Abstract
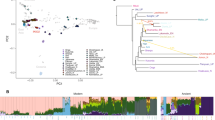
The origins of the First Americans remain contentious. Although Native Americans seem to be genetically most closely related to east Asians1,2,3, there is no consensus with regard to which specific Old World populations they are closest to4,5,6,7,8. Here we sequence the draft genome of an approximately 24,000-year-old individual (MA-1), from Mal’ta in south-central Siberia9, to an average depth of 1×. To our knowledge this is the oldest anatomically modern human genome reported to date. The MA-1 mitochondrial genome belongs to haplogroup U, which has also been found at high frequency among Upper Palaeolithic and Mesolithic European hunter-gatherers10,11,12, and the Y chromosome of MA-1 is basal to modern-day western Eurasians and near the root of most Native American lineages5. Similarly, we find autosomal evidence that MA-1 is basal to modern-day western Eurasians and genetically closely related to modern-day Native Americans, with no close affinity to east Asians. This suggests that populations related to contemporary western Eurasians had a more north-easterly distribution 24,000 years ago than commonly thought. Furthermore, we estimate that 14 to 38% of Native American ancestry may originate through gene flow from this ancient population. This is likely to have occurred after the divergence of Native American ancestors from east Asian ancestors, but before the diversification of Native American populations in the New World. Gene flow from the MA-1 lineage into Native American ancestors could explain why several crania from the First Americans have been reported as bearing morphological characteristics that do not resemble those of east Asians2,13. Sequencing of another south-central Siberian, Afontova Gora-2 dating to approximately 17,000 years ago14, revealed similar autosomal genetic signatures as MA-1, suggesting that the region was continuously occupied by humans throughout the Last Glacial Maximum. Our findings reveal that western Eurasian genetic signatures in modern-day Native Americans derive not only from post-Columbian admixture, as commonly thought, but also from a mixed ancestry of the First Americans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Accessions
Gene Expression Omnibus
Sequence Read Archive
Data deposits
Sequence data for MA-1 and AG-2, produced in this study, are available for download through NCBI SRA accession number SRP029640. Data from the Illumina genotyping analysis generated in this study are available through GEO Series accession number GSE50727; PLINK files can be accessed from http://www.ebc.ee/free_data. In addition, the above data and alignments for the published modern genomes, Denisova genome, Tianyuan individual and the two ancient genomes are available at http://www.cbs.dtu.dk/suppl/malta. Raw reads and alignments for the four modern genomes sequenced in this study are available for demographic research under data access agreement with E.W.
References
Turner, C. G. Advances in the dental search for native american origins. Acta Anthropogenet. 8, 23–78 (1984)
Hubbe, M., Harvati, K. & Neves, W. Paleoamerican morphology in the context of European and East Asian Pleistocene variation: implications for human dispersion into the New World. Am. J. Phys. Anthropol. 144, 442–453 (2011)
Schurr, T. The peopling of the New World: perspectives from molecular anthropology. Annu. Rev. Anthropol. 33, 551–583 (2004)
O’Rourke, D. H. & Raff, J. A. The human genetic history of the Americas: the final frontier. Curr. Biol. 20, R202–R207 (2010)
Lell, J. T. et al. The dual origin and siberian affinities of native american Y chromosomes. Am. J. Hum. Genet. 70, 192–206 (2002)
Starikovskaya, E. B. et al. Mitochondrial DNA diversity in indigenous populations of the southern extent of Siberia, and the origins of Native American haplogroups. Ann. Hum. Genet. 69, 67–89 (2005)
Dulik, M. C. et al. Mitochondrial DNA and Y chromosome variation provides evidence for a recent common ancestry between Native American and Indigenous Altaians. Am. J. Hum. Genet. 90, 229–246 (2012)
Regueiro, M., Alvarez, J., Rowold, D. & Herrera, R. J. On the origins, rapid expansion and genetic diversity of Native Americans from hunting-gatherers to agriculturalists. Am. J. Phys. Anthropol. 150, 333–348 (2013)
Gerasimov, M. M. in The Archaeology and Geomorphology of Northern Asia: Selected Works 5–32 (University of Toronto Press, 1964)
Bramanti, B. et al. Genetic discontinuity between local hunter-gatherers and central Europe’s first farmers. Science 326, 137–140 (2009)
Malmström, H. et al. Ancient DNA reveals lack of continuity between Neolithic hunter-gatherers and contemporary Scandinavians. Curr. Biol. 19, 1758–1762 (2009)
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013)
Owsley, D. W. & Jantz, R. L. in Claiming the Stones-Naming the Bones: Cultural Property and the Negotiation of National and Ethnic Identity (Getty Research Institute, 2002)
