Abstract
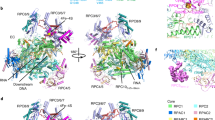
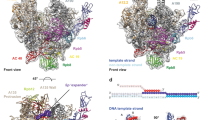
Transcription of ribosomal RNA by RNA polymerase (Pol) I initiates ribosome biogenesis and regulates eukaryotic cell growth. The crystal structure of Pol I from the yeast Saccharomyces cerevisiae at 2.8 Å resolution reveals all 14 subunits of the 590-kilodalton enzyme, and shows differences to Pol II. An ‘expander’ element occupies the DNA template site and stabilizes an expanded active centre cleft with an unwound bridge helix. A ‘connector’ element invades the cleft of an adjacent polymerase and stabilizes an inactive polymerase dimer. The connector and expander must detach during Pol I activation to enable transcription initiation and cleft contraction by convergent movement of the polymerase ‘core’ and ‘shelf’ modules. Conversion between an inactive expanded and an active contracted polymerase state may generally underlie transcription. Regulatory factors can modulate the core–shelf interface that includes a ‘composite’ active site for RNA chain initiation, elongation, proofreading and termination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Roeder, R. G. & Rutter, W. J. Multiple forms of DNA-dependent RNA polymerase in eukaryotic organisms. Nature 224, 234–237 (1969)
Miller, O. L., Jr & Beatty, B. R. Visualization of nucleolar genes. Science 164, 955–957 (1969)
Laferté, A. et al. The transcriptional activity of RNA polymerase I is a key determinant for the level of all ribosome components. Genes Dev. 20, 2030–2040 (2006)
Drygin, D., Rice, W. G. & Grummt, I. The RNA polymerase I transcription machinery: an emerging target for the treatment of cancer. Annu. Rev. Pharmacol. Toxicol. 50, 131–156 (2010)
Schultz, P., Celia, H., Riva, M., Sentenac, A. & Oudet, P. Three-dimensional model of yeast RNA polymerase I determined by electron microscopy of two-dimensional crystals. EMBO J. 12, 2601–2607 (1993)
Bischler, N. et al. Localization of the yeast RNA polymerase I-specific subunits. EMBO J. 21, 4136–4144 (2002)
Cramer, P., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: RNA polymerase II at 2.8 angstrom resolution. Science 292, 1863–1876 (2001)
Cheung, A. C. & Cramer, P. A movie of RNA polymerase II transcription. Cell 149, 1431–1437 (2012)
Kuhn, C. D. et al. Functional architecture of RNA polymerase I. Cell 131, 1260–1272 (2007)
Geiger, S. R. et al. RNA polymerase I contains a TFIIF-related DNA-binding subcomplex. Mol. Cell 39, 583–594 (2010)
Jennebach, S., Herzog, F., Aebersold, R. & Cramer, P. Crosslinking-MS analysis reveals RNA polymerase I domain architecture and basis of rRNA cleavage. Nucleic Acids Res. 40, 5591–5601 (2012)
Korkhin, Y. et al. Evolution of complex RNA polymerases: the complete archaeal RNA polymerase structure. PLoS Biol. 7, e1000102 (2009)
Hirata, A. et al. Archaeal RNA polymerase subunits E and F are not required for transcription in vitro, but a Thermococcus kodakarensis mutant lacking subunit F is temperature-sensitive. Mol. Microbiol. 70, 623–633 (2008)
Kettenberger, H., Armache, K. J. & Cramer, P. Architecture of the RNA polymerase II–TFIIS complex and implications for mRNA cleavage. Cell 114, 347–357 (2003)
Buhler, J. M., Sentenac, A. & Fromageot, P. Isolation, structure, and general properties of yeast ribonucleic acid polymerase A (or I). J. Biol. Chem. 249, 5963–5970 (1974)
Ruan, W., Lehmann, E., Thomm, M., Kostrewa, D. & Cramer, P. Evolution of two modes of intrinsic RNA polymerase transcript cleavage. J. Biol. Chem. 286, 18701–18707 (2011)
Alic, N. et al. Selectivity and proofreading both contribute significantly to the fidelity of RNA polymerase III transcription. Proc. Natl Acad. Sci. USA 104, 10400–10405 (2007)
