Abstract
The recognition of autophagy related 16-like 1 (ATG16L1) as a genetic risk factor has exposed the critical role of autophagy in Crohn’s disease1. Homozygosity for the highly prevalent ATG16L1 risk allele, or murine hypomorphic (HM) activity, causes Paneth cell dysfunction2,3. As Atg16l1HM mice do not develop spontaneous intestinal inflammation, the mechanism(s) by which ATG16L1 contributes to disease remains obscure. Deletion of the unfolded protein response (UPR) transcription factor X-box binding protein-1 (Xbp1) in intestinal epithelial cells, the human orthologue of which harbours rare inflammatory bowel disease risk variants, results in endoplasmic reticulum (ER) stress, Paneth cell impairment and spontaneous enteritis4. Unresolved ER stress is a common feature of inflammatory bowel disease epithelium4,5, and several genetic risk factors of Crohn’s disease affect Paneth cells2,4,6,7,8,9. Here we show that impairment in either UPR (Xbp1ΔIEC) or autophagy function (Atg16l1ΔIEC or Atg7ΔIEC) in intestinal epithelial cells results in each other’s compensatory engagement, and severe spontaneous Crohn’s-disease-like transmural ileitis if both mechanisms are compromised. Xbp1ΔIEC mice show autophagosome formation in hypomorphic Paneth cells, which is linked to ER stress via protein kinase RNA-like endoplasmic reticulum kinase (PERK), elongation initiation factor 2α (eIF2α) and activating transcription factor 4 (ATF4). Ileitis is dependent on commensal microbiota and derives from increased intestinal epithelial cell death, inositol requiring enzyme 1α (IRE1α)-regulated NF-κB activation and tumour-necrosis factor signalling, which are synergistically increased when autophagy is deficient. ATG16L1 restrains IRE1α activity, and augmentation of autophagy in intestinal epithelial cells ameliorates ER stress-induced intestinal inflammation and eases NF-κB overactivation and intestinal epithelial cell death. ER stress, autophagy induction and spontaneous ileitis emerge from Paneth-cell-specific deletion of Xbp1. Genetically and environmentally controlled UPR function within Paneth cells may therefore set the threshold for the development of intestinal inflammation upon hypomorphic ATG16L1 function and implicate ileal Crohn’s disease as a specific disorder of Paneth cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
13 November 2013
Figure 4a was corrected.
References
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012)
Cadwell, K. et al. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 456, 259–263 (2008)
Cadwell, K. et al. Virus-plus-susceptibility gene interaction determines Crohn’s disease gene Atg16L1 phenotypes in intestine. Cell 141, 1135–1145 (2010)
Kaser, A. et al. XBP1 links ER stress to intestinal inflammation and confers genetic risk for human inflammatory bowel disease. Cell 134, 743–756 (2008)
Tréton, X. et al. Altered endoplasmic reticulum stress affects translation in inactive colon tissue from patients with ulcerative colitis. Gastroenterology 141, 1024–1035 (2011)
Kobayashi, K. S. et al. Nod2-dependent regulation of innate and adaptive immunity in the intestinal tract. Science 307, 731–734 (2005)
Zhao, F. et al. Disruption of Paneth and goblet cell homeostasis and increased endoplasmic reticulum stress in Agr2–/– mice. Dev. Biol. 338, 270–279 (2010)
Wehkamp, J. et al. Reduced Paneth cell α-defensins in ileal Crohn’s disease. Proc. Natl Acad. Sci. USA 102, 18129–18134 (2005)
Maloy, K. J. & Powrie, F. Intestinal homeostasis and its breakdown in inflammatory bowel disease. Nature 474, 298–306 (2011)
Levine, B., Mizushima, N. & Virgin, H. W. Autophagy in immunity and inflammation. Nature 469, 323–335 (2011)
Sarkar, S., Ravikumar, B. & Rubinsztein, D. C. Autophagic clearance of aggregate-prone proteins associated with neurodegeneration. Methods Enzymol. 453, 83–110 (2009)
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15, 1101–1111 (2004)
Rouschop, K. M. et al. The unfolded protein response protects human tumor cells during hypoxia through regulation of the autophagy genes MAP1LC3B and ATG5. J. Clin. Invest. 120, 127–141 (2010)
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005)
Boyce, M. et al. A selective inhibitor of eIF2α dephosphorylation protects cells from ER stress. Science 307, 935–939 (2005)
