell lymphoma and low PTEN expression.
a, Overall survival of sublethally irradiated Rag1−/− mice transplanted with 1 × 105 T-ALL cells from primary Vav-tTA;shPten.1522 mice compared to untransplanted mice (n = 5 for both groups, P < 0.003). b, IHC staining for eGFP expression in the indicated tissues from secondary T-ALL transplant recipients. Scale bars, 400 μm, 100 μm for insets. c, Overall survival of PTEN normal (WT) versus PTEN-altered patients with T-ALL analysed from published data on patients with T-ALL15, P = 0.02). PTEN-altered (n = 20) includes patients with PTEN deletion, mutation, underexpression (<0.8 sigma after z scoring) and any combination of such alterations; PTEN normal (n = 62) include all other patients with available data. d, PTEN IHC staining of tissue microarrays of tumour sections from Memorial Sloan Kettering Cancer Center patients with peripheral T-cell lymphomas. Examples of low (top panel) and high (bottom panel) PTEN expression samples are shown. e, Contingency table showing a significant association (P < 0.003; Fisher’s exact test) between low expression of PTEN and intestinal infiltration in PTCL patients. f, Overall survival of Rag1−/− mice transplanted with T-ALL cells from Ptenfl/fl; Lck-cre mice ± Dox (n = 5 for each group). g, Weight of spleen (n = 4) and lymph nodes (n = 8) in Rag1−/− mice transplanted with Vav-tTA;shPten leukaemic cells untreated or treated with Dox for 5 days (± s.d.). h, MRI of Rag1−/− mice transplanted with Vav-tTA;shPten leukaemic cells untreated or treated with Dox for 5 days, 14 days after transplant. Arrows highlight lymph nodes (LN) and increased signals in the liver. Representative images for one out of three analysed mice per condition are shown. i, MRI imaging of the intestine and liver of the same mice as in h are shown. Dashed arrows highlight the liver, full arrows the intestine.
Extended Data Figure 7 Pten reactivation affects multiple pathways and increases apoptosis in tumour cells infiltrating the intestine, but not in the spleen.
a, b, Heatmap of top 30 upregulated (a) and downregulated (b) genes after Pten reactivation as determined by RNA-seq on CD4-sorted leukaemic samples isolated from the spleen. Three mice with Pten knocked down and four mice with reactivated Pten were analysed. Pten is one of the top 50 upregulated genes after reactivation, but is not included on the list. c, Bubblegraph visualization of the most significantly affected pathways as determined by DAVID pathway analysis. y axis represents relative pathway enrichment in Pten reactivated versus Pten knockdown leukaemic cells, and size of the bubble graph is inversely proportional to P value. d, IHC analysis for expression of GFP and PTEN in spleen, lymph node (LN) and liver from shPten-tumour-transplanted mice ± Dox treatment (5 days after start of Dox treatment; n = 3 per group). Representative sections are shown. Scale bars, 100 μm for full images, 20 μm for insets. e, In vivo 5-bromodeoxyuridine (BrdU) uptake in leukaemic cells isolated from the lymph nodes of mice transplanted with Vav-tTA;shPten primary T-ALL tumours ± Dox. n = 3 for each group (± s.d.). f, TUNEL staining of spleen and intestinal sections of Rag1−/− mice serially transplanted with Vav-tTA;shPten leukaemia cells and either left untreated or treated with Dox 24 h before sectioning. Scale bars, 200 μm (×2.5) and 50 μm (×10). g, Quantification of TUNEL-stained sections from the intestinal sections in f. TUNEL positive cells from three representative areas of 1 mm2 from two different intestine sections were counted for each condition (P < 0.01) (± s.d.).