Astakhov, S. N. Paleolit Eniseia: Paleoliticheskie Stoianki Afontovoi Gore v G. Krasnoiarske (Evropaiskii Dom, 1999)
Gamble, C. Interaction and alliance in Palaeolithic society. Man (Lond) 17, 92–107 (1982)
Abramova, Z. L’art Paléolithique d’Europe Orientale et de Sibérie (Jérôme Millon, 1995)
White, R. The women of Brassempouy: a century of research and interpretation. J. Archaeol. Method and Theory 13, 250–303 (2006)
Hansen, A. J., Willerslev, E., Wiuf, C., Mourier, T. & Arctander, P. Statistical evidence for miscoding lesions in ancient DNA templates. Mol. Biol. Evol. 18, 262–265 (2001)
Reich, D. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012)
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012)
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012)
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012)
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010)
Gutenkunst, R. N., Hernandez, R. D., Williamson, S. H. & Bustamante, C. D. Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLoS Genet. 5, e1000695 (2009)
Wall, J. D. et al. Genetic variation in Native Americans, inferred from latino SNP and resequencing data. Mol. Biol. Evol. 28, 2231–2237 (2011)
Fu, Q. et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc. Natl Acad. Sci. USA 110, 2223–2227 (2013)
Lipson, M. et al. Efficient moment-based inference of admixture parameters and sources of gene flow. Mol. Biol. Evol. (2013)
Goebel, T. Pleistocene human colonization of siberia and peopling of the Americas: an ecological approach. Evol. Anthropol. 8, 208–227 (1999)
Brown, M. D. et al. mtDNA haplogroup X: an ancient link between Europe/Western Asia and North America? Am. J. Hum. Genet. 63, 1852–1861 (1998)
Bradley, B. & Stanford, D. The North Atlantic ice-edge corridor: a possible Palaeolithic route to the New World. World Archaeol. 36, 459–478 (2004)
Stafford, T. W., Jr, Jull, A. J. T., Brendel, K., Duhamel, R. & Donahue, D. Study of bone radiocarbon dating accuracy at the University of Arizona NSF accelerator facility for radioisotope analysis. Radiocarbon 29, 24–44 (1987)
Stafford, T. W., Jr, Brendel, K. & Duhamel, R. Radiocarbon, 13C and 15N analysis of fossil bone: removal of humates with XAD-2 resin. Geochim. Cosmochim. Acta 52, 2257–2267 (1988)
Stafford, T. W., Jr, Hare, P. E., Currie, L., Jull, A. J. T. & Donahue, D. Accelerator radiocarbon dating at the molecular level. J. Archaeol. Sci. 18, 35–72 (1991)
Ramsey, C. B. Bayesian analysis of radiocarbon dates. Radiocarbon 51, 337–360 (2009)
Reimer, P. J. et al. IntCal09 and Marine09 radiocarbon age calibration curves, 0-50,000 years cal BP . Radiocarbon 51, 1111–1150 (2009)
Yang, D. Y., Eng, B., Waye, J. S., Dudar, J. C. & Sanders, S. R. Technical note: improved DNA extraction from ancient bones using silica-based spin columns. Am. J. Phys. Anthropol. 105, 539–543 (1998)
Svensson, E. M. et al. Tracing genetic change over time using nuclear SNPs in ancient and modern cattle. Anim. Genet. 38, 378–383 (2007)
Powell, R. & Gannon, F. Purification of DNA by phenol extraction and ethanol precipitation. Oxford Practical Approach Series. http://fds.oup.com/www.oup.co.uk/pdf/pas/9v1-7-3.pdf (2002)
Orlando, L. et al. Recalibrating Equus evolution using the genome sequence of an early Middle Pleistocene horse. Nature 499, 74–78 (2013)
Reich, D. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010)
Lindgreen, S. AdapterRemoval: easy cleaning of next-generation sequencing reads. BMC Res. Notes 5, 337 (2012)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Schubert, M. et al. Improving ancient DNA read mapping against modern reference genomes. BMC Genomics 13, 178 (2012)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
Krause, J. et al. A complete mtDNA genome of an early modern human from Kostenki, Russia. Curr. Biol. 20, 231–236 (2010)
Skoglund, P., Storå, J., Götherström, A. & Jakobsson, M. Accurate sex identification in ancient human remains using DNA shotgun sequencing. J. Archaeol. Sci. 40, 4477–4482 (2013)
Rasmussen, M. et al. An Aboriginal Australian genome reveals separate human dispersals in Asia. Science 334, 94–98 (2011)
Frazer, K. A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
The 1000 Genomes Project Consortium An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012)
Van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–E394 (2009)
Behar, D. M. et al. A “Copernican” reassessment of the human mitochondrial DNA tree from its root. Am. J. Hum. Genet. 90, 675–684 (2012)
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010)
Wei, W. et al. A calibrated human Y-chromosomal phylogeny based on resequencing. Genome Res. 23, 388–395 (2013)
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011)
Hancock, A. M. et al. Adaptations to climate-mediated selective pressures in humans. PLoS Genet. 7, e1001375 (2011)
Rasmussen, M. et al. Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature 463, 757–762 (2010)
International HapMap 3 Consortium Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010)