Novoa, I. et al. Stress-induced gene expression requires programmed recovery from translational repression. EMBO J. 22, 1180–1187 (2003)
Hampe, J. et al. A genome-wide association scan of nonsynonymous SNPs identifies a susceptibility variant for Crohn disease in ATG16L1. Nature Genet. 39, 207–211 (2007)
Cleynen, I. et al. Genetic factors conferring an increased susceptibility to develop Crohn's disease also influence disease phenotype: results from the IBDchip European Project. Gut http://dx.doi.org/10.1136/gutjnl-2011-300777 (2012)
Fujita, N. et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell 19, 2092–2100 (2008)
Mizushima, N. et al. Mouse Apg16L, a novel WD-repeat protein, targets to the autophagic isolation membrane with the Apg12-Apg5 conjugate. J. Cell Sci. 116, 1679–1688 (2003)
Fujita, N. et al. Differential involvement of Atg16L1 in Crohn disease and canonical autophagy: analysis of the organization of the Atg16L1 complex in fibroblasts. J. Biol. Chem. 284, 32602–32609 (2009)
Yang, L., Li, P., Fu, S., Calay, E. S. & Hotamisligil, G. S. Defective hepatic autophagy in obesity promotes ER stress and causes insulin resistance. Cell Metab. 11, 467–478 (2010)
Bertolotti, A. et al. Increased sensitivity to dextran sodium sulfate colitis in IRE1β-deficient mice. J. Clin. Invest. 107, 585–593 (2001)
Cao, S. S. et al. The unfolded protein response and chemical chaperones reduce protein misfolding and colitis in mice. Gastroenterology 144, 989–1000 (2013)
Kaser, A., Zeissig, S. & Blumberg, R. S. Inflammatory bowel disease. Annu. Rev. Immunol. 28, 573–621 (2010)
Rogler, G. et al. Nuclear factor κB is activated in macrophages and epithelial cells of inflamed intestinal mucosa. Gastroenterology 115, 357–369 (1998)
Kaneko, M., Niinuma, Y. & Nomura, Y. Activation signal of nuclear factor-κB in response to endoplasmic reticulum stress is transduced via IRE1 and tumor necrosis factor receptor-associated factor 2. Biol. Pharm. Bull. 26, 931–935 (2003)
Vaishnava, S., Behrendt, C. L., Ismail, A. S., Eckmann, L. & Hooper, L. V. Paneth cells directly sense gut commensals and maintain homeostasis at the intestinal host-microbial interface. Proc. Natl Acad. Sci. USA 105, 20858–20863 (2008)
Kuballa, P., Huett, A., Rioux, J. D., Daly, M. J. & Xavier, R. J. Impaired autophagy of an intracellular pathogen induced by a Crohn’s disease associated ATG16L1 variant. PLoS ONE 3, e3391 (2008)
Deuring, J. J. et al. Genomic ATG16L1 risk allele-restricted Paneth cell ER stress in quiescent Crohn’s disease. Gut http://dx.doi.org/10.1136/gutjnl-2012-303527 (2013)
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem. 277, 33275–33283 (2002)
Shimshek, D. R. et al. Codon-improved Cre recombinase (iCre) expression in the mouse. Genesis 32, 19–26 (2002)
Garabedian, E. M., Roberts, L. J., McNevin, M. S. & Gordon, J. I. Examining the role of Paneth cells in the small intestine by lineage ablation in transgenic mice. J. Biol. Chem. 272, 23729–23740 (1997)
Bry, L. et al. Paneth cell differentiation in the developing intestine of normal and transgenic mice. Proc. Natl Acad. Sci. USA 91, 10335–10339 (1994)
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001)
Iwawaki, T., Akai, R., Yamanaka, S. & Kohno, K. Function of IRE1 alpha in the placenta is essential for placental development and embryonic viability. Proc. Natl Acad. Sci. USA 106, 16657–16662 (2009)
Baert, L. et al. Detection of murine norovirus 1 by using plaque assay, transfection assay, and real-time reverse transcription-PCR before and after heat exposure. Appl. Environ. Microbiol. 74, 543–546 (2008)
Pierce, J. W. et al. Novel inhibitors of cytokine-induced IκBα phosphorylation and endothelial cell adhesion molecule expression show anti-inflammatory effects in vivo. J. Biol. Chem. 272, 21096–21103 (1997)
Satoh, Y., Yamano, M., Matsuda, M. & Ono, K. Ultrastructure of Paneth cells in the intestine of various mammals. J. Electron Microsc. Tech. 16, 69–80 (1990)
Vidal, K., Grosjean, I., Revillard, J. P., Gespach, C. & Kaiserlian, D. Immortalization of mouse intestinal epithelial cells by the SV40-large T gene. Phenotypic and immune characterization of the MODE-K cell line. J. Immunol. Methods 166, 63–73 (1993)
