Abstract
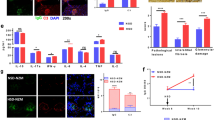
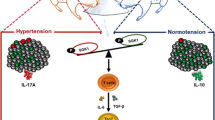
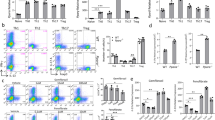
There has been a marked increase in the incidence of autoimmune diseases in the past half-century. Although the underlying genetic basis of this class of diseases has recently been elucidated, implicating predominantly immune-response genes1, changes in environmental factors must ultimately be driving this increase. The newly identified population of interleukin (IL)-17-producing CD4+ helper T cells (TH17 cells) has a pivotal role in autoimmune diseases2. Pathogenic IL-23-dependent TH17 cells have been shown to be critical for the development of experimental autoimmune encephalomyelitis (EAE), an animal model for multiple sclerosis, and genetic risk factors associated with multiple sclerosis are related to the IL-23–TH17 pathway1,2. However, little is known about the environmental factors that directly influence TH17 cells. Here we show that increased salt (sodium chloride, NaCl) concentrations found locally under physiological conditions in vivo markedly boost the induction of murine and human TH17 cells. High-salt conditions activate the p38/MAPK pathway involving nuclear factor of activated T cells 5 (NFAT5; also called TONEBP) and serum/glucocorticoid-regulated kinase 1 (SGK1) during cytokine-induced TH17 polarization. Gene silencing or chemical inhibition of p38/MAPK, NFAT5 or SGK1 abrogates the high-salt-induced TH17 cell development. The TH17 cells generated under high-salt conditions display a highly pathogenic and stable phenotype characterized by the upregulation of the pro-inflammatory cytokines GM-CSF, TNF-α and IL-2. Moreover, mice fed with a high-salt diet develop a more severe form of EAE, in line with augmented central nervous system infiltrating and peripherally induced antigen-specific TH17 cells. Thus, increased dietary salt intake might represent an environmental risk factor for the development of autoimmune diseases through the induction of pathogenic TH17 cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
International Multiple Sclerosis Genetics Consortium & Wellcome Trust Case Control Consortium. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011)
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V. K. IL-17 and Th17 Cells. Annu. Rev. Immunol. 27, 485–517 (2009)
Ascherio, A. & Munger, K. L. Environmental risk factors for multiple sclerosis. Part II: Noninfectious factors. Ann. Neurol. 61, 504–513 (2007)
McGuire, S. Institute of Medicine. 2010. Strategies to reduce sodium intake in the United States. Washington, DC: The National Academies Press. Adv. Nutr. 1, 49–50 (2010)
Appel, L. J. et al. The importance of population-wide sodium reduction as a means to prevent cardiovascular disease and stroke: a call to action from the American Heart Association. Circulation 123, 1138–1143 (2011)
Brown, I. J., Tzoulaki, I., Candeias, V. & Elliott, P. Salt intakes around the world: implications for public health. Int. J. Epidemiol. 38, 791–813 (2009)
Machnik, A. et al. Macrophages regulate salt-dependent volume and blood pressure by a vascular endothelial growth factor-C-dependent buffering mechanism. Nature Med. 15, 545–552 (2009)
Platten, M. et al. Blocking angiotensin-converting enzyme induces potent regulatory T cells and modulates TH1- and TH17-mediated autoimmunity. Proc. Natl Acad. Sci. USA 106, 14948–14953 (2009)
Stegbauer, J. et al. Role of the renin-angiotensin system in autoimmune inflammation of the central nervous system. Proc. Natl Acad. Sci. USA 106, 14942–14947 (2009)
Yang, L. et al. IL-21 and TGF-β are required for differentiation of human TH17 cells. Nature 454, 350–352 (2008)
Junger, W. G., Liu, F. C., Loomis, W. H. & Hoyt, D. B. Hypertonic saline enhances cellular immune function. Circ. Shock 42, 190–196 (1994)
Zielinski, C. E. et al. Pathogen-induced human TH17 cells produce IFN-γ or IL-10 and are regulated by IL-1β. Nature 484, 514–518 (2012)
Zhou, L. & Littman, D. R. Transcriptional regulatory networks in Th17 cell differentiation. Curr. Opin. Immunol. 21, 146–152 (2009)
Ghoreschi, K. et al. Generation of pathogenic TH17 cells in the absence of TGF-β signalling. Nature 467, 967–971 (2010)
Codarri, L. et al. RORγt drives production of the cytokine GM-CSF in helper T cells, which is essential for the effector phase of autoimmune neuroinflammation. Nature Immunol. 12, 560–567 (2011)
El-Behi, M. et al. The encephalitogenicity of TH17 cells is dependent on IL-1- and IL-23-induced production of the cytokine GM-CSF. Nature Immunol. 12, 568–575 (2011)
Reboldi, A. et al. C–C chemokine receptor 6-regulated entry of TH-17 cells into the CNS through the choroid plexus is required for the initiation of EAE. Nature Immunol. 10, 514–523 (2009)
O’Connell, R. M. et al. MicroRNA-155 promotes autoimmune inflammation by enhancing inflammatory T cell development. Immunity 33, 607–619 (2010)
Shapiro, L. & Dinarello, C. A. Osmotic regulation of cytokine synthesis in vitro. Proc. Natl Acad. Sci. USA 92, 12230–12234 (1995)
Go, W. Y., Liu, X., Roti, M. A., Liu, F. & Ho, S. N. NFAT5/TonEBP mutant mice define osmotic stress as a critical feature of the lymphoid microenvironment. Proc. Natl Acad. Sci. USA 101, 10673–10678 (2004)
Kino, T. et al. Brx mediates the response of lymphocytes to osmotic stress through the activation of NFAT5. Sci. Signal. 2, ra5 (2009)
