Abstract
The complexity of the human brain has made it difficult to study many brain disorders in model organisms, highlighting the need for an in vitro model of human brain development. Here we have developed a human pluripotent stem cell-derived three-dimensional organoid culture system, termed cerebral organoids, that develop various discrete, although interdependent, brain regions. These include a cerebral cortex containing progenitor populations that organize and produce mature cortical neuron subtypes. Furthermore, cerebral organoids are shown to recapitulate features of human cortical development, namely characteristic progenitor zone organization with abundant outer radial glial stem cells. Finally, we use RNA interference and patient-specific induced pluripotent stem cells to model microcephaly, a disorder that has been difficult to recapitulate in mice. We demonstrate premature neuronal differentiation in patient organoids, a defect that could help to explain the disease phenotype. Together, these data show that three-dimensional organoids can recapitulate development and disease even in this most complex human tissue.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Götz, M. & Huttner, W. B. The cell biology of neurogenesis. Nature Rev. Mol. Cell Biol. 6, 777–788 (2005)
Zecevic, N., Chen, Y. & Filipovic, R. Contributions of cortical subventricular zone to the development of the human cerebral cortex. J. Comp. Neurol. 491, 109–122 (2005)
Fietz, S. A. et al. OSVZ progenitors of human and ferret neocortex are epithelial-like and expand by integrin signaling. Nature Neurosci. 13, 690–699 (2010)
Hansen, D. V., Lui, J. H., Parker, P. R. L. & Kriegstein, A. R. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561 (2010)
Smart, I. H. M., Dehay, C., Giroud, P., Berland, M. & Kennedy, H. Unique morphological features of the proliferative zones and postmitotic compartments of the neural epithelium giving rise to striate and extrastriate cortex in the monkey. Cereb. Cortex 12, 37–53 (2002)
Shitamukai, A., Konno, D. & Matsuzaki, F. Oblique radial glial divisions in the developing mouse neocortex induce self-renewing progenitors outside the germinal zone that resemble primate outer subventricular zone progenitors. J. Neurosci. 31, 3683–3695 (2011)
Lui, J. H., Hansen, D. V. & Kriegstein, A. R. Development and evolution of the human neocortex. Cell 146, 18–36 (2011)
Fietz, S. A. & Huttner, W. B. Cortical progenitor expansion, self-renewal and neurogenesis — a polarized perspective. Curr. Opin. Neurobiol. 21, 23–35 (2011)
Cox, J., Jackson, A. P., Bond, J. & Woods, C. G. What primary microcephaly can tell us about brain growth. Trends Mol. Med. 12, 358–366 (2006)
Megraw, T. L., Sharkey, J. T. & Nowakowski, R. S. Cdk5rap2 exposes the centrosomal root of microcephaly syndromes. Trends Cell Biol. 21, 470–480 (2011)
Barrera, J. A. et al. CDK5RAP2 regulates centriole engagement and cohesion in mice. Dev. Cell 18, 913–926 (2010)
Lizarraga, S. B. et al. Cdk5rap2 regulates centrosome function and chromosome segregation in neuronal progenitors. Development 137, 1907–1917 (2010)
Pulvers, J. N. et al. Mutations in mouse Aspm (abnormal spindle-like microcephaly associated) cause not only microcephaly but also major defects in the germline. Proc. Natl Acad. Sci. USA 107, 16595–16600 (2010)
Gruber, R. et al. MCPH1 regulates the neuroprogenitor division mode by coupling the centrosomal cycle with mitotic entry through the Chk1-Cdc25 pathway. Nature Cell Biol. 13, 1325–1334 (2011)
Sato, T. et al. Single Lgr5 stem cells build crypt–villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009)
Suga, H. et al. Self-formation of functional adenohypophysis in three-dimensional culture. Nature 480, 57–62 (2011)
Nakano, T. et al. Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell Stem Cell 10, 771–785 (2012)
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011)
Eiraku, M. & Sasai, Y. Self-formation of layered neural structures in three-dimensional culture of ES cells. Curr. Opin. Neurobiol. 22, 768–777 (2012)
Eiraku, M. et al. Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell 3, 519–532 (2008)
Danjo, T. et al. Subregional specification of embryonic stem cell-derived ventral telencephalic tissues by timed and combinatory treatment with extrinsic signals. J. Neurosci. 31, 1919–1933 (2011)