Extended Data Figure 8 AKT and S6 protein phosphorylation is affected by PTEN reactivation in the intestine.
a, IHC staining for phospho-S6 (pS235/236-S6) and phospho-AKT (pS473AKT) of spleen sections from Rag1−/− mice transplanted with Vav-tTA;shPten tumour cells from primary mice and either treated with Dox or left untreated 2 days after treatment begin (n = 3 per group). Scale bars, 100 μm, 20 μm for insets. Representative images are shown. b, IHC staining for pS473-AKT (bottom) in the intestine, showing very low pAKT signal in the intestinal epithelial cells independent of Dox treatment status (arrows; bottom left and right panels), conversely strong staining for pAKT was detected in some of the infiltrating tumour cells (arrow heads). The signal was reduced concomitantly with the overall reduction of the Pten-shRNA-linked GFP signal (top) after 36 h of Dox treatment (+ Dox; right panels). c, Representative histogram of flow cytometric analysis for intracellular pS6 signal in CD4+ cells isolated from spleen and intestine of Rag1−/− mice transplanted with shPten tumour cells and either treated with Dox for 5 days or left untreated. d, Flow cytometric quantification of pS6 signal in CD4+ cells isolated from the intestine (d) and spleens (e) of Rag1−/− mice transplanted with primary shPten tumours and treated ± Dox for 5 days (n = 4 for each condition, P < 0.04 for the intestine and not significant for the spleen by paired t-test). MFI, mean fluorescent intensity, error bars represent s.d.
Extended Data Figure 9 NCr mice display a reduced intestinal tumour infiltration, which is not dependent on the absence of the thymus.
a, Brightfield pictures of the intestinal sites of Rag1−/− and NCr nude mice serially transplanted with shPten tumours (top four panels) and fluorescence images (FI) of cells infiltrating the small intestine in these mice (bottom four panels). Scale bars, 800 μm (top FI panels) and 100 μm (bottom FI panels). Pictures were taken on a Nikon SMZ 1000 stereomicroscope. b, Quantification of the intestinal infiltration in transplanted Rag1−/− or NCr mice by flow cytometry (P < 0.03). c, d, Weight of lymph nodes (P < 0.01) (c) and spleens (P = n.s.) (d) in transplanted Rag1−/− and NCr mice. e, CCL25 expression in the small intestine of Rag1−/− and NCr mice measured by ELISA. Error bars in b–e show s.d. f, Western blot analysis of CCL25 expression in the small intestine of Rag1−/− and NCr mice. g, Overall survival of Rag1−/− and thymectomized Rag1−/− mice after transplant with shPten T-ALL cells (n = 5 per group). h, H&E and immunohistochemical analysis of CD3 expression of spleen, liver and intestine from Rag1−/− and thymectomized Rag1−/− mice transplanted with shPten T-ALL cells. Scale bars, 200 μm for spleen and liver and 100 μm for intestinal samples. i, Flow cytometric measurement of CCR9 expression on shPten leukaemia cells either in the absence of Dox (Pten knocked down) or Dox-treated (Pten reactivated). One representative analysis out of four analysed on/off Dox pairs is shown. A CCR9 negative B-cell line was used as control. j, Immunoblot analysis of PTEN, p-AKT(S473) and ACTB expression in human HBP-ALL T-ALL cells infected with either a control shRNA (shRenilla) or a shRNA targeting PTEN, and either starved or stimulated for 15 min with 500 ng ml−1 CCL25. k, shPten tumour cell migration across a Boyden chamber in the presence or absence of 1 μg ml−1 Dox and 500 ng ml−1 CCL25. One representative experiment of two is shown, samples were run in triplicate; **P < 0.01, ***P < 0.001 by t-test (± s.d.).
