Abstract
Medulloblastoma is a malignant childhood brain tumour comprising four discrete subgroups. Here, to identify mutations that drive medulloblastoma, we sequenced the entire genomes of 37 tumours and matched normal blood. One-hundred and thirty-six genes harbouring somatic mutations in this discovery set were sequenced in an additional 56 medulloblastomas. Recurrent mutations were detected in 41 genes not yet implicated in medulloblastoma; several target distinct components of the epigenetic machinery in different disease subgroups, such as regulators of H3K27 and H3K4 trimethylation in subgroups 3 and 4 (for example, KDM6A and ZMYM3), and CTNNB1-associated chromatin re-modellers in WNT-subgroup tumours (for example, SMARCA4 and CREBBP). Modelling of mutations in mouse lower rhombic lip progenitors that generate WNT-subgroup tumours identified genes that maintain this cell lineage (DDX3X), as well as mutated genes that initiate (CDH1) or cooperate (PIK3CA) in tumorigenesis. These data provide important new insights into the pathogenesis of medulloblastoma subgroups and highlight targets for therapeutic development.
Similar content being viewed by others
Main
Medulloblastoma is the most common malignant childhood brain tumour1. The disease includes four subgroups (sonic hedgehog (SHH) subgroup, WNT subgroup, subgroup 3 and subgroup 4), defined primarily by gene expression profiling, that show differences in karyotype, histology and prognosis2. Studies of genetically engineered mice show that these tumours arise from different cell types: SHH-subgroup medulloblastomas develop from committed cerebellar granule neuron progenitors (GNPs) in Ptch1+/− mice3,4; WNT-subgroup tumours are generated by lower rhombic lip progenitors (LRLPs) in Blbp-Cre;Ctnnb1+/lox(Ex3) ;Tp53flx/flx mice5; whereas subgroup-3 medulloblastomas probably arise from an undefined class of cerebellar progenitors6. The identification of medulloblastoma subgroups has not changed clinical practice. All patients currently receive the same combination of surgery, radiation and chemotherapy. This aggressive treatment fails to cure two thirds of patients with subgroup-3 disease, and probably over-treats children with WNT-subgroup medulloblastoma who invariably survive with long-term cognitive and endocrine side effects2,7. Drugs targeting the genetic alterations that drive each medulloblastoma subgroup could prove more effective and less toxic, but the identity of these alterations remains largely unknown.
The genomic landscape of medulloblastoma
To identify genetic alterations that drive medulloblastoma, we performed whole-genome sequencing (WGS) of DNA from 37 tumours and matched normal blood (discovery cohort). Tumours were subgrouped by gene expression (WNT subgroup, n = 5; SHH subgroup, n = 5; subgroup 3, n = 6; subgroup 4, n = 19; ‘unclassified’ (profiles not available), n = 2; Fig. 1, Supplementary Figs 1–3 and Supplementary Table 1). Validation of all putative somatic alterations including single nucleotide variations (SNVs), insertion/deletions (indels) and structural variations (SVs) identified by CREST8, was conducted for 12 tumours using custom capture arrays and Illumina-based DNA sequencing (Supplementary Table 2). Putative coding alterations and SVs were validated in the remaining 25 discovery cohort cases by polymerase chain reaction (PCR) and Sanger-based sequencing. Mutation frequency was determined in a separate ‘validation cohort’ of 56 medulloblastomas (WNT subgroup, n = 6; SHH subgroup, n = 8; subgroup 3, n = 11; subgroup 4, n = 19; unclassified, n = 12; Fig. 1 and Supplementary Table 1).
Top, clinical, histological, gross chromosomal, nuclear CTNNB1 (nCTNNB1) and cohort (discovery or validation) details of 79 medulloblastomas by subgroup. ER, enrichment. Bottom, genetic alterations detected in 27 genes of particular interest. Colour key at top right. ANOVA (continuous) or Fisher’s exact test (categorical) P value is shown on right. False discovery rate (FDR) estimates of each mutation are shown on right. Slash indicates loss or mutation of wild-type allele, including X chromosome in males. ***P < 0.0005; **P < 0.005; *P < 0.05; NS, not significant. F, female; M, male. amp., amplification; del., deletion; microdel., microdeletion; valid., validation cohort. ND, not done.
