Abstract
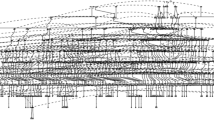
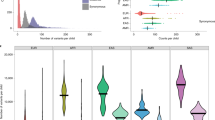
Mutations generate sequence diversity and provide a substrate for selection. The rate of de novo mutations is therefore of major importance to evolution. Here we conduct a study of genome-wide mutation rates by sequencing the entire genomes of 78 Icelandic parent–offspring trios at high coverage. We show that in our samples, with an average father’s age of 29.7, the average de novo mutation rate is 1.20 × 10−8 per nucleotide per generation. Most notably, the diversity in mutation rate of single nucleotide polymorphisms is dominated by the age of the father at conception of the child. The effect is an increase of about two mutations per year. An exponential model estimates paternal mutations doubling every 16.5 years. After accounting for random Poisson variation, father’s age is estimated to explain nearly all of the remaining variation in the de novo mutation counts. These observations shed light on the importance of the father’s age on the risk of diseases such as schizophrenia and autism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Keightley, P. D. Rates and fitness consequences of new mutations in humans. Genetics 190, 295–304 (2012)
Crow, J. F. The origins, patterns and implications of human spontaneous mutation. Nature Rev. Genet. 1, 40–47 (2000)
Kondrashov, A. S. Direct estimates of human per nucleotide mutation rates at 20 loci causing Mendelian diseases. Hum. Mutat. 21, 12–27 (2003)
Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012)
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012)
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012)
Xue, Y. et al. Human Y chromosome base-substitution mutation rate measured by direct sequencing in a deep-rooting pedigree. Curr. Biol. 19, 1453–1457 (2009)
Conrad, D. F. et al. Variation in genome-wide mutation rates within and between human families. Nature Genet. 43, 712–714 (2011)
Roach, J. C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010)
Holm, H. et al. A rare variant in MYH6 is associated with high risk of sick sinus syndrome. Nature Genet. 43, 316–320 (2011)
Rafnar, T. et al. Mutations in BRIP1 confer high risk of ovarian cancer. Nature Genet. 43, 1104–1107 (2011)
Sulem, P. et al. Identification of low-frequency variants associated with gout and serum uric acid levels. Nature Genet. 43, 1127–1130 (2011)
Keightley, P. D. et al. Analysis of the genome sequences of three Drosophila melanogaster spontaneous mutation accumulation lines. Genome Res. 19, 1195–1201 (2009)
Malaspina, D. Paternal factors and schizophrenia risk: de novo mutations and imprinting. Schizophr. Bull. 27, 379–393 (2001)
Croen, L. A., Najjar, D. V., Fireman, B. & Grether, J. K. Maternal and paternal age and risk of autism spectrum disorders. Arch. Pediatr. Adolesc. Med. 161, 334–340 (2007)
Duong, L. et al. Mutations in NRXN1 in a family multiply affected with brain disorders: NRXN1 mutations and brain disorders. Am. J. Med. Genet. 159B, 354–358 (2012)
Gauthier, J. et al. Truncating mutations in NRXN2 and NRXN1 in autism spectrum disorders and schizophrenia. Hum. Genet. 130, 563–573 (2011)
Kirov, G. et al. Comparative genome hybridization suggests a role for NRXN1 and APBA2 in schizophrenia. Hum. Mol. Genet. 17, 458–465 (2008)
Levinson, D. F. et al. Copy number variants in schizophrenia: confirmation of five previous findings and new evidence for 3q29 microdeletions and VIPR2 duplications. Am. J. Psychiatry 168, 302–316 (2011)
Rujescu, D. et al. Disruption of the neurexin 1 gene is associated with schizophrenia. Hum. Mol. Genet. 18, 988–996 (2009)
Boyden, L. M. et al. Mutations in kelch-like 3 and cullin 3 cause hypertension and electrolyte abnormalities. Nature 482, 98–102 (2012)
Lynch, M. Rate, molecular spectrum, and consequences of human mutation. Proc. Natl Acad. Sci. USA 107, 961–968 (2010)
Nachman, M. W. & Crowell, S. L. Estimate of the mutation rate per nucleotide in humans. Genetics 156, 297–304 (2000)
Coulondre, C., Miller, J. H., Farabaugh, P. J. & Gilbert, W. Molecular basis of base substitution hotspots in Escherichia coli. Nature 274, 775–780 (1978)
Kong, A. et al. Recombination rate and reproductive success in humans. Nature Genet. 36, 1203–1206 (2004)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Acknowledgements
This research was partly funded by The National Institutes of Health grant MH071425 (K.S.); the European Community’s Seventh Framework Programme, PsychCNVs project, grant agreement HEALTH-F2-2009-223423, and NextGene project, grant agreement IAPP-MC-251592; The European Community IMI grant EU-AIMS, grant agreement 115300.
Author information
Authors and Affiliations
Contributions
A.K. and K.S. planned and directed the research. A.K. wrote the first draft and together with K.S., S.B., P.S., A.H. and U.T. wrote the final version. O.T.M. and U.T. oversaw the sequencing and laboratory work. G. Masson, G. Magnusson and G.S. processed the raw sequencing data. A.K. and M.L.F. analysed the data, with W.S.W.W., H.H., G.B.W., S.S., G.T. and D.F.G. providing assistance. P.S. and S.A.G. performed functional annotations. S.B. analysed the mutations with respect to sequence content. A.S., Aslaug J. and Adalbjorg J. did the Sanger sequencing. A.H. investigated the contribution of demographics.
Corresponding authors
Ethics declarations
Competing interests
The authors from deCODE Genetics are employees of or own stock options in deCODE Genetics. W.S.W.W. is an employee of Illumina Inc., a public company that develops and markets systems for genetic analysis; she receives stocks as part of her compensation.
Supplementary information
Supplementary Information
This file contains Supplementary Text, additional references, Supplementary Table 2 and Supplementary Figure 1. (PDF 937 kb)
Supplementary Data
This file contains Supplementary Table 1 which shows information for each of the 4,933 de novo mutations individually. They correspond to the summary in Supplementary Table 2. The positions are based on Human Assembly Build 36. (XLS 523 kb)
Rights and permissions
About this article
Cite this article
Kong, A., Frigge, M., Masson, G. et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488, 471–475 (2012). https://doi.org/10.1038/nature11396
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11396
This article is cited by
-
Reconstructing the ancestral gene pool to uncover the origins and genetic links of Hmong–Mien speakers
BMC Biology (2024)
-
Meta-analysis of 46,000 germline de novo mutations linked to human inherited disease
Human Genomics (2024)
-
Assessment of parental mosaicism rates in neurodevelopmental disorders caused by apparent de novo pathogenic variants using deep sequencing
Scientific Reports (2024)
-
Neuronal knockdown of Cullin3 as a Drosophila model of autism spectrum disorder
Scientific Reports (2024)
-
Association of Paternal Age Alone and Combined with Maternal Age with Perinatal Outcomes: A Prospective Multicenter Cohort Study in China
Journal of Epidemiology and Global Health (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.