Abstract
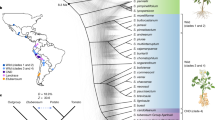
Potato (Solanum tuberosum L.) originates from the Andes and evolved short-day-dependent tuber formation as a vegetative propagation strategy. Here we describe the identification of a central regulator underlying a major-effect quantitative trait locus for plant maturity and initiation of tuber development. We show that this gene belongs to the family of DOF (DNA-binding with one finger) transcription factors1 and regulates tuberization and plant life cycle length, by acting as a mediator between the circadian clock and the StSP6A mobile tuberization signal2. We also show that natural allelic variants evade post-translational light regulation, allowing cultivation outside the geographical centre of origin of potato. Potato is a member of the Solanaceae family and is one of the world’s most important food crops. This annual plant originates from the Andean regions of South America3. Potato develops tubers from underground stems called stolons. Its equatorial origin makes potato essentially short-day dependent for tuberization and potato will not make tubers in the long-day conditions of spring and summer in the northern latitudes. When introduced in temperate zones, wild material will form tubers in the course of the autumnal shortening of day-length. Thus, one of the first selected traits in potato leading to a European potato type4 is likely to have been long-day acclimation for tuberization. Potato breeders can exploit the naturally occurring variation in tuberization onset and life cycle length, allowing varietal breeding for different latitudes, harvest times and markets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
The sequence of the StCDF1.2 gene variant is available in GenBank (accession number KC284850). All other sequences from potato are available at GenBank or the Potato Genome Sequencing Consortium website (http://www.potatogenome.net).
References
Yanagisawa, S. Dof domain proteins: plant-specific transcription factors associated with diverse phenomena unique to plants. Plant Cell Physiol. 45, 386–391 (2004)
Navarro, C. et al. Control of flowering and storage organ formation in potato by FLOWERING LOCUS T. Nature 478, 119–122 (2011)
Spooner, D. M., McLean, K., Ramsay, G., Waugh, R. & Bryan, G. J. A single domestication for potato based on multilocus amplified fragment length polymorphism genotyping. Proc Natl Acad. Sci. USA 102, 14694–14699 (2005)
Salaman, R. N. The History and Social Influence of the Potato (Cambridge Univ. Press, 1989)
Bäurle, I. & Dean, C. The timing of developmental transitions in plants. Cell 125, 655–664 (2006)
Koornneef, M., Hanhart, C. J. & van der Veen, J. H. A genetic and physiological analysis of late flowering mutants in Arabidopsis thaliana. Mol. Gen. Genet. 229, 57–66 (1991)
Kojima, S. et al. Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1 under short-day conditions. Plant Cell Physiol. 43, 1096–1105 (2002)
Lifschitz, E. & Eshed, Y. Universal florigenic signals triggered by FT homologues regulate growth and flowering cycles in perennial day-neutral tomato. J. Exp. Bot. 57, 3405–3414 (2006)
Pin, P. A. et al. An antagonistic pair of FT homologs mediates the control of flowering time in sugar beet. Science 330, 1397–1400 (2010)
Suárez-López, P. et al. CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 410, 1116–1120 (2001)
Yanovsky, M. J. & Kay, S. A. Molecular basis of seasonal time measurement in Arabidopsis. Nature 419, 308–312 (2002)
Park, D. H. et al. Control of circadian rhythms and photoperiodic flowering by the Arabidopsis GIGANTEA gene. Science 285, 1579–1582 (1999)
Imaizumi, T., Schultz, T. F., Harmon, F. G., Ho, L. A. & Kay, S. A. FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science 309, 293–297 (2005)
Sawa, M., Nusinow, D. A., Kay, S. A. & Imaizumi, T. FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science 318, 261–265 (2007)
Song, Y. H., Smith, R. W., To, B. J., Millar, A. J. & Imaizumi, T. FKF1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science 336, 1045–1049 (2012)
González-Schain, N. D., Diaz-Mendoza, M., Zurczak, M. & Suarez-Lopez, P. Potato CONSTANS is involved in photoperiodic tuberization in a graft-transmissible manner. Plant J. 70, 678–690 (2012)