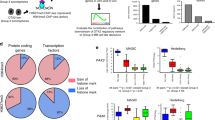
WGS of the discovery cohort detected 22,887 validated or high-quality somatic sequence mutations (SNVs and indels), 536 validated or curated SVs, and 5,802 copy number variations (CNVs; 92% concordant with 6.0 SNP mapping arrays; Supplementary Tables 3–6 and Supplementary Figs 4–7). In all but five tumours with the highest mutation rates, >50% of SNVs were C→T/G→A transitions (Supplementary Fig. 8). The mean missense:silent mutation ratio was 3.6:1 and 40% of all missense mutations were predicted to be deleterious, suggesting a selective pressure for SNVs that affect protein coding (Supplementary Table 5). Global patterns of total SNVs and amplifications varied significantly among medulloblastoma subgroups, even when corrected for age and sex, supporting the notion that these tumours are distinct pathological entities (Fig. 1 and Supplementary Fig. 6). Custom capture-based analysis of the allele frequency of all somatic mutations in 12 medulloblastomas allowed us to predict the ancestry of certain genetic alterations, suggesting that aneuploidy precedes widespread sequence mutation in medulloblastomas with highly mutated genomes (Supplementary Figs 9–11).
Novel CNVs and SVs are rare in medulloblastoma
The repertoire of focally amplified or deleted genes seems to be very limited in medulloblastoma. We detected expected2 gains of MYC, MYCN and OTX2 in subgroups 3 and 4, but no novel recurrent amplifications (Fig. 1, Supplementary Fig. 12 and Supplementary Table 7). In keeping with recent reports9, high-level amplification of MYCN in subgroup-3 sample no. 16 (sample numbering as in Fig. 1) was generated by chromothripsis; although chromothripsis was observed infrequently (n = 2/37 of the discovery cohort; Supplementary Fig. 13).
Focal homo- or heterozygous deletions of genes previously implicated in medulloblastoma were also detected (for example, PTCH1, PTEN; Fig. 1)10,11 but novel recurrent focal deletions were rare. Three subgroup-4 tumours (nos 11–13) and one unclassified tumour deleted DDX31, AK8 and TSC1 at chromosome 9q34.14 in concert with OTX2 amplification, suggesting that these alterations are cooperative (P < 0.0005, Fisher’s exact test). The breakpoint in this deletion occurs in DDX31, and two samples contained a missense mutation (subgroup 4, no. 15) and complex rearrangement (unidentified case SJMB026) in this gene, suggesting that DDX31 is the target of these alterations (Supplementary Fig. 14).
Over 50% of SVs detected by WGS broke the coding region of at least one gene, but less than 2% (n = 6/314, excluding two tumours with excessive SVs) encode potential in-frame fusion proteins (Supplementary Fig. 15); none affect the same gene or signal pathway. Therefore, fusion proteins are likely to be an uncommon transforming mechanism in medulloblastoma.
Although germline mutations in TP53, PTCH1, APC and CREBBP predispose to medulloblastoma11,12,13,14, only 23 mutations previously associated with cancer were detected in discovery cohort germ lines. Only one of these—in a known case of Turcot’s syndrome—was accompanied by a somatic mutation (germline APC Y935*/somatic deletion; WNT subgroup no. 11; Supplementary Table 8). Thus, inherited forms of medulloblastoma seem to be rare in our cohort.
Novel mutations in medulloblastoma subgroups
Because SVs and CNVs are unlikely to drive most medulloblastomas, we investigated whether recurrent (more than two samples) somatic SNVs and/or indels might target discrete genes and pathways. This analysis identified 49 genes, across all 93 tumours, which were targeted by non-silent, recurrent, somatic mutations; 84% (n = 41/49) have not yet been implicated in medulloblastoma (Supplementary Tables 9 and 10). Several of these congregated in disease subgroups and converged on specific cell pathways (Fig. 1, Supplementary Fig. 8 and Supplementary Table 11).
Histone methylation is deregulated in subgroups 3 and 4
The H3K27 trimethyl mark (H3K27me3) represses lineage-specific genes in stem cells15 (Supplementary Fig. 8). H3K27me3 is written by the polycomb repressive complex 2 (PRC2) that includes the methylase EZH2 (refs 16, 17) and is erased during differentiation by the demethylase KDM6A18. As H3K27me3 is erased, chromatin remodellers recruited to H3K4me3 promote differentiation, for example, CHD7 (refs 19, 20). This process is tightly controlled during development and deregulated in cancers; EZH2 is mutated in lymphomas21 and upregulated in breast22 and prostate23 cancer, while biallelic inactivation of KDM6A (chromosome Xp11.2) or KDM6A and its paralogue UTY (chromosome Yq11), occurs in adult female and male cancers, respectively24.