Hempel, S. L., Buettner, G. R., O’Malley, Y. Q., Wessels, D. A. & Flaherty, D. M. Dihydrofluorescein diacetate is superior for detecting intracellular oxidants: comparison with 2',7'-dichlorodihydrofluorescein diacetate, 5(and 6)-carboxy-2',7'-dichlorodihydrofluorescein diacetate, and dihydrorhodamine 123. Free Radic. Biol. Med. 27, 146–159 (1999)
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009)
Wei, Y., Sinha, S. & Levine, B. Dual role of JNK1-mediated phosphorylation of Bcl-2 in autophagy and apoptosis regulation. Autophagy 4, 949–951 (2008)
Carchman, E. H., Rao, J., Loughran, P. A., Rosengart, M. R. & Zuckerbraun, B. S. Heme oxygenase-1-mediated autophagy protects against hepatocyte cell death and hepatic injury from infection/sepsis in mice. Hepatology 53, 2053–2062 (2011)
Hu, P., Han, Z., Couvillon, A. D., Kaufman, R. J. & Exton, J. H. Autocrine tumor necrosis factor alpha links endoplasmic reticulum stress to the membrane death receptor pathway through IRE1α-mediated NF-κB activation and down-regulation of TRAF2 expression. Mol. Cell. Biol. 26, 3071–3084 (2006)
Urano, F. et al. Coupling of stress in the ER to activation of JNK protein kinases by transmembrane protein kinase IRE1. Science 287, 664–666 (2000)
Barrett, J. C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn’s disease. Nature Genet. 40, 955–962 (2008)
Tashiro, E. et al. Trierixin, a novel Inhibitor of ER stress-induced XBP1 activation from Streptomyces sp. 1. Taxonomy, fermentation, isolation and biological activities. J. Antibiot. 60, 547–553 (2007)
Rioux, J. D. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nature Genet. 39, 596–604 (2007)
Parkes, M. et al. Sequence variants in the autophagy gene IRGM and multiple other replicating loci contribute to Crohn’s disease susceptibility. Nature Genet. 39, 830–832 (2007)
Acknowledgements
We thank L. Glimcher, A. Goldberg, J. Yuan, M. Parkes, A. Franke, H. Tilg, M. Pasparakis, K. Vlantis, A.-H. Lee and C. L. Bevins for discussion of the project, are grateful to J. Gordon, L. Hooper and K. Rajewsky for providing critical reagents, and thank O. Will for initial handling of the Atg16l1 colony and help with DSS colitis. A. Kaser began work for this study at the Department of Internal Medicine II, Innsbruck Medical University, A-6020 Innsbruck, Austria. This work was supported by NIH grants DK044319, DK051362, DK053056, DK088199, the Harvard Digestive Diseases Center (HDDC) (DK0034854) (R.S.B.); the European Research Council under the European Community’s Seventh Framework Programme (FP7/2007-2013)/ERC Grant agreement no. 260961 (A.K.); the National Institute for Health Research Cambridge Biomedical Research Centre (A.K.); the Austrian Science Fund and Ministry of Science P21530-B18 and START Y446-B18 (A.K.); the Addenbrooke’s Charitable Trust (A.K. and L.N.); BMBF NGFN Animal Model grant (P.R.), the DFG Cluster of Excellence Inflammation at Interfaces (S.S. and P.R.); EU SysmedIBD grant (P.R), the Hans-Dietrich Bruhn Memorial Foundation (R.B.); DFG grants RO2994/5-1 (P.R.) and SFB 877 project B9 (P.R. and S.S.); fellowships from Inflammatory Bowel Disease Working Group (M.F.T.), Crohn’s and Colitis Foundation of America (M.B.F.), European Crohn’s and Colitis Organization (T.E.A.), Crohn’s in Childhood Research Association (A.K. and L.N.), National Research Foundation of Korea funded by the Korean government KRF-2008-357-E00022 no. 2011-0009018 (H. -J.K.).
Author information
Authors and Affiliations
Contributions
T.E.A., M.F.T., L.N. and H.-J.K. performed most experiments, together with J.B., E.M.-N., M.T., S.H., M.B.F., S.B.-B., T.R., R.B. and M.-N.K. J.L.C. helped prepare the manuscript. S.J.H. and J.H. contributed electron microscopic analysis, A.C. provided expertise in autophagy assessment, and R.L. in histology of Paneth cells. S.S. and P.R. designed, generated and analysed an essential mouse strain. K.K., T.I. and S.J.M. provided an essential mouse strain. J.N.G. assessed intestinal inflammation. A.K. and R.S.B. devised and coordinated the project, and together with T.E.A. and M.F.T. wrote the manuscript and designed the experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Autophagy induction in Xbp1-deficient intestinal epithelial cells.
