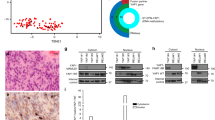
Abstract
Members of the nuclear factor-κB (NF-κB) family of transcriptional regulators are central mediators of the cellular inflammatory response. Although constitutive NF-κB signalling is present in most human tumours, mutations in pathway members are rare, complicating efforts to understand and block aberrant NF-κB activity in cancer. Here we show that more than two-thirds of supratentorial ependymomas contain oncogenic fusions between RELA, the principal effector of canonical NF-κB signalling, and an uncharacterized gene, C11orf95. In each case, C11orf95–RELA fusions resulted from chromothripsis involving chromosome 11q13.1. C11orf95–RELA fusion proteins translocated spontaneously to the nucleus to activate NF-κB target genes, and rapidly transformed neural stem cells—the cell of origin of ependymoma—to form these tumours in mice. Our data identify a highly recurrent genetic alteration of RELA in human cancer, and the C11orf95–RELA fusion protein as a potential therapeutic target in supratentorial ependymoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
Sequence and array data were deposited in the European Bioinformatics Institute (EBI) under accession number EGAS00001000254.
References
Kleihues, P. et al. The WHO classification of tumors of the nervous system. J. Neuropathol. Exp. Neurol. 61, 215–225; discussion. 226–219 (2002)
Merchant, T. E. et al. Conformal radiotherapy after surgery for paediatric ependymoma: a prospective study. Lancet Oncol. 10, 258–266 (2009)
Modena, P. et al. Identification of tumor-specific molecular signatures in intracranial ependymoma and association with clinical characteristics. J. Clin. Oncol. 24, 5223–5233 (2006)
Taylor, M. D. et al. Radial glia cells are candidate stem cells of ependymoma. Cancer Cell 8, 323–335 (2005)
Puget, S. et al. Candidate genes on chromosome 9q33–34 involved in the progression of childhood ependymomas. J. Clin. Oncol. 27, 1884–1892 (2009)
Johnson, R. A. et al. Cross-species genomics matches driver mutations and cell compartments to model ependymoma. Nature 466, 632–636 (2010)
Witt, H. et al. Delineation of two clinically and molecularly distinct subgroups of posterior fossa ependymoma. Cancer Cell 20, 143–157 (2011)
Atkinson, J. M. et al. An integrated in vitro and in vivo high-throughput screen identifies treatment leads for ependymoma. Cancer Cell 20, 384–399 (2011)
Wang, J. et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nature Methods 8, 652–654 (2011)
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011)
Nolan, G. P., Ghosh, S., Liou, H. C., Tempst, P. & Baltimore, D. DNA binding and IκB inhibition of the cloned p65 subunit of NF-κB, a rel-related polypeptide. Cell 64, 961–969 (1991)
Hansen, S. K., Baeuerle, P. A. & Blasi, F. Purification, reconstitution, and I kappa B association of the c-Rel-p65 (RelA) complex, a strong activator of transcription. Mol. Cell. Biol. 14, 2593–2603 (1994)
DiDonato, J. A., Mercurio, F. & Karin, M. NF-κB and the link between inflammation and cancer. Immunol. Rev. 246, 379–400 (2012)
Perkins, N. D. The diverse and complex roles of NF-κB subunits in cancer. Nature Rev. Cancer 12, 121–132 (2012)
Baud, V. & Karin, M. Is NF-κB a good target for cancer therapy? Hopes and pitfalls. Nature Rev. Drug Discov. 8, 33–40 (2009)
Chaturvedi, M. M., Sung, B., Yadav, V. R., Kannappan, R. & Aggarwal, B. B. NF-κB addiction and its role in cancer: ‘one size does not fit all’. Oncogene 30, 1615–1630 (2011)
Guttridge, D. C., Albanese, C., Reuther, J. Y., Pestell, R. G. & Baldwin, A. S. NF-κB controls cell growth and differentiation through transcriptional regulation of cyclin d1. Mol. Cell. Biol. 19, 5785–5799 (1999)
Hinz, M. et al. NF-κB function in growth control: regulation of cyclin d1 expression and G0/G1-to-S-phase transition. Mol. Cell. Biol. 19, 2690–2698 (1999)
Kiefel, H., Pfeifer, M., Bondong, S., Hazin, J. & Altevogt, P. Linking L1CAM-mediated signaling to NF-κB activation. Trends Mol. Med. 17, 178–187 (2011)
Kiefel, H. et al. EMT-associated up-regulation of L1CAM provides insights into L1CAM-mediated integrin signalling and NF-κB activation. Carcinogenesis 33, 1919–1929 (2012)
Courtois, G. & Gilmore, T. D. Mutations in the NF-κB signaling pathway: implications for human disease. Oncogene 25, 6831–6843 (2006)