Hypergeometric distribution analyses revealed selective mutation of histone modifiers in subgroup-3 and -4 medulloblastomas (Supplementary Table 11). Six subgroup-4, one subgroup-3, and one unclassified medulloblastoma contained novel inactivating mutations in KDM6A (Figs 1 and 2 and Supplementary Figs 8 and 16). The single female with a KDM6A splice-site mutation showed a deletion of the second allele that escapes X inactivation25 (subgroup 4, no. 15), and 57% (n = 4/7) of KDM6A-mutant male medulloblastomas deleted chromosome Y, compared with only 6% (n = 3/51) of male, KDM6A wild-type tumours (P < 0.005, Fisher’s exact test; Fig. 1). Thus, a two-hit model of KDM6A-UTY tumour suppression seems to operate in subgroup-4 medulloblastomas. Notably, mutations in six other KDM family members (KDM1A, KDM3A, KDM4C, KDM5A, KDM5B and KDM7A) were detected exclusively in subgroup-3 and -4 tumours, implicating broad disruption of lysine demethylation in these medulloblastomas (Fig. 1, Supplementary Table 11 and Supplementary Fig. 16).
a, Top row, SNP profiles of chromosome 7 (Chr 7) copy number in medulloblastomas (samples as Fig. 1; asterisk indicates subgroup-3 cases). Second row, expression of EZH2. Subgroup-3 and -4 tumours are ordered left to right by expression level, dagger indicates median expression point (Bonferroni-corrected P value of EZH2 expression versus chromosome 7 gain). Third row, mutation status of KDM6A, CHD7 and ZMYM3 (P value, Fisher’s exact test mutations versus EZH2 expression). Fourth row, H3K27me3 immunohistochemistry (numbers indicate colorimetry, P value ANOVA). GP4-7 indicates case subgroup-4, no.7. a.u., arbitrary units. N/A, not available. b, H3K27me3 expression in mouse Blbp-Cre;Ctnnb1+/lox(Ex3) ;Tp53flx/flx (WNT-subgroup), Ptch1+/− ;Tp53−/− (SHH-subgroup) and Myc;Ink4c−/− (subgroup-3) medulloblastomas (right) and developing hindbrain (left). High-power views of E14.5 LRL (i) and upper rhombic lip (URL) (ii). EGL, external germinal layer; IGL, internal granule layer. Scale bar, 50 µm. White arrows in P7 cerebellum (CB) pinpoint H3K27me3 cells in the EGL.
Subgroup-3 and -4 medulloblastomas also gained and overexpressed EZH2 (chromosome 7q35-34), which writes H3K27me3, and contained novel inactivating mutations in effectors and regulators of the H3K4me3 mark26 (Fig. 2a and Supplementary Fig. 8). Gain of chromosome 7q was significantly enriched among subgroup-3 and -4 medulloblastomas (P < 0.005, Fisher’s exact test) and correlated directly with EZH2 expression. Indeed, EZH2 was the eighth most significantly overexpressed gene on chromosome 7 among subgroup-3 and -4 medulloblastomas that gained chromosome 7q relative to those with diploid chromosome 7 (P < 0.005, Bonferroni correction). Nonsense and frameshift mutations were detected in CHD7 in four subgroup-3 and -4 tumours. ZMYM3 (chromosome Xq13.1), which participates in a protein complex with KDM1A to regulate gene expression at the H3K4me3 mark27, was targeted by novel frameshift, nonsense and missense mutations in three male subgroup-4 medulloblastomas. All three tumours with mutations in ZMYM3 also mutated KDM6A (subgroup 4, nos 19, 20) or KDM1A (subgroup 4, no. 21), suggesting that these alterations are cooperative. Remarkably, KDM6A, CHD7 and ZMYM3 mutations were confined to subgroups 3 and 4, and clustered in samples with sub-median EZH2 expression levels (Fig. 2a; P < 0.05, Fisher’s exact test). These data suggest that subgroup-3 and -4 medulloblastomas retain a stem-like epigenetic state by aberrantly writing (EZH2 upregulation) or preserving (KDM6A-UTY inactivation) H3K27me3, or disrupting H3K4me3 associated transcription (CHD7 and ZMYM3 inactivation). Indeed, human and mouse subgroup-3 and -4 medulloblastomas contained significantly more H3K27me3 than did WNT- or SHH-subgroup tumours (Fig. 2b). Thus, gain of EZH2 and loss of KDM6A probably maintains H3K27me3 in subgroup-3 and -4 medulloblastomas.
Finally, we looked to see if the differential expression of H3K27me3 among medulloblastoma subgroups reflects ancestral chromatin marking in the progenitors that generate these tumours (Fig. 2b). Relatively low levels of H3K27me3 were detected in LRLPs and committed GNPs, which generate WNT- and SHH-subgroup medulloblastomas, respectively3,4,5, potentially explaining why mutations that preserve this epigenetic mark are absent from these tumours. We recently showed that subgroup-3 medulloblastomas arise from a rare fraction of cerebellar progenitors6. We are currently investigating whether these progenitors are found among the H3K27me3-positive cells seen in the external germinal layer (Fig. 2b).