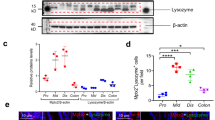
a, Xbp1 expression in shCtrl and shXbp1 MODE-K cells (n = 7/6; unpaired Student’s t-test; mean ± s.e.m.). b, c, Immunoblot of shCtrl and shXbp1 MODE-K cells. d, Immunoblot after autophagosome–lysosome fusion inhibition via bafilomycin in silenced MODE-K intestinal epithelial cells. e, Silenced MODE-K intestinal epithelial cells after GFP–LC3 reporter transfection (scale bars, 5 μm) with green punctae per cell quantification in f (n = 14; unpaired Student’s t-test; mean ± s.e.m.). g, TEM of shCtrl and shXbp1 cells. Note double-membraned structure with engulfed contents characteristic of autophagosomes (white arrows; n = 10). Scale bars, 0.5 μm. h, Quantification of occupied area and average size of autophagic vacuoles from g (n = 10; unpaired Student’s t-test; mean ± s.e.m.). i, j, Densitometry of Fig. 1a (i; n = 5/4; unpaired Student’s t-test; mean ± s.e.m.) and Fig. 1b (j; n = 3; unpaired Student’s t-test; mean ± s.e.m.). k, Low-magnification (×1,380 , original magnification here and in the remainder of this legend) TEM image of Paneth cells from wild-type mice (1), demonstrating the abundant endoplasmic reticulum (ER) and characteristic secretory granules at the apical, lumenally (L) oriented side. (2) Higher magnification (×5,520) of inset ‘2’ in (1) demonstrating typical secretory granules in Paneth cells from wild-type mice. (3) High magnification (×20,700) of inset ‘3’ in (1) illustrating a double membrane structure characteristic of an autophagosome (white arrow) in close proximity to the ER and a mitochondrion. (4) Low magnification (×2,160) TEM image of Paneth cell remnants present in Xbp1ΔIEC small intestinal crypts, which lack expansion of the ER and exhibit only minuscule granule remnants. (5) Higher magnification (×9,000) of inset ‘5’ in (4), demonstrating degradative autophagic vacuoles (black arrows), in close proximity to mitochondria, and the virtual absence of ER membranes. (6) High-power (×14,400) magnification of inset ‘6’ in (4), illustrating a double-membrane structure (white arrow) characteristic of autophagosomes, and a degradative autophagic vacuole (black arrow). ER, endoplasmic reticulum; L, lumen; M, mitochondrion; N, nucleus. Scale bars represent 2 μm, 0.5 μm and 200 nm, respectively. Results represent three (b, c) or two (d, k) independent experiments. *P < 0.05, ***P < 0.001.
Extended Data Figure 2 Autophagy in Xbp1-deficient intestinal epithelial cells is induced by eIF2α phosphorylation but independent of JNK or oxidative stress pathways.
a, Timeline for salubrinal experiment shown in Fig. 1j. b, Representative indirect immunofluorescence images of GFP–LC3 punctae accumulation (green) in small intestinal sections co-stained with anti-lysozyme antibody (red) treated as in a (n = 3). DAPI, blue. Scale bars, 10 μm. c, Immunoblot of crypt lysates after 3 days of tamoxifen or vehicle administration (n = 3). d, e, Immunoblot of primary intestinal epithelial cell scrapings (d); densitometry in e (n = 3; unpaired Student’s t-test; mean ± s.e.m.). f, g, Immunoblot of shXbp1 and shCtrl MODE-K cells. ER stress-induced Jun N-terminal kinase-1 (JNK1) has previously been connected in other cellular model systems to autophagy activation through phosphorylation of B-cell leukaemia 2 (Bcl-2) and its dissociation from beclin 143, as have oxidative stress/free radicals and haem oxygenase-1 (HO-1) activation44. h, Intracellular ROS determined by dichlorofluorescein assay and mean fluorescent intensity (MFI) after vehicle or dichlorofluorescein diacetate (DCF-DA) treatment. i, Immunoblot of shXbp1 and shCtrl MODE-K cells after administration of the JNK inhibitor SP600125 (0, 5 or 25 μM) for 4 h. Note the absence of an effect of SP600125 treatment on the conversion of LC3-I to LC3-II or the levels of p-eIF2α, thereby excluding a major contribution of the JNK pathway to autophagy induction in the presence of intestinal-epithelial-cell-associated XBP1 deficiency. j, Immunoblot of shXbp1 and shCtrl MODE-K cells after N-acetylcysteine (NAC), glutathione (GSH) or vehicle treatment for 16 h. Note the absence of an effect of the free radical scavengers on either of these markers of UPR-induced autophagy (LC3-II or p-eIF2α). Results represent three (f–j) independent experiments. *P < 0.05.
Extended Data Figure 3 ER stress-induced activation of PERK/eIF2α induces autophagy in Xbp1-deficiency.
a, Immunoblot of shXbp1 and shCtrl MODE-K cells co-silenced with Perk (siPerk) or scrambled siRNA. b, Cumulative densitometry of the immunoblot in Fig. 1g and two additional experiments (not shown, n = 10; unpaired Student’s t-test; mean ± s.e.m.). c, p-eIF2α immunohistochemistry of small intestinal epithelium. Scale bars, 50 μm. d, mRNA expression levels of Map1lc3b (LC3b) and Atg7 relative to Gapdh in shCtrl and shXbp1 MODE-K cells (n = 3; unpaired Student’s t-test; mean ± s.e.m.). e, Accumulation of GFP–LC3 punctae after salubrinal and 3-day tamoxifen treatment according to timeline in Extended Data Fig. 2a (n = 5). Scale bars, 5 μm. f, Immunoblot of primary epithelial cell scrapings from small intestine upon vehicle or salubrinal treatment according to the timeline in g (n = 2). Note the increased relative levels of LC3 conversion and CHOP, a transcriptional target of ATF4, in salubrinal-treated mice. g, Timeline of salubrinal experiment. Results are reported in f and Fig. 1k. h, Gadd34 expression in shCtrl and shXbp1 MODE-K cells co-silenced for Gadd34 (siGadd34) and control (siCtrl) (one-way ANOVA with post-hoc Holm’s corrected unpaired Student’s t-test; mean ± s.e.m.). i, Cells as in h were immunoblotted. j, Immunoblot of crypt lysates (n = 2/4/2). Results represent three (a, h, i) independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 4 Absence of Atg7 in intestinal epithelium disrupts autophagy and exacerbates spontaneous enteritis of Xbp1 deficiency and development of conditional Atg16l1 knockout mouse.