Prescott, E. M. et al. Transcriptional termination by RNA polymerase I requires the small subunit Rpa12p. Proc. Natl Acad. Sci. USA 101, 6068–6073 (2004)
Arimbasseri, A. G., Rijal, K. & Maraia, R. J. Transcription termination by the eukaryotic RNA polymerase III. Biochim. Biophys. Acta 1829, 318–330 (2013)
Van Mullem, V., Landrieux, E., Vandenhaute, J. & Thuriaux, P. Rpa12p, a conserved RNA polymerase I subunit with two functional domains. Mol. Microbiol. 43, 1105–1113 (2002)
Beckouet, F. et al. Two RNA polymerase I subunits control the binding and release of Rrn3 during transcription. Mol. Cell. Biol. 28, 1596–1605 (2008)
Chen, Z. A. et al. Architecture of the RNA polymerase II–TFIIF complex revealed by cross-linking and mass spectrometry. EMBO J. 29, 717–726 (2010)
Eichner, J., Chen, H. T., Warfield, L. & Hahn, S. Position of the general transcription factor TFIIF within the RNA polymerase II transcription preinitiation complex. EMBO J. 29, 706–716 (2010)
Fernández-Tornero, C. et al. Conformational flexibility of RNA polymerase III during transcriptional elongation. EMBO J. 29, 3762–3772 (2010)
Tagami, S. et al. Crystal structure of bacterial RNA polymerase bound with a transcription inhibitor protein. Nature 468, 978–982 (2010)
Weixlbaumer, A., Leon, K., Landick, R. & Darst, S. A. Structural basis of transcriptional pausing in bacteria. Cell 152, 431–441 (2013)
Cheung, A. C., Sainsbury, S. & Cramer, P. Structural basis of initial RNA polymerase II transcription. EMBO J. 30, 4755–4763 (2011)
Wang, D., Bushnell, D. A., Westover, K. D., Kaplan, C. D. & Kornberg, R. D. Structural basis of transcription: role of the trigger loop in substrate specificity and catalysis. Cell 127, 941–954 (2006)
Peyroche, G. et al. The recruitment of RNA polymerase I on rDNA is mediated by the interaction of the A43 subunit with Rrn3. EMBO J. 19, 5473–5482 (2000)
Armache, K. J., Mitterweger, S., Meinhart, A. & Cramer, P. Structures of complete RNA polymerase II and its subcomplex, Rpb4/7. J. Biol. Chem. 280, 7131–7134 (2005)
Jasiak, A. J., Armache, K. J., Martens, B., Jansen, R. P. & Cramer, P. Structural biology of RNA polymerase III: subcomplex C17/25 X-ray structure and 11 subunit enzyme model. Mol. Cell 23, 71–81 (2006)
Smid, A., Riva, M., Bouet, F., Sentenac, A. & Carles, C. The association of three subunits with yeast RNA polymerase is stabilized by A14. J. Biol. Chem. 270, 13534–13540 (1995)
Beckouët, F., Mariotte-Labarre, S., Peyroche, G., Nogi, Y. & Thuriaux, P. Rpa43 and its partners in the yeast RNA polymerase I transcription complex. FEBS Lett. 585, 3355–3359 (2011)
Blattner, C. et al. Molecular basis of Rrn3-regulated RNA polymerase I initiation and cell growth. Genes Dev. 25, 2093–2105 (2011)
Knutson, B. A. & Hahn, S. Yeast Rrn7 and human TAF1B are TFIIB-related RNA polymerase I general transcription factors. Science 333, 1637–1640 (2011)
Naidu, S., Friedrich, J. K., Russell, J. & Zomerdijk, J. C. TAF1B is a TFIIB-like component of the basal transcription machinery for RNA polymerase I. Science 333, 1640–1642 (2011)
Milkereit, P., Schultz, P. & Tschochner, H. Resolution of RNA polymerase I into dimers and monomers and their function in transcription. Biol. Chem. 378, 1433–1443 (1997)
Milkereit, P. & Tschochner, H. A specialized form of RNA polymerase I, essential for initiation and growth-dependent regulation of rRNA synthesis, is disrupted during transcription. EMBO J. 17, 3692–3703 (1998)
Fath, S. et al. Differential roles of phosphorylation in the formation of transcriptional active RNA polymerase I. Proc. Natl Acad. Sci. USA 98, 14334–14339 (2001)