Muguruma, K. et al. Ontogeny-recapitulating generation and tissue integration of ES cell-derived Purkinje cells. Nature Neurosci. 13, 1171–1180 (2010)
Mariani, J. et al. Modeling human cortical development in vitro using induced pluripotent stem cells. Proc. Natl Acad. Sci. USA 109, 12770–12775 (2012)
Xia, X. & Zhang, S.-C. Differentiation of neuroepithelia from human embryonic stem cells. Methods Mol. Biol. 549, 51–58 (2009)
Swanson, L. W. Mapping the human brain: past, present, and future. Trends Neurosci. 18, 471–474 (1995)
Bedogni, F. et al. Tbr1 regulates regional and laminar identity of postmitotic neurons in developing neocortex. Proc. Natl Acad. Sci. USA 107, 13129–13134 (2010)
Yoo, A. S. et al. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476, 228–231 (2011)
Lessard, J. et al. An essential switch in subunit composition of a chromatin remodeling complex during neural development. Neuron 55, 201–215 (2007)
Willardsen, M. I. & Link, B. A. Cell biological regulation of division fate in vertebrate neuroepithelial cells. Dev. Dyn. 240, 1865–1879 (2011)
Chenn, A. & McConnell, S. K. Cleavage orientation and the asymmetric inheritance of Notch1 immunoreactivity in mammalian neurogenesis. Cell 82, 631–641 (1995)
Konno, D. et al. Neuroepithelial progenitors undergo LGN-dependent planar divisions to maintain self-renewability during mammalian neurogenesis. Nature Cell Biol. 10, 93–101 (2008)
Yingling, J. et al. Neuroepithelial stem cell proliferation requires LIS1 for precise spindle orientation and symmetric division. Cell 132, 474–486 (2008)
Postiglione, M. P. et al. Mouse inscuteable induces apical-basal spindle orientation to facilitate intermediate progenitor generation in the developing neocortex. Neuron 72, 269–284 (2011)
Smart, I. H. Proliferative characteristics of the ependymal layer during the early development of the mouse neocortex: a pilot study based on recording the number, location and plane of cleavage of mitotic figures. J. Anat. 116, 67–91 (1973)
Zamenhof, S. Quantitative studies of mitoses in fetal rat brain: orientations of the spindles. Brain Res. 428, 143–146 (1987)
LaMonica, B. E., Lui, J. H., Hansen, D. V. & Kriegstein, A. R. Mitotic spindle orientation predicts outer radial glial cell generation in human neocortex. Nature Commun 4, 1665 (2013)
Hevner, R. F. et al. Tbr1 regulates differentiation of the preplate and layer 6. Neuron 29, 353–366 (2001)
Shafit-Zagardo, B. & Kalcheva, N. Making sense of the multiple MAP-2 transcripts and their role in the neuron. Mol. Neurobiol. 16, 149–162 (1998)
Frotscher, M. Cajal-Retzius cells, Reelin, and the formation of layers. Curr. Opin. Neurobiol. 8, 570–575 (1998)
Tsai, L.-H. & Gleeson, J. G. Nucleokinesis in neuronal migration. Neuron 46, 383–388 (2005)
Gaspard, N. et al. An intrinsic mechanism of corticogenesis from embryonic stem cells. Nature 455, 351–357 (2008)
De Carlos, J. A. & O’Leary, D. D. Growth and targeting of subplate axons and establishment of major cortical pathways. J. Neurosci. 12, 1194–1211 (1992)
Chédotal, A. Further tales of the midline. Curr. Opin. Neurobiol. 21, 68–75 (2011)
Sato, T. R., Gray, N. W., Mainen, Z. F. & Svoboda, K. The functional microarchitecture of the mouse barrel cortex. PLoS Biol. 5, e189 (2007)
Bond, J. et al. A centrosomal mechanism involving CDK5RAP2 and CENPJ controls brain size. Nature Genet. 37, 353–355 (2005)
Pagnamenta, A. T. et al. A novel nonsense CDK5RAP2 mutation in a Somali child with primary microcephaly and sensorineural hearing loss. Am. J. Med. Genet. 158A, 2577–2582 (2012)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Okita, K., Ichisaka, T. & Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 448, 313–317 (2007)
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nature Biotechnol. 25, 681–686 (2007)
Hu, B.-Y. & Zhang, S.-C. Directed differentiation of neural-stem cells and subtype-specific neurons from hESCs. Methods Mol. Biol. 636, 123–137 (2010)
Matsuda, T. & Cepko, C. L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl Acad. Sci. USA 101, 16–22 (2004)
Siegenthaler, J. A. et al. Retinoic acid from the meninges regulates cortical neuron generation. Cell 139, 597–609 (2009)
Tremml, G., Singer, M. & Malavarca, R. Culture of mouse embryonic stem cells. Curr. Protoc. Stem Cell Biol. 1, Unit–1C.4 (2008)
Acknowledgements