Novel mutations in WNT-subgroup medulloblastomas
WNT-subgroup medulloblastomas contained mutations in epigenetic regulators that are different to those seen in subgroup-3 and -4 disease. CTNNB1, the principal effector of the WNT pathway, forms a transcription factor with the T-cell factor/lymphoid enhancer factor (TCF/LEF)28. The carboxy terminus of CTNNB1 then recruits a series of protein complexes that remodel chromatin and promote transcription at WNT-responsive genes (Supplementary Fig. 8). These include: histone acetyltransferases (for example, CREBBP and TRRAP–TIP60 complexes)28,29; ATPases of the SWI/SNF family (for example, SMARCA4)30; and the mediator complex that coordinates RNA polymerase II placement (for example, MED13)31. As expected, >70% (n = 8/11) of WNT-subgroup medulloblastomas contained mutations that stabilize CTNNB1 (Fig. 1 and Supplementary Fig. 8; P < 0.0001, Fisher’s exact test)32,33. A single subgroup-3 case (no. 5) also showed a mutation in CTNNB1, but this mutation has not been reported in cancer, did not upregulate nuclear CTNNB1 (Fig. 1) and is of unclear relevance. Remarkably, six WNT-subgroup medulloblastomas showed mutations in chromatin modifiers that are recruited to TCF/LEF WNT-responsive genes by CTNNB1 (Fig. 1 and Supplementary Fig. 8). Four WNT-subgroup tumours contained heterozygous missense mutations in the helicase domain of SMARCA4 (P < 0.002, Fisher’s exact test), two samples, including one with a SMARCA4 mutation (no. 5), contained nonsense mutations in CREBBP (WNT-subgroup enrichment, P < 0.02, Fisher’s exact test), and missense mutations in TRRAP and MED13 were detected in a single WNT-subgroup medulloblastoma each. Thus, in addition to stabilization of CTNNB1, the development of WNT-subgroup medulloblastoma may require disruption of chromatin remodelling at WNT-responsive genes.
A small number of WNT-subgroup medulloblastomas lack mutations in CTNNB1 or APC, suggesting that alternative mechanisms drive aberrant WNT signals in these tumours. Three WNT-subgroup medulloblastomas in our series contained wild-type CTNNB1 (nos 1, 10 and 11; Fig. 1). Sample no. 11 inactivated APC as the sole case of Turcot’s syndrome in our study, but this tumour and sample no. 10 also contained novel missense mutations in CDH1 (R63G, V329F; WNT-subgroup enrichment, P < 0.05, Fisher’s exact test; Fig. 1). CDH1 sequesters CTNNB1 at the cell membrane34, and mutations that disrupt this interaction promote WNT signalling in adult cancers35,36. The functional consequences of CDH1(R63G) and CDH1(V329F) remain to be determined, but their restriction to WNT-subgroup tumours, mutual exclusivity with CTNNB1 mutations, and adjacency to residues mutated in breast cancer (http://www.sanger.ac.uk/genetics/CGP/cosmic/), suggest they might promote aberrant WNT signals in medulloblastoma.
We showed previously in mice that mutant Ctnnb1 initiates WNT-subgroup medulloblastoma by arresting the migration of LRLPs from the embryonic dorsal brainstem to the pontine grey nucleus (PGN)5. Therefore, to test whether disruption of CDH1 might substitute for mutant CTNNB1 in medulloblastoma, we used short hairpin (sh)RNAs to knockdown Cdh1 in embryonic day (E)14.5 mouse LRLPs (Fig. 3a–c). Deletion of Cdh1 expression upregulated Tcf/Lef-mediated gene transcription in LRLPs and more than doubled their self-renewal capacity (Fig. 3b). Furthermore, in utero electroporation of LRLPs with Cdh1 shRNAs impeded their migration from the dorsal brainstem to the PGN with an efficiency similar to that of mutant Ctnnb1 (Fig. 3d, e; see Supplementary Methods). These data support the hypothesis that CDH1 suppresses the formation of WNT-subgroup medulloblastoma by regulating WNT-signals in LRLPs.
a, b, Isolated Olig3+/Wnt1+ LRLPs were transduced in b with mutant Ctnnb1 (above hashed line) or the indicated shRNA-RFP (red fluorescence protein) construct (below hashed line). LRLPs were also transduced (+) or not (−) with a Tcf/Lef-enhanced green fluorescence (Tcf) reporter. Numbers on right show clonal percentage 2′ to 3′ passage neurosphere formation (± standard deviation (s.d.)). N/A, not applicable. Scale bar, 10 μm. c, Knockdown of genes targeted by shRNA relative to control transduced cells. Data show mean ± s.d. d, Immunofluorescence of P1 mouse hindbrains electroporated in utero at E14.5 with GFP (to control for equivalence of electroporation between embryos control) and the indicated construct. High-power views of indicated areas are shown right. Cells targeted by Ddx3x shRNA are present 48 h after electroporation but ablated by P1. Scale bars, 200 µm. e, Heatmap showing the distribution of GFP+/RFP+ cells in eletroporated mice at P1. Median distance migrated by cells and P values of migration distance and cell number relative to controls is shown. ****P < 0.00005; ***P < 0.0005; ** P < 0.005; *P < 0.05. Red and green text reports significant increase or decrease, respectively, relative to control.