Gerber, J. et al. Site specific phosphorylation of yeast RNA polymerase I. Nucleic Acids Res. 36, 793–802 (2008)
Miller, G. et al. hRRN3 is essential in the SL1-mediated recruitment of RNA Polymerase I to rRNA gene promoters. EMBO J. 20, 1373–1382 (2001)
Yuan, X., Zhao, J., Zentgraf, H., Hoffmann-Rohrer, U. & Grummt, I. Multiple interactions between RNA polymerase I, TIF-IA and TAFI subunits regulate preinitiation complex assembly at the ribosomal gene promoter. EMBO Rep. 3, 1082–1087 (2002)
Clemente-Blanco, A. et al. Cdc14 inhibits transcription by RNA polymerase I during anaphase. Nature 458, 219–222 (2009)
Cheung, A. C. & Cramer, P. Structural basis of RNA polymerase II backtracking, arrest and reactivation. Nature 471, 249–253 (2011)
Epshtein, V., Cardinale, C. J., Ruckenstein, A. E., Borukhov, S. & Nudler, E. An allosteric path to transcription termination. Mol. Cell 28, 991–1001 (2007)
Nielsen, S., Yuzenkova, Y. & Zenkin, N. Mechanism of eukaryotic RNA polymerase III transcription termination. Science 340, 1577–1580 (2013)
Sainsbury, S., Niesser, J. & Cramer, P. Structure and function of the initially transcribing RNA polymerase II–TFIIB complex. Nature 493, 437–440 (2013)
Soutourina, J., Wydau, S., Ambroise, Y., Boschiero, C. & Werner, M. Direct interaction of RNA polymerase II and mediator required for transcription in vivo . Science 331, 1451–1454 (2011)
Lawson, C. L. et al. Catabolite activator protein: DNA binding and transcription activation. Curr. Opin. Struct. Biol. 14, 10–20 (2004)
Zuo, Y., Wang, Y. & Steitz, T. A. The mechanism of E. coli RNA polymerase regulation by ppGpp is suggested by the structure of their complex. Mol. Cell 50, 430–436 (2013)
Tschochner, H. A novel RNA polymerase I-dependent RNase activity that shortens nascent transcripts from the 3′ end. Proc. Natl Acad. Sci. USA 93, 12914–12919 (1996)
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010)
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007)
Cowtan, K. Recent developments in classical density modification. Acta Crystallogr. D 66, 470–478 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983)
Bischler, N. et al. Specific interaction and two-dimensional crystallization of histidine tagged yeast RNA polymerase I on nickel-chelating lipids. Biophys. J. 74, 1522–1532 (1998)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
Geiger, S. R., Kuhn, C. D., Leidig, C., Renkawitz, J. & Cramer, P. Crystallization of RNA polymerase I subcomplex A14/A43 by iterative prediction, probing and removal of flexible regions. Acta Crystallogr. F 64, 413–418 (2008)
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Acknowledgements
We thank C. Kuhn, who obtained the first Pol I crystals. We thank C. Bäjen, S. Benkert, S. Geiger, S. Jennebach, T. Gubbey and K. Maier. We thank the crystallization facility (Conti Department) and the Jentsch Department of the Max-Planck-Institut for Biochemistry. Part of this work was performed at the European Synchrotron Radiation Facility at Grenoble, France, and at the Swiss Light Source at the Paul-Scherrer-Institut, Villigen, Switzerland. C.E. was supported by a PhD student fellowship of the Boehringer Ingelheim Fonds, the Elite Network Bavaria program ‘Protein Dynamics in Health and Disease’, and the Graduate Research Academy ‘RNA Biology’ of SFB960. S.S. was supported by a postdoctoral fellowship of the Alexander-von-Humboldt Foundation. P.C. was supported by the Deutsche Forschungsgemeinschaft (SFB646, TR5, GraKo1721, SFB960, CIPSM, NIM), an Advanced Grant of the European Research Council, the Jung-Stiftung, and the Vallee Foundation.