Godfraind, C. et al. Distinct disease-risk groups in pediatric supratentorial and posterior fossa ependymomas. Acta Neuropathol. 124, 247–257 (2012)
Chen, L.-F. & Greene, W. C. Shaping the nuclear action of NF-κB. Nature Rev. Mol. Cell Biol. 5, 392–401 (2004)
Zhong, H., Voll, R. E. & Ghosh, S. Phosphorylation of NF-κB p65 by PKA stimulates transcriptional activity by promoting a novel bivalent interaction with the coactivator CBP/p300. Molecular Cell 1, 661–671 (1998)
Okazaki, T. et al. Phosphorylation of serine 276 is essential for p65 NF-κB subunit-dependent cellular responses. Biochem. Biophys. Res. Commun. 300, 807–812 (2003)
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012)
Robinson, G. et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 488, 43–48 (2012)
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013)
Acknowledgements
This research was supported as part of the St. Jude Children’s Research Hospital, Washington University Pediatric Cancer Genome Project. This work was supported by grants from the National Institutes of Health (R01CA129541, P01CA96832 and P30CA021765 to R.J.G), the Collaborative Ependymoma Research Network (CERN), and by the American Lebanese Syrian Associated Charities (ALSAC). We are grateful to S. Temple for the gift of reagents and the staff of the Hartwell Center for Bioinformatics and Biotechnology, Animal Imaging Center, and Flow Cytometry and Cell Sorting Shared Resource at St. Jude Children’s Research Hospital for technical assistance.
Author information
Authors and Affiliations
Contributions
M.P., K.M.M., C.P., R.W., J.D.D., R.L., R.G.T., T.N.P., R.T., E.W., B.T., W.O., K.G., M.R., X.C., P.N., E.H., D.F., G.W., S.S., J.E., K.B., D.Y., B.V., H.L.M., J.B., P.G., R.H., J.M., G.S., L.D., C.L., K.O., D.Z., R.S.F., L.L.F. Yo.L., Yu.L., A.G., A.A.S., F.B. and T.M. contributed to the design and conduct of experiments and to the writing. E.R.M., R.K.W., J.R.D. and D.R.G. contributed to experimental design and to the writing. J.Z., D.W.E. and R.J.G. conceived the research and designed, directed and wrote the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Next-generation sequencing coverage of ependymoma samples.
a, Coverage for whole-genome sequenced cases. Percent of the genome (left) and exome (right) covered at 10×, 20× and 30× depth in tumour and germline samples. b, RNA-seq coverage. Coverage below the red line is considered poor quality; those with 20× depth, above the green line, are considered very high quality.
Extended Data Figure 2 Comparison of genomic aberrations among ependymomas analysed by whole genome sequencing.
a, The large majority of ependymomas have less than ten coding SNVs. Samples with more than 20 coding SNVs and their corresponding sample number from Fig. 1 are shown. b, Comparison of total number of structural variations in posterior fossa and supratentorial (ST) samples (Wilcox ranked sum test, P = 0.0006). c, Circos plot depicting SVs discovered across all supratentorial (red, outer plot) and posterior fossa (blue, inner plot) ependymomas. Each dot represents a validated or putative SV breakpoint detected by CREST in the WGS discovery cohort. Note the highly focal clustering of SVs on Chr11q in supratentorial ependymomas.
Extended Data Figure 3 Chromothripsis in supratentorial ependymomas resulting in C11orf95–RELA translocations.
Circos plots for the eight supratentorial (ST) ependymomas analysed using WGS that contained C11orf95–RELA translocations (sample numbers as Fig. 1). From the outer ring to the inner ring: chromosome, CNV calls, softclip count histogram, SVs (red, both sides with ≥10 softclips; blue, one side with >10 softclips; grey, <10 supporting softclips on either side).
Extended Data Figure 4 C11orf95–RELA translocations.
a, Breakpoints of structural rearrangements (red loops) at 11q13.1 in tumours ST1 to ST8. b, Exemplary C11orf95–RELA translocation and fusion transcript in sample ST5. Top, RNA-seq coverage; middle, DNA sequence across the fusion breakpoint; bottom, RNA sequence. c, Derivative chromosome generated by chromothripsis in tumour ST6 highlighting the locations of C11orf95 ‘break-apart’ FISH probes. Yellow block arrows represent chromosome fragments rearranged by chromothripsis. Numbers indicate fragment order on normal chromosome 11. FISH result, right. Arrows, transcription orientation.