Extended Data Figure 10 CCR9 inactivation by shRNA knockdown or by pharmacologic inhibition attenuates intestinal tumour infiltration.
a, CCR9 expression on the surface of shPten tumour cells either infected with a control shRNA (shRenilla) (left) or with a shRNA targeting Ccr9 (right) as measured by flow cytometry, compared to uninfected cells, respectively. b, Flow cytometry-based quantification of CCR9 suppression in shCcr9-infected shPten T-ALL cells compared to shRenilla-infected cells, n = 5 for each cohort (± s.d.). c, Raw percentage of shRenilla/shCcr9-expressing shPten T-ALL cells ± s.d. in different tissue compartments of mice 12 days after transplantation, determined by flow cytometry, n = 5 for each cohort. P < 0.0005 (intestine). d, IHC analysis for mCherry (left, shRenilla-mCherry; right, shCcr9-mCherry)-expressing cells in tissue sections of mice from c. Spleen, liver and intestinal sections of mice transplanted with shRenilla- or shCcr9-infected T-ALL cells were analysed for mCherry expression. Representative stains from one mouse out of three analysed mice are shown. Scale bars, 100 μm, insets 20 μm. e, IHC staining for eGFP expression in representative sections of small intestine, liver and spleen of Vav-tTA;shPten-tumour-bearing mice treated with vehicle or the CCR9 inhibitor CCX8037 (n = 3). Scale bars, 400 μm (×2.5) and 100 μm (×10). f, Flow cytometric quantification of intestinal tumour infiltration in Rag1−/− mice transplanted with Vav-tTA;shPten leukaemia cells and treated with vehicle (n = 4) or a small molecule inhibitor of CCR9 (n = 5). *P < 0.05 by t-test (± s.d.). g, Immunoblot analysis of p-AKT expression 15 min after stimulation of shPten leukaemia cells with CCL25 in the absence or presence of indicated concentrations of CCX8037. h, IHC analysis of eGFP and p-AKT signal in representative sections of small intestine from Vav-tTA;shPten-tumour-bearing mice treated with vehicle or the CCR9 inhibitor CCX8037. Scale bars, 100 μm, 25 μm for insets.
Supplementary information
3D reconstruction of 18F-FDG PET/CT imaging of a mouse with PTEN-inactivated shPten T-ALL (without Dox treatment)
18F-FDG uptake at 12 days after transplantation with 1x105 shPten T-ALL cells without Dox treatment (PTEN knocked down) was measured by combined PET/CT imaging on a Inveon MicroPET/CT system (Siemens, NY, USA). 3D reconstruction was performed using the Inveon Workplace software package (Siemens Medical Solutions USA, PA, USA). Significant uptake was observed in the intestine, liver and spleen. Additionally, there is physiologic tracer enrichment in the heart and bladder. (MP4 2717 kb)
3D reconstruction of 18F-FDG PET/CT imaging of a mouse with PTEN-reactivated shPten T-ALL (with Dox treatment)
18F-FDG uptake at 12 days after transplantation with 1x105 shPten T-ALL cells (4 days after begin of Dox treatment to reexpress PTEN) was measured by combined PET/CT imaging on a Inveon MicroPET/CT system (Siemens, NY, USA). 3D reconstruction was performed using the Inveon Workplace software package (Siemens Medical Solutions USA, PA, USA). Significant uptake was observed in the spleen, whereas liver and intestine show only a weak signal. Also in the Dox treated mice, there is physiologic tracer enrichment in the heart and bladder. (MP4 2890 kb)
Rights and permissions
About this article
Cite this article
Miething, C., Scuoppo, C., Bosbach, B. et al. PTEN action in leukaemia dictated by the tissue microenvironment. Nature 510, 402–406 (2014). https://doi.org/10.1038/nature13239
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13239
This article is cited by
-
CCR9 initiates epithelial–mesenchymal transition by activating Wnt/β-catenin pathways to promote osteosarcoma metastasis
Cancer Cell International (2021)
-
Targeting chemokines for acute lymphoblastic leukemia therapy
Journal of Hematology & Oncology (2021)
-
Emerging role of PTEN loss in evasion of the immune response to tumours
British Journal of Cancer (2020)
-
Abrogation of glutathione peroxidase-1 drives EMT and chemoresistance in pancreatic cancer by activating ROS-mediated Akt/GSK3β/Snail signaling
Oncogene (2018)
-
Off and back-on again: a tumor suppressor’s tale
Oncogene (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.