Author information
Authors and Affiliations
Contributions
C.E. planned and carried out experiments and crystal structure determination. S.S. advised on experimental and crystallographic work. A.C.C. performed computational crystallographic analysis. D.K. contributed to computational crystallography and model building. P.C. designed and supervised research and prepared the manuscript, with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Activity of purified Pol I in vitro.
a, Pol I is active in RNA extension and cleavage. Purified S. cerevisiae Pol I and Pol II can extend RNA in a DNA–RNA scaffold in the presence of nucleoside triphosphate substrates (NTPs). In the absence of nucleoside triphosphate substrates, Pol I cleaves RNA. Transcription assays were performed as described9 using the DNA–RNA scaffold shown. After running a 20% acrylamide UREA gel, RNA was detected by fluorescence. b, Dynamic light scattering is consistent with a concentration-dependent Pol I dimer–monomer equilibrium in solution. Note that the sample with the higher Pol I concentration shows an increased hydrodynamic radius and apparent (estimated) molecular mass for the predominant peak by mass that accounts for over 96%. The method cannot distinguish monomeric and dimeric species. Thus, the estimated molecular mass observed at 3.5 mg ml−1 Pol I concentration (769 kDa) arises from a mixture of monomers (590 kDa) and dimers (1,180 kDa). c, De novo RNA synthesis activity on tailed DNA template. Assays were performed as described47 using the DNA template shown. After running a 20% acrylamide UREA gel, de novo synthesized, radioactive RNA was detected by phosphoimaging.
Extended Data Figure 2 Structure-based alignment of A190 and Rpb1.
Invariant and conserved residues are highlighted in green and light green, respectively. Secondary structure elements are indicated above and below the alignment for A190 and Rpb1, respectively (cylinders, helices; arrows, strands). Residues that form different folds in Pol I or form Pol-II-specific folds are in green or red, respectively (compare Fig. 1c).
Extended Data Figure 3 Structure-based alignment of A135 and Rpb2.
Invariant and conserved residues are highlighted in green and light green, respectively. Secondary structure elements are indicated above and below the alignment for A135 and Rpb2, respectively (cylinders, helices; arrows, strands). Residues that form different folds in Pol I or form Pol-II-specific folds are in green or red, respectively (compare Fig. 1c).
Extended Data Figure 4 Detailed comparison of A190–A135 with Rpb1–Rpb2.
a, Comparison of A190 domain structures (top) that differ significantly from their corresponding Pol II regions (bottom). Labelling of corresponding secondary structure elements is as for Pol II (ref. 7). New or lacking secondary structure elements are labelled. New elements were named according to the preceding Pol II element with small letters added alphabetically for subsequent elements. b, Comparison of A135 domain structures (top) that differ significantly from their corresponding Pol II regions (bottom) as in Extended Data Fig. 3.
Extended Data Figure 5 Structure of the subassembly AC40–AC19–Rpb10–Rpb12.
a, Schematic of domain structures. AC40 and AC19 were aligned with homologous Pol II subunits Rpb3 and Rpb11, respectively. Organization and annotation as in Fig. 2a. b, Ribbon view from the ‘back’ of the enzyme7 (left, Pol I; right, Pol II). c, d, Structure-based alignments of AC40 and Rpb3 (c) and of AC19 and Rpb11 (d). Invariant and conserved residues are highlighted in green and light green, respectively. Secondary structure elements are indicated above and below the alignment for AC40 and Rpb3, respectively (cylinders, helices; arrows, strands). Residues that form different folds in Pol I or form Pol-II-specific folds are in green or red, respectively (compare Fig. 1c).