Chen, S. et al. Tonicity-dependent induction of Sgk1 expression has a potential role in dehydration-induced natriuresis in rodents. J. Clin. Invest. 119, 1647–1658 (2009)
Ortells, M. C. et al. Transcriptional regulation of gene expression during osmotic stress responses by the mammalian target of rapamycin. Nucleic Acids Res. 40, 4368–4384 (2012)
Woehrle, T. et al. Hypertonic stress regulates T cell function via pannexin-1 hemichannels and P2X receptors. J. Leukoc. Biol. 88, 1181–1189 (2010)
Waldegger, S., Barth, P., Raber, G. & Lang, F. Cloning and characterization of a putative human serine/threonine protein kinase transcriptionally modified during anisotonic and isotonic alterations of cell volume. Proc. Natl Acad. Sci. USA 94, 4440–4445 (1997)
Bell, L. M. et al. Hyperosmotic stress stimulates promoter activity and regulates cellular utilization of the serum- and glucocorticoid-inducible protein kinase (Sgk) by a p38 MAPK-dependent pathway. J. Biol. Chem. 275, 25262–25272 (2000)
Waldegger, S. et al. h-sgk serine-threonine protein kinase gene as transcriptional target of transforming growth factor β in human intestine. Gastroenterology 116, 1081–1088 (1999)
Webster, M. K., Goya, L., Ge, Y., Maiyar, A. C. & Firestone, G. L. Characterization of sgk, a novel member of the serine/threonine protein kinase gene family which is transcriptionally induced by glucocorticoids and serum. Mol. Cell. Biol. 13, 2031–2040 (1993)
Waldegger, S., Gabrysch, S., Barth, P., Fillon, S. & Lang, F. h-sgk serine-threonine protein kinase as transcriptional target of p38/MAP kinase pathway in HepG2 human hepatoma cells. Cell. Physiol. Biochem. 10, 203–208 (2000)
Sherk, A. B. et al. Development of a small-molecule serum- and glucocorticoid-regulated kinase-1 antagonist and its evaluation as a prostate cancer therapeutic. Cancer Res. 68, 7475–7483 (2008)
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006)
Astier, A. L. et al. RNA interference screen in primary human T cells reveals FLT3 as a modulator of IL-10 levels. J. Immunol. 184, 685–693 (2010)
Reich, M. et al. GenePattern 2.0. Nature Genet. 38, 500–501 (2006)
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003)
Wenzel, K. et al. Potential relevance of α1-adrenergic receptor autoantibodies in refractory hypertension. PLoS ONE 3, e3742 (2008)
Noubade, R. et al. Activation of p38 MAPK in CD4 T cells controls IL-17 production and autoimmune encephalomyelitis. Blood 118, 3290–3300 (2011)
Engel, F. B. et al. p38 MAP kinase inhibition enables proliferation of adult mammalian cardiomyocytes. Genes Dev. 19, 1175–1187 (2005)
Bettelli, E. et al. Myelin oligodendrocyte glycoprotein-specific T cell receptor transgenic mice develop spontaneous autoimmune optic neuritis. J. Exp. Med. 197, 1073–1081 (2003)
Acknowledgements
The authors would like to thank S. Bhela, M. Zaidi, P. Quass, M. Mroz and S. Seubert for technical assistance and F. C. Luft for critical reading of the manuscript. We are grateful to J.-P. David and S. Teufel for providing Mx-Cre+ p38αfl/fl mice. This work was supported by a National MS Society Collaborative Research Center Award CA1061-A-18, National Institutes of Health Grants P01 AI045757, U19 AI046130, U19 AI070352, and P01 AI039671, and by a Jacob Javits Merit award (NS2427) from the National Institute of Neurological Disorders and Stroke, the Penates Foundation and the Nancy Taylor Foundation for Chronic Diseases, Inc. (to D.A.H.). R.A.L. was supported by the ELAN programme, University of Erlangen. D.N.M. was supported by the German Research Foundation (DFG) and the German Center for Cardiovascular Research (DZHK). J.T. was supported by the Interdisciplinary Center for Clinical Research at University of Erlangen and the German Research Foundation.
Author information
Authors and Affiliations
Contributions
M.K. designed the study, planned and performed experiments, analysed data and wrote the manuscript. A.M. planned and performed experiments, analysed data and wrote the manuscript. J.T. and H.K. interpreted data and supported the work with key suggestions and editing the manuscript. N.Y. analysed data. R.A.L. planned experiments, analysed data and wrote the manuscript. D.N.M. designed the study, planned experiments, analysed data and wrote the manuscript. D.A.H. designed the study, planned experiments, analysed data, and wrote the manuscript. M.K., D.N.M. and D.A.H. co-directed the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-15. (PDF 11533 kb)
Rights and permissions
About this article
Cite this article
Kleinewietfeld, M., Manzel, A., Titze, J. et al. Sodium chloride drives autoimmune disease by the induction of pathogenic TH17 cells. Nature 496, 518–522 (2013). https://doi.org/10.1038/nature11868
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11868
This article is cited by
-
Triggers for the onset and recurrence of psoriasis: a review and update
Cell Communication and Signaling (2024)
-
Meningeal interleukin-17-producing T cells mediate cognitive impairment in a mouse model of salt-sensitive hypertension
Nature Neuroscience (2024)
-
Immune and inflammatory mechanisms in hypertension
Nature Reviews Cardiology (2024)
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
The underlying mechanisms of DNA methylation in high salt memory in hypertensive vascular disease
Scientific Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.