WNT-subgroup medulloblastomas were also enriched for novel, recurrent somatic missense mutations in the DEAD-box RNA helicase DDX3X at chromosome Xp11.3 (P < 0.0001, Fisher’s exact test; Fig. 1). DDX3X regulates several critical cell processes including chromosome segregation37, cell cycle progression38, gene transcription and translation39. Previously reported cancer-associated mutations in DDX3X disrupt the ATPase activity of the protein, but seven of eight mutations identified in our series clustered in the DEAD-box domain (Supplementary Information and Supplementary Fig. 8). Structural modelling predicts that these mutations interfere with nucleic acid binding, possibly altering specificity and/or affinity for RNA substrates, rather than inactivating DDX3X (Supplementary Figs 17–22). Indeed, the wild-type allele of DDX3X that escapes X inactivation25 was retained by two of three DDX3X-mutant female medulloblastomas, and knockdown of Ddx3x halved the self-renewal rate of mouse LRLPs, suggesting that this protein is important for the proliferation and/or maintenance of the LRLP lineage (Fig. 3b).
To understand better the role of DDX3X in WNT-subgroup medulloblastoma, we used our in utero migration assay to assess the impact of Ddx3x shRNAs, mutant Ddx3xT275M (identified in WNT-subgroup sample no. 9), or mutant Ddx3xG325E (WNT sample no. 8) on LRLPs. Remarkably, although Ddx3x shRNAs were expressed abundantly in E14.5 brainstem cells within 48 h of electroporation, ≤0.5% of Ddx3x-shRNA-positive cells were present by postnatal day (P)1, confirming the critical importance of this gene to maintain the LRLP lineage (Fig. 3d, e). In contrast, mice electroporated with either mutant Ddx3xT275M or Ddx3xG325E consistently contained ∼50% more labelled cells at P1 than did controls, although these cells migrated normally (Fig. 3d, e and data not shown). Thus, mutations in DDX3X may contribute to WNT-subgroup medulloblastoma by increasing LRLP proliferation rather than perturbing the migration of their daughter cells. Notably, comparable knockdown in utero of Mll2, Gabrg1 and Kdm6a that were selectively mutated in non-WNT medulloblastomas had no apparent impact on LRLPs; supporting the value of our assay for assessing WNT-subgroup specific mutations and underscoring the importance of cell context for functional studies of genes mutated in cancer subgroups.
PIK3CA mutations promote WNT-subgroup medulloblastoma
Cancer-associated, activating mutations in PIK3CA were detected in a single case each of WNT-subgroup (PIK3CA(Q546K)), SHH-subgroup (PIK3CA(H1047R)) and subgroup-4 (PIK3CA(N345K)) medulloblastoma (Fig. 1 and Supplementary Fig. 23). Although PIK3CA mutations are common in adult cancers40 and reported in medulloblastoma41, their role in tumorigenesis remains controversial. In particular it is not known if these mutations initiate or progress cancer. To test this, we generated mice that express a conditional allele of the Pik3caE545K mutation. Mice harbouring Pik3caE545K or Pik3caE545K and Tp53flx/flx were bred with Blbp-Cre, which drives efficient recombination in LRLPs5. Blbp-Cre;Pik3caE545K mice, with or without Tp53flx/flx , survived tumour free for a median of 212 days with no evidence of aberrant LRLP migration (Fig. 4a and data not shown). In stark contrast, 100% (n = 11/11) of Blbp-Cre;Ctnnb1+/lox(Ex3) ;Tp53+/flx ;Pik3caE545K mice developed WNT-subgroup medulloblastomas by 3 months of age; only 4% (n = 2/54) of Blbp-Cre;Ctnnb1+/lox(Ex3) ;Tp53+/flx mice develop WNT-subgroup medulloblastoma by 11 months (Fig. 4a, b). Pik3ca wild-type and mutant mouse medulloblastomas displayed similar ‘classic’ histologies and nuclear Ctnnb1+, but Pik3caE545K mutant tumours contained greater AKT pathway activity as measured by pS6 and p4EBP1 immunostaining. Thus mutations in PIK3CA probably activate the AKT pathway to progress, rather than initiate, WNT-subgroup medulloblastoma.
a, Tumour-free survival of mice of the indicated genotype. All mice carry the Blbp-cre allele. Log rank P < 0.0001. b, Haematoxylin and eosin (H&E) and immunohistochemical stains of indicated tumours. Scale bar, 50 µm.