a, Immunoblot of primary intestinal epithelial cell scrapings (n = 4/3/3/3). Note the loss of ATG5, ATG12 and ATG16L1 proteins in ATG7-deficient mice. b, Representative high magnification TEM images (n = 2). Note intact ER in wild-type mice; severe distortion of ER in Atg7ΔIEC mice; autophagic vacuoles with the characteristic double membrane (white arrow) in Xbp1ΔIEC mice; and the absence of ER and autophagic vacuoles in Atg7/Xbp1ΔIEC mice. Scale bars, 200 nm. ER, endoplasmic reticulum. c, Quantification of autophagic vacuoles (n = 10; unpaired Student’s t-test; mean ± s.e.m.). d, High-magnification (×400) haematoxylin and eosin images. Note knife-like extension of inflammation along blood vessels through whole thickness of muscularis propria into the serosa in Atg7/Xbp1ΔIEC mice. Scale bars, 10 μm. e, Linear regression analysis for the correlation of inflammation with age. Each dot represents a single animal (R2 and P value for deviation from 0 are shown). f, Deep inflammation score (as described in Methods) in Atg7/Xbp1ΔIEC mice plotted at indicated ages in weeks (n = 13/17/9; median shown; Kruskal–Wallis with post-hoc Holm’s-corrected Mann–Whitney U-test). g, Atg16l1 intestinal-epithelial-cell-specific knockout design. Exon 1 of the Atg16l1 gene was flanked by two loxP sites and Atg16l1 deletion was mediated by Cre recombinase (Cre) under control of the Villin (V-) promoter (V-cre+;Atg16l1fl/fl or Atg16l1ΔIEC). See Methods for a detailed description. h, Immunohistochemical staining (brown) with anti-ATG16L1 antibody (MBL) (n = 2). Note cytoplasmic intestinal-epithelial-cell-specific ATG16L1 immunostaining in wild-type but not Atg16l1ΔIEC mice with retained ATG16L1 expression in lamina propria mononuclear cells in Atg16l1ΔIEC mice. Scale bars, 50 μm. i, Indirect immunofluorescence microscopy (green) with anti-ATG16L1 antibody (Cell Signaling Technology) (n = 2). Note the similar staining pattern as in h with cytoplasmic intestinal-epithelial-cell-specific ATG16L1 immunoreactivity in wild-type but not Atg16l1ΔIEC mice and retained ATG16L1 expression in lamina propria mononuclear cells in Atg16l1ΔIEC mice. Dashed line denotes crypt unit. Scale bars, 20 μm. j, Schematic representation of lysozyme+ granule allocation patterns (green): normal (D0), disordered (D1), depleted (D2), and diffuse (D3)2. k, l, Lysozyme immunofluorescence (green) in crypts of Atg16l1ΔIEC and wild-type mice (k), quantified in l (n = 3; unpaired Student’s t-test; mean ± s.e.m.) according to granule allocation patterns shown in j. Dashed line denotes crypt unit. DAPI, blue. Scale bars, 5 μm. m, n, Quantification of Paneth cells based on the number of granular vesicles stained with toluidine blue (m) as shown in n (n = 4; unpaired Student’s t-test; mean ± s.e.m.). Scale bars, 5 μm. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 5 Atg16l1 intestinal-epithelial-cell-specific deletion abrogates UPR-induced autophagy and exacerbates ER stress-induced spontaneous intestinal inflammation due to Xbp1 deficiency.
a, Immunoblot of primary intestinal epithelial cell scrapings. Note identical GAPDH loading control as in Fig. 3b (n = 3). b, Representative high-magnification TEM images (n = 2). Note intact ER in wild-type mice; severe distortion of ER in Atg16l1ΔIEC mice; autophagic vacuoles with the characteristic double membrane (arrow) in Xbp1ΔIEC mice; and absence of ER and autophagic vacuoles in Atg16l1/Xbp1ΔIEC mice. Scale bar, 200 nm. ER, endoplasmic reticulum. c, Quantification of autophagic vacuoles (n = 10 unpaired Student’s t-test; mean ± s.e.m.). d, Representative images of p62 immunostaining on small intestinal sections (n = 3). Brown staining indicates p62 with maximal signal intensity in the region of Paneth cells. Scale bars, 20 μm. e, High-resolution haematoxylin and eosin images. Note the extension of the inflammation into muscularis propria (arrow) in Atg16l1/Xbp1ΔIEC mice. Scale bars, 20 μm. f, Deep inflammation score (as described in Methods) in mice assessed at 18 weeks of age (n = 11; median shown; Kruskal–Wallis with post-hoc Dunn’s correction). g, Enteritis histology score (n = 14; 7–10-week-old mice housed in Innsbruck; median shown; Kruskal–Wallis with post-hoc Holm’s-corrected Mann–Whitney U-test). h, qRT–PCR of grp78 in primary intestinal epithelial cell scrapings (n = 7; unpaired Student’s t-test; mean ± s.e.m.). i, Disease activity index during DSS colitis (n = 7/4, two-way ANOVA with post-hoc Bonferroni; mean ± s.e.m.). j, Representative endoscopic images from the colon at day 5 of DSS treatment (n = 3). k, Representative haematoxylin and eosin staining of colonic sections from 5-day DSS-treated and untreated mice (n = 7/4). Scale bars, 50 μm. l, Colitis score after 5 days of DSS administration (n = 7/4, Mann–Whitney U-test). *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 6 ER stress-induced enteritis of Xbp1 deficiency exacerbated by autophagy impairment correlates with age and cell death in intestinal epithelial cells and is dependent on IRE1α-mediated NF-κB activation.