We are grateful to members of the Knoblich laboratory for technical expertise and feedback, A. Peer, P. Moeseneder and N. Corsini for experimental support and M. Repic for help with establishing organoid electroporations. We also thank the Stem Cell and BioOptics core facilities of IMBA/IMP for technical support. We would especially like to thank the patients and their families for participating in this study. We would also like to thank S. McGurk for providing control MRI images. M.A.L. received funding from an EMBO post-doctoral fellowship and a Helen Hay Whitney post-doctoral fellowship. Work in A.P.J.’s laboratory is supported by the Medical Research Council, a starter grant from the European Research Council (ERC) and the Lister Institute for Preventative Medicine. This research was also supported in part by Wellcome Trust grant WT098051. Work in J.A.K.’s laboratory is supported by the Austrian Academy of Sciences, the Austrian Science Fund (FWF) (projects Z153-B09 and I552-B19) and an advanced grant from ERC.
Author information
Authors and Affiliations
Contributions
M.A.L. and J.A.K. conceived the project and experimental design and wrote the manuscript. M.A.L. performed experiments and analysed data. M.R., C.-A.M. and D.W. performed experiments and analysed data under the supervision of J.A.K., J.M.P. and A.P.J. L.S.B., M.E.H. and T.H. performed patient diagnosis and provided MRIs coordinated by A.P.J. J.A.K. directed and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Generation of cerebral organoids from multiple human pluripotent stem cells.
a, Haematoxylin and eosin staining of cerebral organoids compared with stationary culture reveals overall larger tissues with substructure reminiscent of brain regions such as forebrain cortex (arrows) and choroid plexus (arrowhead). b, Higher magnification images of haematoxylin and eosin stained organoids revealing layering reminiscent of the cerebral cortical molecular layer (bar), as well as tissue reminiscent of meninges (arrowheads) and choroid plexus (arrows). c, TUNEL staining (green) revealing cell death in the interior regions (arrows) of the cerebral organoid with cortical regions developing along the exterior. DAPI marks nuclei (blue). d, Haematoxylin and eosin staining of organoids generated from human H9 ES cells as well as human iPS cells display similar size and complex morphology as well as the presence of advanced forebrain tissues, shown at higher magnification in the bottom panels. e, Staining for N-cadherin (green) and newborn neurons (DCX, red) in tissues generated from both human H9 ES cells and human iPS cells reveals similar organization and intact apical basal polarity in both types of tissues. Scale bars, 0.5 mm (a), 100 μm (b, c, d bottom panels, e), and 0.5 mm (d top panels).
Extended Data Figure 2 Neural identity during differentiation of cerebral organoids.
a, RT–PCR for the pluripotency markers OCT4 and NANOG as well as neural identity markers SOX1 and PAX6 in undifferentiated human ES cells and following differentiation at 9 days, revealing induction of neural identity with decreased pluripotent identity at 9 days of differentiation. b, Immunohistochemistry for the forebrain/midbrain marker OTX1/2 (green) and the hindbrain marker GBX2 (red) at 16 days of differentiation, revealing primarily fore/midbrain identity with adjacent regions of hindbrain reminiscent of the mid–hindbrain boundary (arrows). DAPI marks nuclei (blue). c, Staining for the cortical lobe markers LMO4 (frontal and occipital marker, green) and TSHZ2 (occipital marker, red). Note the expected nuclear staining (arrows and arrowheads) for both in one region (top panels) suggesting occipital identity, whereas only LMO4 staining (arrowheads) is clearly evident in another region (bottom panels), suggesting frontal identity. DAPI marks nuclei (blue). d, Staining for the ventral marker NKX2-1 (red) and the cortical interneuron marker calretinin (green) on an organoid containing both ventral (arrowheads) and dorsal (top left) regions within one section. Images on the right are higher magnification stitched images of the region outlined in the lower magnification image at left. Calretinin interneurons can be seen between the two regions with typical morphology of migration and redirection towards the dorsal cortex (arrows). Scale bars, 100 μm.