SHH-subgroup medulloblastomas
Four of thirteen SHH-subgroup medulloblastomas contained expected biallelic inactivating alterations in SUFU or PTCH1. What drives aberrant SHH signals in the remaining cases remains unclear. These tumours contained mutations in MLL2, TP53 and PTEN that have been reported previously in medulloblastoma42; but these mutations occur in other subgroups and are not known to activate SHH signals. Two SHH-subgroup tumours (nos 11 and 12) contained identical novel T48M mutations in the GABAA (γ-aminobutyric acid, subtype A) receptor, γ1, which is predicted to be deleterious (Fig. 1 and Supplementary Table 9). Disruption of GABAA receptors can enhance neural stem cell proliferation43, suggesting that these mutations might deregulate the proliferation of GNPs that generate SHH-subgroup medulloblastomas.
Discussion
We have identified several, new, recurrent, somatic mutations in specific subgroups of medulloblastoma. Alterations affecting EZH2, KDM6A, CHD7 and ZMYM3 seem to disrupt chromatin marking of genes in subgroup-3 and -4 tumours. Further epigenetic studies will be required to uncover the identity of these genes, but evidence suggests these may include OTX2, MYC and MYCN44,45. As amplification of these genes was detected almost exclusively in subgroup-3 and -4 tumours that lacked mutations in KDM6A, CHD7 or ZMYM3, it is tempting to speculate that these genetic alterations target common transforming pathways. A recent study detected recurrent mutations in three other chromatin remodellers in medulloblastoma42: SMARCA4, MLL2 and MLL3, but this study did not include details of tumour subgroup. Here, we show that mutations in SMARCA4, CREBBP, TRRAP and MED13 are enriched in WNT-subgroup medulloblastomas; thereby uncovering potential cooperative mutations in chromatin remodellers and their binding-partner oncogene, CTNNB1. Thus, disruptions in the epigenetic machinery of medulloblastoma are likely to be subgroup specific and may cooperate with other oncogenic mutations. The low incidence of MLL2 mutations detected in our study relative to previous work42 probably reflects differences in study populations (see Supplementary Results).
Although medulloblastoma is more prevalent in males, especially with subgroup-3 and -4 disease46, the reason for this sex bias is unknown. One potential explanation is the location of medulloblastoma oncogenes or tumour suppressor genes on chromosome X47. Three of the most recurrently mutated genes detected in our study are located on chromosome X, of which two (ZMYM3 and KDM6A) were observed almost exclusively in males. Mutation in these genes might explain some of the male sex bias in medulloblastoma. The third mutated X chromosome gene, DDX3X, is more likely to be a WNT-subgroup medulloblastoma oncogene. Three of four female medulloblastomas carried heterozygous mutations in DDX3X that escape X inactivation25, and our functional data indicate that mutations in this gene provide a proliferative advantage to LRLPs that generate these tumours.
Our findings also have important implications for drug development. Inhibitors of the epigenetic machinery, especially those that maintain H3K27me3—for example, EZH2 methylase—may be useful treatments for subgroup-3 and -4 disease. These tumours include the most aggressive forms of medulloblastoma, for which treatment options are limited. Mutations that activate PIK3CA and DDX3X in WNT-subgroup tumours might also be targeted with novel therapeutic strategies48,49. Future clinical trials of drugs that target these mutant proteins must recruit the appropriate patient populations, as we demonstrate that mutations show subgroup specificity in medulloblastoma. Our accurate mouse models of WNT-subgroup, SHH-subgroup and subgroup-3 medulloblastoma should help with future studies of the biological and therapeutic importance of the novel genetic alterations described in this study.
Methods Summary
Human tumour and matched blood samples were obtained with informed consent through an institutional review board approved protocol at St Jude Children’s Research Hospital. WGS and analysis of WGS data were performed as previously described50. Details of sequence coverage, custom capture and other validation procedures are provided in Supplementary Information (Supplementary Tables 12–15). Immunohistochemistry and immunofluorescence of human and mouse tissues were performed using routine techniques and primary antibodies of the appropriate tissues as described (Supplementary Methods). Medulloblastoma mRNA and DNA profiles were generated using Affymetrix U133v2 and SNP 6.0 arrays, respectively (Supplementary Methods). Real-time PCR with reverse transcriptase (RT–PCR) analysis of genes targeted in mouse LRLPs by shRNAs were performed as described previously32. LRLPs were isolated and transduced with indicated lentiviruses in stem cell cultures or targeted in utero with shRNAs or mutant cDNA sequences by electroporation as described5 (Supplementary Information). Mice harbouring a Cre-inducible Pik3caE545K allele were generated using homologous recombination: a lox-puro-STOP-lox cassette was introduced immediately upstream of the exon containing the initiation codon, exon 9 was replaced with an exon containing the E545K mutation. Pik3caE545K mice were bred with Blbp-Cre;Ctnnb1lox(Ex3)/lox(Ex3) and Tp53flx/flx mice to generate progeny of the appropriate genotype and subjected to clinical surveillance.