a, Representative images of TUNEL-labelled intestinal epithelium (brown) (n = 7). Scale bars, 50 μm. b, Correlation of TUNEL+ cells with the severity of inflammation. Linear regression analysis of Atg16l1/Xbp1ΔIEC (left) and Atg7/Xbp1ΔIEC (right) with significant R2 and P value for deviation from zero shown (left panel, n = 5/5/7/7; right panel, n = 5). c, Linear least square regression analysis for the correlation of enteritis histology score with cell death and age of animals by genotype. Each dot represents a single animal (grey, wild type; yellow, Atg16l1ΔIEC; blue, Xbp1ΔIEC; red, Atg16l1/Xbp1ΔIEC mice) and the plane represents the linear regression for the enteritis histology score as a function of age and TUNEL labelling for Atg16l1/Xbp1ΔIEC mice (n = 6/13/12/12). Note that the severity of inflammation significantly correlates with numbers of TUNEL+ intestinal epithelial cells and age only in Atg16l1/Xbp1ΔIEC mice (R2 = 0.602, P = 0.016). The three-dimensional (3D) plot is also available online in video format (Supplementary Video 1). Regression analysis was performed using the R package lessR (http://cran.r-project.org/web/packages/lessR/index.html), last accessed May 2013. d, shCtrl or shXbp1 MODE-K cells were co-silenced for Atg16l1 (siAtg16l1) or with scrambled siRNA (siCtrl), and analysed by flow cytometry for annexin V and propidium iodide (PI) uptake (one-way ANOVA with post-hoc Bonferroni; mean ± s.e.m.). e, Densitometry of the immunoblot shown in Fig. 3b (n = 3; one-way ANOVA with post-hoc Holm’s corrected unpaired Student’s t-test; mean ± s.e.m.). f, Immunohistochemical staining for p-IκBα in the small intestinal epithelium (n = 3). Scale bars, 20 μm. g, NF-κB consensus sequence binding assay after stimulation of shCtrl and shXbp1 MODE-K cells for 5 and 20 min with indicated concentrations of TNF. h, Nfkbia expression, a prototypic NF-κB-transactivated gene, after TNF stimulation of shCtrl and shXbp1 MODE-K cells (mean ± s.e.m.). i, Representative images of TUNEL-labelled sections after administration of BAY11-7082 or vehicle (n = 3/4/4). Scale bars, 50 μm. j, Enteritis histology score of mice treated with the NF-κB inhibitor BAY11-7082 or vehicle (n = 7/9/8; median shown; Kruskal–Wallis with post-hoc Holm’s-corrected Mann–Whitney U-test). k, Expression of Nfkbia in Ern1- and control-silenced shXbp1 and control MODE-K cells after TNF stimulation (mean ± s.e.m.). Results represent three (h, k) or two (d, g) independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 7 Rapamycin induces autophagy in Paneth cells, reduces the severity of enteritis and the number of TUNEL+ intestinal epithelial cells in Xbp1ΔIEC but not in Atg7/Xbp1ΔIEC or Atg16l1/Xbp1ΔIEC mice.
a, b, Quantification of GFP–LC3 punctae accumulation after 3-day tamoxifen treatment and induction by rapamycin (a; n = 10; one-way ANOVA with post-hoc Bonferroni; mean ± s.e.m.) and representative images in b (n = 3). Scale bars, 5 μm. c, Densitometry of the immunoblot in Fig. 4a (n = 3; one-way ANOVA with post-hoc Holm’s-corrected unpaired Student’s t-test; mean ± s.e.m.). d, Quantification of TUNEL+ intestinal epithelial cells per cm of gut in mice treated with rapamycin or vehicle (n = 5; unpaired Student’s t-test; mean ± s.e.m.). e, Representative images of TUNEL-labelled sections from rapamycin- or vehicle-treated mice. Scale bars, 50 μm. f, g, Quantification of TUNEL+ intestinal epithelial cells per cm of gut in mice treated with rapamycin or vehicle (f; n = 4/4/5/5/5/5; g; n = 5; one-way ANOVA with post-hoc Holm’s-corrected unpaired Student’s t-test; mean ± s.e.m.). h, i, Enteritis histology scores of rapamycin- or vehicle-treated mice of respective genotypes (h; n = 8/9/6/9/12/13; i, n = 8/7/6/7/7/6) (median shown; Kruskal–Wallis with post-hoc Holm’s-corrected Mann–Whitney U-test). *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 8 Autophagy induction in Paneth cells of Xbp1ΔPC mice.