Extended Data Figure 3 RG organization and morphology.
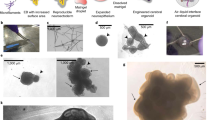
a, Staining for the chromatin remodelling BAF components BAF53A (also known as ACTL6A) (green, top panels) and BAF53B (also known as ACTL6B) (green, bottom panels) in serial sections of the same tissue showing the neural-progenitor-specific BAF53A expressed in VZ RGs, whereas the neuron-specific BAF53B is expressed in DCX+ (red) neurons outside the VZ. b, Higher magnification image of p-vimentin staining (green) of a dividing radial glia revealing the long basal process typical of radial glial morphology. c, Schematic of electroporation technique. Plasmid DNA was injected into fluid-filled cavities within the organoid and an electric pulse was applied to electroporate cells (radial glial progenitors) adjacent to the cavity. This resulted in several regions of electroporation (top right, GFP in green) and high efficiency of electroporation of RGs (bottom, GFP in green). d, GFP-electroporated progenitors (arrows) in an early stage tissue (18 days) revealing neuroepithelial morphology. e, GFP-electroporated tissue at 30 days, revealing RG (arrows) with typical bipolar morphology (arrowheads). f, GFP-electroporated tissue at 36 days revealing more advanced thicker cortical region with RG (arrow) exhibiting long apical and basal processes (arrowheads). g, p-Vimentin (green) staining revealing a mitotic cell at the apical surface during anaphase (arrow) with a horizontal orientation of division.
Extended Data Figure 4 Spatial organization and characteristics of cortical neuron identities.
a, Staining for the preplate marker TBR1 (green) and the deep-layer marker CTIP2 (red) at day 30, revealing rudimentary spatial separation reminiscent of the early stages of cortical plate development. b, Immunohistochemistry for the early-born neuron marker CTIP2 (green) and later-born neuron marker BRN2 (red) reveals independent neuron populations exhibiting rudimentary separation at 30 days of differentiation. c, GFP (green)-electroporated neuronal axon 5 days after electroporation displaying complex morphology and axon branching (arrowheads). d, GFP (green)-electroporated cortical neurons (arrows) 5 days after electroporation extend long-range axons with evidence of axon bundling (arrowheads) similar to that seen in pyramidal tracts. e, Single-cell tracings of calcium surges in individual neurons (ROI, outlined in left panel) as measured by change in fluorescence (arbitrary units).
Extended Data Figure 5 Human features of cortical development are not recapitulated in mouse organoids.
a, Low-magnification image of the region shown in Fig. 5a revealing the presence of a separated region of oRGs (demarcated by arrowheads) that appear separate from the VZ in all regions (brackets) but more separated and with a layer of TUJ1+ fibres in between in thicker parts of the cortical tissue (larger bracket). The entire organoid can be seen in Fig. 1c. b, Low-magnification image of a cerebral organoid derived from mouse ES cells stained for neurons (Tuj1, green) and neural progenitors (Sox2, red) revealing overall smaller organoid size as well as smaller cortical regions (arrows) than in humans. c, Higher magnification of a region of cortical identity in mouse cerebral organoids stained for RG progenitors (Sox2, red) revealing the presence of only a few oRGs (arrowheads) that do not organize into a separate layer such as that seen in humans.
Extended Data Figure 6 Patient growth parameters.
a, All growth parameters were significantly reduced both at birth and postnatally, with all z-scores less than −2 s.d. from the population mean for age and sex (dashed line). Weight (wgt), height (hgt) and head circumference (occipitofrontal circumference, ofc) at birth and at current age of 3.5 years of age. Head circumference was much more severely affected than height and weight, indicating that brain volume was disproportionately reduced as a result of more severe growth restriction. b, CDK5RAP2 (red) is absent from the centrosome in patient fibroblasts. Immunofluorescent microscopy images of patient (A3842) and control cells, also stained with the centriolar marker CPAP (green).