References
Central Brain Tumor Registry of the United States Statistical report: primary brain tumors in the United States, 1995–1999. https:// http://www.cbtrus.org/reports/2002/2002report.pdf (CBTRUS, 2006)
Taylor, M. D. et al. Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol. 123, 465–472 (2012)
Schüller, U. et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh-induced medulloblastoma. Cancer Cell 14, 123–134 (2008)
Yang, Z. J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145 (2008)
Gibson, P. et al. Subtypes of medulloblastoma have distinct developmental origins. Nature 468, 1095–1099 (2010)
Kawauchi, D. et al. A mouse model of the most aggressive subgroup of human medulloblastoma. Cancer Cell 21, 168–180 (2012)
Mulhern, R. K. et al. Neurocognitive consequences of risk-adapted therapy for childhood medulloblastoma. J. Clin. Oncol. 23, 5511–5519 (2005)
Wang, J. et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nature Methods 8, 652–654 (2011)
Rausch, T. et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148, 59–71 (2012)
Castellino, R. C. et al. Heterozygosity for Pten promotes tumorigenesis in a mouse model of medulloblastoma. PLoS ONE 5, e10849 (2010)
Hahn, H. et al. Mutations of the human homolog of Drosophila patched in the nevoid basal cell carcinoma syndrome. Cell 85, 841–851 (1996)
Malkin, D. et al. Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science 250, 1233–1238 (1990)
Hamilton, S. R. et al. The molecular basis of Turcot’s syndrome. N. Engl. J. Med. 332, 839–847 (1995)
Taylor, M. D. et al. Medulloblastoma in a child with Rubenstein-Taybi syndrome: case report and review of the literature. Pediatr. Neurosurg. 35, 235–238 (2001)
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007)
Cao, R. et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298, 1039–1043 (2002)
Czermin, B. et al. Drosophila Enhancer of Zeste/ESC complexes have a histone H3 methyltransferase activity that marks chromosomal Polycomb sites. Cell 111, 185–196 (2002)
Agger, K. et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 (2007)
Schnetz, M. P. et al. Genomic distribution of CHD7 on chromatin tracks H3K4 methylation patterns. Genome Res. 19, 590–601 (2009)
Sauvageau, M. & Sauvageau, G. Polycomb group proteins: multi-faceted regulators of somatic stem cells and cancer. Cell Stem Cell 7, 299–313 (2010)
Morin, R. D. et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nature Genet. 42, 181–185 (2010)
Kleer, C. G. et al. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc. Natl Acad. Sci. USA 100, 11606–11611 (2003)
Varambally, S. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002)
van Haaften, G. et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nature Genet. 41, 521–523 (2009)
Yang, F., Babak, T., Shendure, J. & Disteche, C. M. Global survey of escape from X inactivation by RNA-sequencing in mouse. Genome Res. 20, 614–622 (2010)
Christensen, J. et al. RBP2 belongs to a family of demethylases, specific for tri- and dimethylated lysine 4 on histone 3. Cell 128, 1063–1076 (2007)
Lee, M. G., Wynder, C., Cooch, N. & Shiekhattar, R. An essential role for CoREST in nucleosomal histone 3 lysine 4 demethylation. Nature 437, 432–435 (2005)
Mosimann, C., Hausmann, G. & Basler, K. β-Catenin hits chromatin: regulation of Wnt target gene activation. Nature Rev. Mol. Cell Biol. 10, 276–286 (2009)
Hecht, A., Vleminckx, K., Stemmler, M. P., van Roy, F. & Kemler, R. The p300/CBP acetyltransferases function as transcriptional coactivators of β-catenin in vertebrates. EMBO J. 19, 1839–1850 (2000)
Barker, N. et al. The chromatin remodelling factor Brg-1 interacts with β-catenin to promote target gene activation. EMBO J. 20, 4935–4943 (2001)
Carrera, I., Janody, F., Leeds, N., Duveau, F. & Treisman, J. E. Pygopus activates Wingless target gene transcription through the mediator complex subunits Med12 and Med13. Proc. Natl Acad. Sci. USA 105, 6644–6649 (2008)
Thompson, M. C. et al. Genomics identifies medulloblastoma subgroups that are enriched for specific genetic alterations. J. Clin. Oncol. 24, 1924–1931 (2006)
Kool, M. et al. Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PLoS ONE 3, e3088 (2008)
Orsulic, S., Huber, O., Aberle, H., Arnold, S. & Kemler, R. E-cadherin binding prevents β-catenin nuclear localization and β-catenin/LEF-1-mediated transactivation. J. Cell Sci. 112, 1237–1245 (1999)