Extended Data Figure 6 A12.2 and A49–A34.5.
a, Subunit A12.2 amino acid sequence alignment. The sequence of yeast S. cerevisiae A12.2 (Sc) was aligned with that of the human subunit (Homo sapiens; Hs), and the N- and C-terminal domains were aligned with their counterparts in the Pol II subunit Rpb9 and the Pol II elongation factor TFIIS14, respectively. b, Interaction of A49–A34.5 with Pol I core domains. The view is from the front of the enzyme7 (Fig. 1). Different Pol I subunit domains are coloured as in Figs 2 and 3.
Extended Data Figure 7 Shift in domain positions between Pol I and Pol II.
Individual domain fragments of Pol I were isolated and superposed onto their counterparts in Pol II and subsequently recombined to form a ‘pseudo-Pol II’ model. This pseudo-Pol II model was then superposed onto the complete Pol I structure over a single common domain (indicated by column headings). The resulting root mean squared deviation (r.m.s.d.) Cα (Å) value for every individual domain (indicated by row headings) was calculated using PyMol and coloured according to value (green to yellow = 0.0 to 3.0 Å, orange = 3.0 to 4.0 Å, and red >4.0 Å).
Extended Data Figure 8 Structure and conservation of the connector and expander.
a, The connector binds the adjacent polymerase. The electron density is not continuous between the end of the A43 N-terminal part (residue Ile 250) and the beginning of the A43 connector in the adjacent polymerase. The connector cannot be assigned to A43 from the same polymerase because the observed distance between the last residue in the stalk and the first residue in the connector (>72 Å) cannot be spanned with only 12 residues. By contrast, the distance from Ile 250 to the connector in the cleft of the neighbouring polymerase is 18 Å and easily spanned by the missing residues. We note that dimeric forms of Pol I were previously observed in two-dimensional arrays59. b, Amino acid sequence alignment of the connector region in S. cerevisiae (Sc); Candida glabrata (Cg); Homo sapiens (Hs) and Mus musculus (Mm). Secondary structure elements are indicated above the alignment (K5 helix, cylinder; strands D1, D2; arrows). Residues that are involved in the interface with the Pol I clamp and cleft and showed a buried surface area in excess of 40 Å2 are indicated with an asterisk above the alignment. Buried surfaces were calculated with the PISA server60. A structure-based alignment of the A14–A43 stalk residues was published9,61 and is not included here. c, Amino acid sequence alignment of the expander region in S. cerevisiae, C. glabrata, H. sapiens and M. musculus. Secondary structure elements are indicated above the alignment (helices, cylinders; strands, arrows). Residues that are involved in the interface with the Pol I cleft and showed a buried surface area in excess of 40 Å2 are indicated with an asterisk above the alignment. Buried surfaces were calculated with the PISA server60. d, Previously obtained crosslinks11 of the expander element map to the Pol I cleft. Crosslinks to A190 and A135 are indicated in grey and wheat respectively.
Supplementary information
Conformational differences between Pol I and Pol II illustrated by polymerase module movement
The video illustrates the differences in conformation between RNA polymerases I and II, and begins with a top view of Pol II. The core, shelf, clamp, stalk and lobe modules are coloured grey, magenta, orange, yellow and blue respectively. The structure morphs from a Pol II to Pol I conformation, highlighting their differences in cleft width. The Pol I specific subunits are then shown briefly before being faded out and the structure morphs back to the Pol II conformation. The video was generated using UCSF chimera. (MOV 13395 kb)
Rights and permissions
About this article
Cite this article
Engel, C., Sainsbury, S., Cheung, A. et al. RNA polymerase I structure and transcription regulation. Nature 502, 650–655 (2013). https://doi.org/10.1038/nature12712
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12712
This article is cited by
-
Structures of transcription preinitiation complex engaged with the +1 nucleosome
Nature Structural & Molecular Biology (2023)
-
Structural insights into nuclear transcription by eukaryotic DNA-dependent RNA polymerases
Nature Reviews Molecular Cell Biology (2022)
-
RNA-Polymerase I: einer spezialisierten Transkriptionsmaschine auf der Spur
BIOspektrum (2022)
-
Conserved strategies of RNA polymerase I hibernation and activation
Nature Communications (2021)
-
Structure and function of virion RNA polymerase of a crAss-like phage
Nature (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.