Extended Data Figure 7 Characterization of patient-derived iPS cells and cerebral organoids.
a, iPS cells derived from A3842 patient skin fibroblasts exhibit typical ES-cell-like morphology. Four lines were chosen for analysis on the basis of this typical morphology and pluripotency. b, Alkaline phosphatase staining (blue) of patient-derived iPS cell colonies revealing pluripotency. c, Representative early organoid culture of patient (line 1M) and control using the protocol and timing established for normal human ES cells. Patient organoids were much smaller and failed to thrive, therefore the protocol was slightly modified with increased starting cell number to produce neural tissues. d, Patient-derived tissues using increased starting cell number displayed neuroepithelium but did not form thick fluid-filled cortical tissues compared with control-derived tissues. Patient-derived tissues also display outgrowth with neural morphology compared with control. e, Staining of patient and control organoids at an early stage (day 22) for neurons (TUJ1, green) and RG (SOX2, red) revealed smaller progenitor zones (arrowheads) and increased neurons in patient-derived tissues (lines 1M and 14B are shown here). f, Quantification of the percentage of SOX2+ progenitors and TUJ1+ neurons in cerebral cortical regions of control and two lines of patient-derived tissues (1M and 14B) at the early stage of day 22. Error bars are ± s.e.m. ***P < 0.001 compared with control, Student’s t-test. n = 4 tissues for each line. g, Bright-field image of patient-derived tissues (line 14B) electroporated with either GFP alone (left panel) or a GFP and CDK5RAP2 expression construct (right panel). Note the presence of larger neuroepithelial tissue (arrows) in CDK5RAP2-electroporated tissue compared with control. h, GFP staining (green) in GFP control (left) and CDK5RAP2 co-electroporated patient-derived tissues (14B) revealing the presence of multiple GFP+ neurons (arrowheads) in control 6 days after electroporation, whereas CDK5RAP2-electroporated tissues display multiple GFP+ RG (arrows).
Extended Data Figure 8 shRNA-mediated knockdown of CDK5RAP2 in human organoids.
a, Western blot for endogenous CDK5RAP2 in 293T cells transfected with four different shRNAs against CDK5RAP2. shRNAs 1 and 2 are most efficient, whereas shRNA 4 leads to a modest reduction in protein. α-Tubulin is shown as a loading control. b, Higher magnification of results in Fig. 6i showing neuronal morphology of CDK5RAP2 shRNA (GFP, green, arrowheads)-electroporated cells. These exhibit increased DCX (blue) expression with an expansion of the zone of DCX positivity (bars) and a loss of SOX2 (red) compared with scrambled electroporated or adjacent non-electroporated tissue (arrows). c, Quantification of the percentage of GFP+ electroporated cells exhibiting SOX2+ progenitor identity or DCX+ neuronal identity in scrambled control- or shRNA-co-electroporated tissues. ***P < 0.001 compared to control, Student’s t-test, n = 4 tissues for each shRNA. Error bars are ± s.e.m.
Supplementary information
Supplementary Information
This file contains Supplementary Text. (PDF 83 kb)
Interkinetic nuclear migration in cerebral organoids
Live imaging of GFP electroporated organoid revealing movement of nuclei along apical and basal processes of RG. Arrow marks one RG in particular with clear IKNM. Time shown in hrs:min. (MOV 1777 kb)
Calcium surges in neurons of cerebral organoids
Live imaging of Fluo-4 signal in a human cerebral organoid revealing spontaneous calcium surges in individual neurons (arrows). Time shown in min:sec. (MOV 4247 kb)
False colour heat map of spontaneous neural activity
False colour heat map of a zoomed in region of Supplemental Video 2 showing spontaneous calcium surges. Time shown in min:sec. (MOV 5755 kb)
Rights and permissions
About this article
Cite this article
Lancaster, M., Renner, M., Martin, CA. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013). https://doi.org/10.1038/nature12517
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12517
This article is cited by
-
A multicellular brain spheroid model for studying the mechanisms and bioeffects of ultrasound-enhanced drug penetration beyond the blood‒brain barrier
Scientific Reports (2024)
-
Schizophrenia endothelial cells exhibit higher permeability and altered angiogenesis patterns in patient-derived organoids
Translational Psychiatry (2024)
-
Cerebral organoids display dynamic clonal growth and tunable tissue replenishment
Nature Cell Biology (2024)
-
Comparing stem cells, transdifferentiation and brain organoids as tools for psychiatric research
Translational Psychiatry (2024)
-
Eavesdropping on brain organoids
Nature Biotechnology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.