Extended Data Figure 5 Translocation fusions: transcripts and predicted protein products detected in ependymoma.
a, Reverse transcription PCR products of the indicated transcripts detected in tumour samples (sample numbers as in Fig. 1a). b, Predicted protein products of wild-type translocation partners (top) and fusion products (bottom).
Extended Data Figure 6 Maps of fusion C11orf95–RELA products.
RNA-seq contig maps demonstrating the various fusion products generated through splicing of the primary C11orf95–RELA translocation transcript.
Extended Data Figure 7 Analysis of C11orf95–RELA translocation and expression in formalin-fixed paraffin-embedded human and mouse ependymoma.
a, Formalin-fixed paraffin-embedded (FFPE) cohort (top) and results (middle) of break-apart FISH and RT–PCR analysis of the C11orf95–RELA translocation and transcript, respectively. Tumours with ‘unavailable’ data had insufficient material for analysis. b, GFAP, p-Ser276-RELA, CCND1 and L1CAM immunohistochemistry in human and mouse RELAFUS1–7-positive and -negative cases. The mouse tumours recapitulate the ‘vascular-variant’ of human supratentorial ependymoma (see haematoxylin and eosin (H&E) images). This subtype is characterized by a branching network of capillaries (white arrows) and cytoplasmic clearing (black arrows). ***P < 0.0005. Scale bar, 50 μm
Extended Data Figure 8 C11orf95–RELA fusion protein spontaneously activates an NF-κB transcriptional reporter.
NF-κB–green fluorescence reporter (GFP) activity in 293T cells transduced with the indicated virus, treated for 60 min with TNF (50 ng ml–1) or vehicle control.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Materials, Supplementary Results, Supplementary Figures 1-15 additional references. (PDF 3496 kb)
Supplementary Tables
This file contains Supplementary Tables 1-6 and comprises Table 1: Clinical, pathological and validated genetic abnormalities of all 177 tumors in the study cohort ; Table 2: Coverage data for tumours analyzed by whole genome sequencing; Table 3: Coverage data for tumors analyzed by RNASeq; able 4: Tier 1 validated and putative high quality SNV and indel mutations; Table 5: Tier 2 validated and putative high quality SNV and indel mutations; and Table S6: Tier 3 validated and putative high-quality SNVs and indel mutations. (ZIP 954 kb)
Supplementary Tables
This file contains Supplementary Tables 7-12 and comprises Table 7: Numbers of high quality validated and putative Tier1 SNVs and indel mutations detected by WGS; Table 8: Copy number variations in tumors from the discovery series analyzed by WGS; Table 9: Copy number alterations detected by SNP 6 arrays; Table 10: Summary of All Genetic Aberrations Found by WGS; Table 11: Counts of structural variations detected by CREST in samples analyzed by WGS; and Table 12: Statistical analysis of chromothripsis status within 11q13.1. (ZIP 525 kb)
Supplementary Tables
This file contains Supplementary Tables 13-18 and comprises Table 13: Validated and uncovered fusion builder predictions from WGS; Table 14: Primers used for SV validation; Table 15: iFISH probes; Table 16a: C11orf95-RELA fusions predicted by RNA-seq; Table 16b: Fusions involving C11orf95 but not RELA; Table 16c: Additional fusions not involving C11orf95; Table 17a: Significantly activated transcription regulators identified by Ingenuity Pathway Analysis; Table 17b: Significant networks identified by analysis of differential gene expression data or RNASeq data & highlighting NF-kB; Table 18a: Pathway analysis of genes upregulated by RELAWT in NSCs and Table 18b: Pathway analysis of genes upregulated by RELAFUS1 in NSCs (ZIP 107 kb)
Rights and permissions
About this article
Cite this article
Parker, M., Mohankumar, K., Punchihewa, C. et al. C11orf95–RELA fusions drive oncogenic NF-κB signalling in ependymoma. Nature 506, 451–455 (2014). https://doi.org/10.1038/nature13109
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13109
This article is cited by
-
Distinct DNA methylation signatures associated with blood lipids as exposures or outcomes among survivors of childhood cancer: a report from the St. Jude lifetime cohort
Clinical Epigenetics (2023)
-
Nuclear condensates of YAP fusion proteins alter transcription to drive ependymoma tumourigenesis
Nature Cell Biology (2023)
-
Integrating network pharmacology and experimental validation to decipher the mechanism of action of Jingfang Granule in the treatment of viral myocarditis
Naunyn-Schmiedeberg's Archives of Pharmacology (2023)
-
A clinicopathological analysis of supratentorial ependymoma, ZFTA fusion-positive: utility of immunohistochemical detection of CDKN2A alterations and characteristics of the immune microenvironment
Brain Tumor Pathology (2023)
-
Etiology of oncogenic fusions in 5,190 childhood cancers and its clinical and therapeutic implication
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.