Risinger, J. I., Berchuck, A., Kohler, M. F. & Boyd, J. Mutations of the E-cadherin gene in human gynecologic cancers. Nature Genet. 7, 98–102 (1994)
Becker, K.-F. et al. E-Cadherin gene mutations provide clues to diffuse type gastric carcinomas. Cancer Res. 54, 3845–3852 (1994)
Pek, J. W. & Kai, T. DEAD-box RNA helicase Belle/DDX3 and the RNA interference pathway promote mitotic chromosome segregation. Proc. Natl Acad. Sci. USA 108, 12007–12012 (2011)
Lai, M. C., Chang, W. C., Shieh, S. Y. & Tarn, W. Y. DDX3 regulates cell growth through translational control of cyclin E1. Mol. Cell. Biol. 30, 5444–5453 (2010)
Schröder, M. Human DEAD-box protein 3 has multiple functions in gene regulation and cell cycle control and is a prime target for viral manipulation. Biochem. Pharmacol. 79, 297–306 (2010)
Samuels, Y. et al. High frequency of mutations of the PIK3CA gene in human cancers. Science 304, 554 (2004)
Broderick, D. K. et al. Mutations of PIK3CA in anaplastic oligodendrogliomas, high-grade astrocytomas, and medulloblastomas. Cancer Res. 64, 5048–5050 (2004)
Parsons, D. W. et al. The genetic landscape of the childhood cancer medulloblastoma. Science 331, 435–439 (2011)
Andäng, M. et al. Histone H2AX-dependent GABAA receptor regulation of stem cell proliferation. Nature 451, 460–464 (2008)
Pasini, D. et al. Coordinated regulation of transcriptional repression by the RBP2 H3K4 demethylase and Polycomb-Repressive Complex 2. Genes Dev. 22, 1345–1355 (2008)
Khan, A., Shover, W. & Goodliffe, J. M. Su(z)2 antagonizes auto-repression of Myc in Drosophila, increasing Myc levels and subsequent trans-activation. PLoS ONE 4, e5076 (2009)
Northcott, P. A. et al. Medulloblastoma comprises four distinct molecular variants. J. Clin. Oncol. 29, 1408–1414 (2011)
Spatz, A., Borg, C. & Feunteun, J. X-chromosome genetics and human cancer. Nature Rev. Cancer 4, 617–629 (2004)
Lindqvist, L. et al. Selective pharmacological targeting of a DEAD box RNA helicase. PLoS One 3, e1583 (2008)
Engelman, J. A. Targeting PI3K signalling in cancer: opportunities, challenges and limitations. Nature Rev. Cancer 9, 550–562 (2009)
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012)
Acknowledgements
This research was supported as part of the St Jude Children’s Research Hospital, Washington University Pediatric Cancer Genome Project. This work was supported by grants from the National Institutes of Health (R01CA129541, P01CA96832 and P30CA021765; R.J.G.), the Collaborative Ependymoma Research Network (CERN), Musicians against Childhood Cancer (MACC), The Noyes Brain Tumour Foundation, and by the American Lebanese Syrian Associated Charities (ALSAC). We are grateful to S. Temple for the gift of reagents and the staff of the Hartwell Center for Bioinformatics and Biotechnology and ARC at St Jude Children’s Research Hospital for technical assistance.
Author information
Authors and Affiliations
Contributions
G.R., M.P., T.A.K., C.L., X.C., L.D., T.N.P., E.H., L.W., X.Z., N.Ch., R.H., N.Cu., R.T., J.W., G.W., M.R., X.H., J.B., P.G., J.M., J.E., B.V., A.O.-T., T.L., S.Po., S.Pa., D.Z., D.K. and D.F. contributed to the design and conduct of experiments and to the writing. S.J.B., R.K., M.F.R., R.S.F., L.L.F., D.J.D., K.O. and E.R.M. contributed to experimental design and to the writing. A.G., D.W.E., C.C.L., E.B., T.H., S.G. and R.C. provided clinical expertise. R.K.W., J.R.D., J.Z. and R.J.G. conceived the research and contributed to the design, direction and reporting of the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Results and Supplementary References – see Table of Contents for details. (PDF 5466 kb)
Supplementary Tables 1
This file contains the data for Supplementary Tables 1-4. (XLS 1830 kb)
Supplementary Tables 2
This file contains the data for Supplementary Tables 5-7. (XLS 7933 kb)
Supplementary Tables 3
This file contains the data for Supplementary Table 8. (XLS 48 kb)
Supplementary Tables 4
This file contains the data for Supplementary Tables 9-15. (XLS 209 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
Robinson, G., Parker, M., Kranenburg, T. et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 488, 43–48 (2012). https://doi.org/10.1038/nature11213
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11213
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.