Rodríguez-Falcón, M., Bou, J. & Prat, S. Seasonal control of tuberization in potato: conserved elements with the flowering response. Annu. Rev. Plant Biol. 57, 151–180 (2006)
Visker, M. H. et al. Can the QTL for late blight resistance on potato chromosome 5 be attributed to foliage maturity type? Theor. Appl. Genet. 106, 317–325 (2003)
The Potato Genome Sequencing Consortium. Genome sequence and analysis of the tuber crop potato. Nature 475, 189–195 (2011)
Abelenda, J. A., Navarro, C. & Prat, S. From the model to the crop: genes controlling tuber formation in potato. Curr. Opin. Biotechnol. 22, 287–292 (2011)
Martínez-García, J. F., Virgos-Soler, A. & Prat, S. Control of photoperiod-regulated tuberization in potato by the Arabidopsis flowering-time gene CONSTANS. Proc. Natl Acad. Sci. USA 99, 15211–15216 (2002)
Takase, T. et al. LOV KELCH PROTEIN2 and ZEITLUPE repress Arabidopsis photoperiodic flowering under non-inductive conditions, depending on FLAVIN-BINDING KELCH REPEAT F-BOX1. Plant J. 67, 608–621 (2011)
Inui, H., Ogura, Y. & Kiyosue, T. Overexpression of Arabidopsis thaliana LOV KELCH REPEAT PROTEIN 2 promotes tuberization in potato (Solanum tuberosum cv. May Queen).. FEBS Lett. 584, 2393–2396 (2010)
Karimi, M., Inzé, D. & Depicker, A. GATEWAYTM vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002)
Gady, A. L. F. et al. Implementation of two high through-put techniques in a novel application: detecting point mutations in large EMS mutated plant populations. Plant Methods 5, 13 (2009)
Calderon-Villalobos, L. I. et al. LucTrap vectors are tools to generate luciferase fusions for the quantification of transcript and protein abundance in vivo. Plant Physiol. 141, 3–14 (2006)
Voinnet, O., Rivas, S., Mestre, P. & Baulcombe, D. An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J. 33, 949–956 (2003)
Acknowledgements
We would like to thank K. Kaufman, G. Angenent, N. Mansoori Zangir, J. Uitdewilligen and R. Hutten for help and advice in the planning and execution of the project. Thanks also go to V. Ghasemi Omran and I. Pereira-Bertram for their expert technical assistance and to T. Bisseling for critical reading of the manuscript. We gratefully acknowledge the Dutch potato breeding companies, Argrico Research B.V., HZPC Holland B.V. and Meijer B.V. for the execution of the field trials. The project was supported by the Technology Foundation STW (grant no. WGC.7795 and WPB.5283) and FP6 integrated Project EU-Sol PL016214. M.d.M.C.G. received additional support from Almeria University Fund.
Author information
Authors and Affiliations
Contributions
B.K., J.A.A., M.d.M.C.G., M.O., J.M.d.B., K.K., B.M.H., C.S. and C.W.B.B. performed the experiments and analysed the results. B.K., S.P., H.J.v.E., J.M.d.B., R.G.F.V. and C.W.B.B. designed the experiments, C.W.B.B. wrote the manuscript and J.A.A., S.P., B.M.H., B.K., M.d.M.C.G., H.J.v.E., J.M.d.B., K.K. and R.G.F.V. gave critical feedback on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary Figures 1-7, Supplementary Tables 1-3 and Supplementary References. (PDF 1871 kb)
Rights and permissions
About this article
Cite this article
Kloosterman, B., Abelenda, J., Gomez, M. et al. Naturally occurring allele diversity allows potato cultivation in northern latitudes. Nature 495, 246–250 (2013). https://doi.org/10.1038/nature11912
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11912
This article is cited by
-
Transcriptional and Post-transcriptional Regulation of Tuberization in Potato (Solanum tuberosum L.)
Journal of Plant Growth Regulation (2024)
-
High-Density Linkage Map Constructed from a Skim Sequenced Diploid Potato Population Reveals Transmission Distortion and QTLs for Tuber Yield and Pollen Shed
Potato Research (2024)
-
BEL transcription factors in prominent Solanaceae crops: the missing pieces of the jigsaw in plant development
Planta (2024)
-
Potato and sweetpotato breeding at the International Potato Center: approaches, outcomes and the way forward
Theoretical and Applied Genetics (2024)
-
Investigating the genetic components of tuber bruising in a breeding population of tetraploid potatoes
BMC Plant Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.