Abstract
To correlate the variable clinical features of oestrogen-receptor-positive breast cancer with somatic alterations, we studied pretreatment tumour biopsies accrued from patients in two studies of neoadjuvant aromatase inhibitor therapy by massively parallel sequencing and analysis. Eighteen significantly mutated genes were identified, including five genes (RUNX1, CBFB, MYH9, MLL3 and SF3B1) previously linked to haematopoietic disorders. Mutant MAP3K1 was associated with luminal A status, low-grade histology and low proliferation rates, whereas mutant TP53 was associated with the opposite pattern. Moreover, mutant GATA3 correlated with suppression of proliferation upon aromatase inhibitor treatment. Pathway analysis demonstrated that mutations in MAP2K4, a MAP3K1 substrate, produced similar perturbations as MAP3K1 loss. Distinct phenotypes in oestrogen-receptor-positive breast cancer are associated with specific patterns of somatic mutations that map into cellular pathways linked to tumour biology, but most recurrent mutations are relatively infrequent. Prospective clinical trials based on these findings will require comprehensive genome sequencing.
Similar content being viewed by others
Main
Oestrogen-receptor-positive breast cancer exhibits highly variable prognosis, histological growth patterns and treatment outcomes. Neoadjuvant aromatase inhibitor treatment trials provide an opportunity to document oestrogen-receptor-positive breast cancer phenotypes in a setting where sample acquisition is easy, prospective consent for genomic analysis can be obtained and responsiveness to oestrogen deprivation therapy is documented1. We therefore conducted massively parallel sequencing (MPS) on 77 samples accrued from two neoadjuvant aromatase inhibitor clinical trials2,3. Forty-six cases underwent whole-genome sequencing (WGS) and 31 cases underwent exome sequencing, followed by extensive analysis for somatic alterations and their association with aromatase inhibitor response. Case selection for discovery was based on the levels of the tumour proliferation marker Ki67 in the surgical specimen, because high cellular proliferation despite aromatase inhibitor treatment identifies poor prognosis tumours exhibiting oestrogen-independent growth4 (Supplementary Fig. 1). Twenty-nine samples had Ki67 levels above 10% (‘aromatase-inhibitor-resistant tumours’, median Ki67 21%, range 10.3–80%) and 48 were at or below 10% (‘aromatase-inhibitor-sensitive tumours’, median Ki67 1.2%, range 0–8%). Cases were also classified as luminal A or B by gene expression profiling3. We subsequently examined interactions between Ki67 biomarker change, histological categories, intrinsic subtype and mutation status in selected recurrently mutated genes in 310 cases overall. Pathway analysis was applied to contrast the signalling perturbations in aromatase-inhibitor-sensitive versus aromatase-inhibitor-resistant tumours.
Results
The mutation landscape of luminal-type breast cancer
Using paired-end MPS, 46 tumour and normal genomes were sequenced to at least 30-fold and 25-fold haploid coverage, respectively, with diploid coverage of at least 95% based on concordance with SNP array data (Supplementary Table 1). Candidate somatic events were identified using multiple algorithms5,6, and were then verified by hybridization capture-based validation that targeted all putative somatic single-nucleotide variants (SNVs) and small insertions/deletions (indels) that overlap coding exons, splice sites and RNA genes (tier 1), high-confidence SNVs and indels in non-coding conserved or regulatory regions (tier 2), as well as non-repetitive regions of the human genome (tier 3). In addition, somatic structural variants and germline structural variants that potentially affect coding sequences (Supplementary Information) were assessed. Digital sequencing data from captured target DNAs from the 46 tumour and normal pairs (Supplementary Table 2 and Supplementary Information) confirmed 81,858 mutations (point mutations and indels) and 773 somatic structural variants. The average numbers of somatic mutations and structural variants were 1,780 (range 44–11,619) and 16.8 (range 0–178) per case, respectively (Supplementary Table 3). Tier 1 point mutations and small indels predicted for all 46 cases also were validated using both 454 and Illumina sequencing (Supplementary Information). BRC25 was a clear outlier with only 44 validated tiers 1–3 mutations, all at low allele frequencies (ranging from 5% to 26.8%). This sample probably had low tumour content despite histopathology assessment, but the data are included to avoid bias.