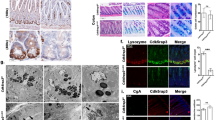
a, Representative TEM images of small intestinal crypts from Defa6-cre+ (wild type) and Defa6-cre+;Xbp1fl/fl (Xbp1ΔPC) mice. (1) Low magnification TEM image of Paneth cells at the base of a small intestinal crypt in wild-type mice, demonstrating abundant ER and characteristic secretory granules at the apical, lumenally oriented side. Scale bar, 5 μm. (2) Higher magnification of inset ‘2’ in (1) with numerous secretory granules. Scale bar, 1 μm. (3) High magnification of inset ‘3’ in (1) with intact ER. Scale bar, 200 nm. (4) Low-magnification TEM image of Paneth cells in an Xbp1ΔPC small intestinal crypt. Note the reduced number of secretory granules, the exfoliating hypomorphic Paneth cell into crypt lumen (‘E’, black dashed outline) and a transmigrating polymorphonuclear cell through the mucosa (white dashed outline). Scale bar, 5 μm. (5) Higher magnification of inset ‘5’ in (4), demonstrating abnormalities in secretory granule maturation with numerous hypodense granules (*) and a distorted ER. Note the accumulation of electron-dense cargo indicative of degradative autophagic vacuoles (black arrow) in close proximity to the nucleus. Scale bar, 1 μm. (6) High magnification of the inset ‘6’ in (4), demonstrating degradative autophagic vacuoles filled with electron-dense material (black arrow). Note the double membrane structures characteristic for autophagosomes (white arrowheads). Scale bar, 200 nm. (7) Low-magnification TEM image of Paneth cells in an Xbp1ΔPC small intestinal crypt. Note the disorganized ER, hypodense secretory granules (*), empty area after exfoliation of a Paneth cell into the crypt lumen (‘E’, black dashed outline) similar to image (4). (8) High magnification of inset ‘8’ identified in (7), demonstrating an autophagic vacuole (black arrow) surrounded by a markedly distorted ER. Scale bar, 200 nm. (9) High magnification of the inset ‘9’ in (7), with additional examples of degradative autophagic vacuoles accompanied by a distorted ER. Representative of two independent experiments. Scale bar, 200 nm. ER, endoplasmic reticulum; L, lumen; M, mitochondrion; N, nucleus. b, Densitometry of the immunoblot in Fig. 4d (n = 2; unpaired Student’s t-test; mean ± s.e.m.). *P < 0.05.
Extended Data Figure 9 Phenotypic consequences of Xbp1 deletion in Paneth cells and characterization of murine norovirus status.
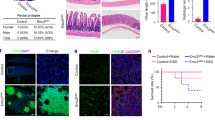
a, b, TUNEL-labelled ileal sections (a) with quantification of TUNEL+ cells per cm gut shown in b (n = 7/11; unpaired Student’s t-test; mean ± s.e.m.). Scale bars, 50 μm. c, d, Ki67 immunohistochemical staining of ileal sections (c) with quantification of Ki67+ cells per total intestinal epithelial cell count along the crypt–villus axis shown in d (n = 5; unpaired Student’s t-test; mean ± s.e.m.). Scale bars, 20 μm. e, f, BrdU-labelled ileal sections (e) after 24 h with quantification of BrdU+ cells per total intestinal epithelial cell count along the crypt–villus axis in f (n = 5/3; unpaired Student’s t-test; mean ± s.e.m.). Scale bars, 50 μm. g, h, PAS-stained ileal sections (g) with quantification of PAS+ cells per total intestinal epithelial cell count along the crypt–villus axis shown in h (n = 5; unpaired Student’s t-test; mean ± s.e.m.). Scale bars, 50 μm. i, Enteritis histology score of indicated genotypes (n = 3; median shown; Kruskal–Wallis). j, Taqman qRT–PCR for MNV of faecal samples in three different animal facilities (n = 3; mean ± s.e.m.). Note detectable MNV genome copy numbers in Boston (Atg7/Xbp1ΔIEC, Fig. 2c) and Innsbruck (Atg16l1/Xbp1ΔIEC, Extended Data Fig. 5g) colonies, and absence of MNV after re-derivation of colonies into the MNV-free Cambridge facility (Atg16l1/Xbp1ΔIEC, Fig. 2e), yet similar levels of enteritis in Xbp1ΔIEC and Atg16l1/Xbp1ΔIEC mice maintained in Innsbruck (MNV+) and Cambridge (MNV−) (Fig. 2b, e and Extended Data Fig. 5g) and relatively more severe disease in Xbp1ΔIEC mice from Boston (MNV+) (Fig. 2c). Sentinel mice from the Cambridge colony also tested negative for MNV by serological analysis (data not shown). These observations suggest that the inflammatory phenotypes observed are not necessarily dependent on MNV. *P < 0.05.