Li, J. Z. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008)
Skoglund, P. & Jakobsson, M. Archaic human ancestry in East Asia. Proc. Natl Acad. Sci. USA 108, 18301–18306 (2011)
Patterson, N., Price, A. L. & Reich, D. Population structure and Eigenanalysis. PLoS Genet. 2, e190 (2006)
Skoglund, P. et al. Origins and Genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012)
Surakka, I. et al. Founder population-specific HapMap panel increases power in GWA studies through improved imputation accuracy and CNV tagging. Genome Res. 20, 1344–1351 (2010)
International HapMap3 Consortium Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010)
Busing, F. M. T. A., Meijer, E. & Van der Leeden, R. Delete-m Jackknife for Unequal m. Stat. Comput. 9, 3–8 (1999)
Durand, E. Y., Patterson, N., Reich, D. & Slatkin, M. Testing for ancient admixture between closely related populations. Mol. Biol. Evol. 28, 2239–2252 (2011)
Acknowledgements
We thank the Hermitage State Museum for providing access to the Mal’ta and Afontova Gora-2 human remains. We also thank the Danish National High-Throughput DNA Sequencing Centre and T. Reisberg for technical assistance. This work was supported by the Danish National Research Foundation and the Lundbeck Foundation (E.W. and M.R.) and the Arctic Social Sciences Program, National Science Foundation (grant PLR-1003725 to K.E.G.). R.V., M.M., M.K., E.M., K.T., S.Ro. and R.M. were supported by the European Regional Development Fund (European Union) through the Centre of Excellence in Genomics to Estonian Biocentre and University of Tartu and Estonian Basic Research grant SF0270177As08. M.M. thanks the Estonian Science Foundation grant no. 8973 and Baltic-American Freedom Foundation Research Scholarship program and M.I.V. thanks the Government of Russian Federation grant no. 14.B25.31.0033 (to E. I. Rogaev). M.D. was supported by the US National Science Foundation (grant DBI-1103639). Computational analyses were carried out at the High Performance Computing Center, University of Tartu, and the Swedish National Infrastructure for Computing (SNIC-UPPMAX, project b2012063).
Author information
Authors and Affiliations
Contributions
E.W. and K.E.G. conceived the project. E.W. headed the project. E.W. and M.R. designed the experimental research project setup. S.D. and K.E.G. provided access to the Mal’ta and Afontova Gora-2 samples, and K.E.G. provided archaeological context for the samples. T.W.S. Jr performed AMS dating. E.B. and O.B. (Tajik individual), E.K. and S.L. (Mari and Avar individuals) provided modern DNA extracts for complete genome sequencing. E.K. and S.L. (Kazakh, Kirghiz, Uzbek and Mari individuals), L.P.O. (Selkup individuals), S.A.F. (Even, Dolgan and Yakut individuals) and M.I.V. (Altai individuals) provided access to modern DNA extracts for genotyping. R.V. carried out Illumina chip analysis on modern samples. P.F.C. performed DNA extraction from the Indian individual. M.R. performed the ancient extractions and library constructions on the modern and ancient samples —the latter with input from L.O. M.R. coordinated the sequencing. M.R. and S.Ra. performed mapping of MA-1 and AG-2 data sets with input from L.O. S.Ra., T.S.-P. and S.B. provided super-computing resources, developed the next-generation sequencing pipeline and performed mapping and genotyping for all the modern genomes. M.R. performed DNA damage analysis with input from L.O. M.M. performed the admixture analysis. M.M., E.M., K.T. and R.V. performed the mtDNA analysis. M.M., M.K., S.Ro., T.K., R.V. and R.M. performed the Y-chromosome analysis. A.A. and I.M. performed the autosomal contamination estimates, error rate estimates, D-statistics tests based on sequence reads and ngsAdmix analyses. P.S. performed biological sexing, mtDNA contamination estimates, PCA, TreeMix, MixMapper, D-statistic tests based on allele frequencies, f3-statistics and phenotypic analyses, and analysis of AG-2 using nucleotide misincorporation patterns under the supervision of R.N. and M.J. M.R., P.S. and E.W. wrote the majority of the manuscript with critical input from R.N., M.J., M.M., K.E.G., A.A., I.M. and M.D. M.M., A.A. and I.M. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary Figures, Supplementary Tables and additional references – see contents page for details. (PDF 13232 kb)
Rights and permissions
About this article
Cite this article
Raghavan, M., Skoglund, P., Graf, K. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014). https://doi.org/10.1038/nature12736
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12736
This article is cited by
-
The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes
Scientific Data (2024)
-
Population genomics of post-glacial western Eurasia
Nature (2024)
-
Exploring regional aspects of 3D facial variation within European individuals
Scientific Reports (2023)
-
Ancient human DNA recovered from a Palaeolithic pendant
Nature (2023)
-
Ancient woman’s DNA recovered from a 20,000-year-old pendant
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.