Extended Data Figure 10 Model of the interplay between UPR-induced autophagy, NF-κB signalling and inflammation.
a, Deletion of Xbp1 in the intestinal epithelium, specifically in Paneth cells, leads to ER stress and activation of the PERK–eIF2α branch of the UPR. ATF4, a transcriptional mediator of this pathway, transactivates genes essential for autophagosome formation, such as Map1lc3b (LC3b) and Atg7, which catalyses the creation of the ATG12–ATG5 conjugate that stabilizes ATG16L1 through complex formation21. UPR-induced autophagy in the intestinal epithelium is essential for restoration of homeostasis and restraint of ER stress-induced intestinal inflammation due to XBP1 deficiency. Activation of the UPR in the setting of XBP1 deficiency results in activation of IRE1α, resulting in the recruitment of TRAF2 and activation of IKK2 leading to IκBα degradation4,27,45,46. As shown here, UPR-mediated autophagy serves an important role in restraining NF-κB activation, conceivably by removing hyperinflammatory ER membranes containing activated IRE1α. Pharmacological augmentation of this compensatory autophagy-dependent mechanism via inhibition of eIF2α dephosphorylation through salubrinal, or via the mTOR inhibitor rapamycin, results in amelioration of UPR-induced enteritis, which is driven by the commensal microbiota, NF-κB, and TNF-R1 signalling in intestinal epithelial cells and myeloid cells, whereby the ligand TNF may originate from XBP1-deficient intestinal epithelial cells4. b, ATG16L1 deficiency in intestinal epithelial cells leads to ER stress as revealed through upregulation of the chaperone GRP78 in intestinal epithelial cells, increased expression of GRP78 protein in Paneth cells, increased IRE1α expression and increased splicing of Xbp1 mRNA in intestinal crypts as well as increased intestinal epithelial cell death. This leads to increased sensitivity of the epithelium to environmental triggers (for example, dextran sodium sulphate) that further challenge the UPR and its compensatory pathways. c, Deficiency of ATG16L1 or ATG7 in the intestinal epithelium results in abrogation of the compensatory autophagic mechanism that restrains IRE1α activity, conceivably via removal of hyperinflammatory ER membranes, and further fosters intestinal epithelial cell death in the context of ER stress due to Xbp1 deficiency, resulting in spontaneous transmural small intestinal inflammation that is associated with further increases in NF-κB activation and cell death via the mechanisms described in a. The UPR allows for responses to a variety of signals that have an impact on protein folding, including genetic (for example, rare XBP1 variants, ORMDL3 as risk factor of inflammatory bowel disease4,47), environmental (for example, low O2 tension in the intestinal tract) and microbial factors (for example, microbial toxins such as trierixin48), which determines the level of ER stress in the intestinal epithelium. UPR-induced autophagy function provides a buffer to cope with different levels of ER stress and vice versa. However, in the presence of genetic risk variants, such as ATG16L1 (refs 1, 17, 49) or IRGM (ref. 50), which are relatively prevalent in the general population, this compensatory mechanism is impaired, resulting in development and/or exacerbation of intestinal inflammation in the setting of unabated ER stress.
Supplementary information
Supplementary Tables
This file contains Supplementary Table 1. (PDF 132 kb)
Inflammation significantly correlates with age and TUNEL+ IECs in Atg16l1/Xbp1ΔIEC mice
Three-dimensional linear least square regression analysis for the correlation of enteritis histology score with cell death and age of animals by genotype. Each dot represents a single animal (grey, Wt; yellow, Atg16l1ΔIEC; blue, Xbp1ΔIEC; red, Atg16l1/Xbp1ΔIEC mice) and the plane represents the linear regression for the histological score as a function of age and TUNEL labeling for Atg16l1/Xbp1ΔIEC mice. Note that the severity of inflammation significantly correlates with numbers of TUNEL+ IECs and age in Atg16l1/Xbp1ΔIEC mice (R2=0.602, p=0.016) but not any other genotype (n=6/12/12/12). Regression analysis was performed using the R package lessR (http://cran.r-project.org/web/packages/lessR/index.html), last accessed May 2013. (MP4 2380 kb)
Rights and permissions
About this article
Cite this article
Adolph, T., Tomczak, M., Niederreiter, L. et al. Paneth cells as a site of origin for intestinal inflammation. Nature 503, 272–276 (2013). https://doi.org/10.1038/nature12599
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12599
This article is cited by
-
Loss of Mptx2 alters bacteria composition and intestinal homeostasis potentially by impairing autophagy
Communications Biology (2024)
-
Paneth cells in farm animals: current status and future direction
Journal of Animal Science and Biotechnology (2023)
-
A stromal lineage maintains crypt structure and villus homeostasis in the intestinal stem cell niche
BMC Biology (2023)
-
ERdj5 protects goblet cells from endoplasmic reticulum stress-mediated apoptosis under inflammatory conditions
Experimental & Molecular Medicine (2023)
-
Trans-Golgi protein TVP23B regulates host-microbe interactions via Paneth cell homeostasis and Goblet cell glycosylation
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.