The overall mutation rate was 1.18 validated mutations per megabase (Mb) (tier 1: 1.05; tier 2: 1.14; tier 3: 1.20). The mutation rate for tier 1 was higher than that observed for acute myeloid leukaemia (0.18–0.23)6,7, but lower than that reported for hepatocellular carcinoma (1.85)8, malignant melanoma (6.65)9 and lung cancers (3.05–8.93)10,11 (Supplementary Table 4). The background mutation rate (BMR) across the 21 aromatase-inhibitor-resistant tumours was 1.62 per Mb, nearly twice that of the 25 aromatase-inhibitor-sensitive tumours at 0.824 per Mb (P = 0.02, one-sided t-test). A trend for more somatic structural variations in the aromatase-inhibitor-resistant group was also observed, as the validated somatic structural variation frequency in the 21 aromatase-inhibitor-resistant tumour genomes was 21.69 versus an average of 12.76 in 25 aromatase-inhibitor-sensitive tumours (P = 0.16, one-sided t-test) (Fig. 1). If ten TP53 mutated cases were excluded, the background mutation rate still tended to be higher in the aromatase-inhibitor-resistant group (P = 0.08). To demonstrate that a single-tumour core biopsy produced representative genomic data, whole-genome sequencing of two pre-treatment biopsies was conducted for 5 of the 46 cases. The frequency of mutations in the paired specimens showed high concordance in all cases (correlation coefficiency ranged from 0.74 to 0.95) (Supplementary Fig. 2) and a somatic mutation was infrequently detected in only one of the two samples (4.65% overall).
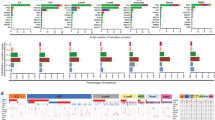
Circos plots44 indicate validated somatic mutations comprising tier 1 point mutations and indels, genome-wide copy number alterations, and structural rearrangements in six representative genomes. Three on-treatment Ki67 less than or at 10% (top panel: BRC15, BRC17 and BRC22) and three on-treatment Ki67 greater than 10% (bottom panel: BRC44, BRC47 and BRC50) cases are shown. Significantly mutated genes are highlighted in red. No purity-based copy number corrections were used for plotting copy number.
Significantly mutated genes in luminal breast cancer
The discovery effort was extended by studying 31 additional cases by exome sequencing, producing an additional 1,371 tier 1 mutations. In total the 77 cases yielded 3,355 tier 1 somatic mutations, including 3,208 point mutations, 1 dinucleotide mutation and 146 indels, ranging from 1 to 28 nucleotides. The point mutations included 733 silent, 2,145 missense, 178 nonsense, 6 read-through, 69 splice-site mutations and 77 in RNA genes (Supplementary Table 5). Of 2,145 missense mutations, 1,551 were predicted to be deleterious by SIFT12 and/or PolyPhen13. The MuSiC package45 was applied to determine the significance of the difference between observed versus expected mutation events in each gene, on the basis of the background mutation rate. This identified 18 significantly mutated genes with a convolution false discovery rate (FDR) < 0.26 (Table 1 and Supplementary Table 6). The list contains genes previously identified as mutated in breast cancer (PIK3CA14, TP5315, GATA312, CDH113, RB116, MLL317, MAP3K118 and CDKN1B19) as well as genes not previously observed in clinical breast cancer samples, including TBX3, RUNX1, LDLRAP1, STNM2, MYH9, AGTR2, STMN2, SF3B1 and CBFB.
Thirteen mutations (3 nonsense, 6 frame-shift indels, 2 in-frame deletions and 2 missense) were identified in MAP3K1 (Table 1 and Fig. 2), a serine/threonine kinase that activates the ERK and JNK kinase pathways through phosphorylation of MAP2K1 and MAP2K4 (ref. 20). Of interest, a missense (S184L) and a splice-region mutation (e2+3 probably affecting splicing) in MAP2K4 were observed in two tumours with no MAP3K1 mutation (Fig. 2). Single nonsynonymous mutations in MAP3K12, MAP3K4, MAP4K3, MAP4K4, MAPK15 and MAPK3 were also detected (Supplementary Table 5). TBX3 harboured three small indels (one insertion and two deletions). TBX3 affects expansion of breast cancer stem-like cells through regulation of FGFR21. Two truncating mutations in the tumour suppressor CDKN1B were identified19. Four missense RUNX1 mutations were observed, with three in the RUNT domain clustered within the 8 amino acid putative ATP-binding site (R166Q, G168E and R169K). RUNX1 is a transcription factor affected by mutation and translocation in the M2 subtype of acute myeloid leukaemia22 and is implicated in tethering the oestrogen receptor to promoters independently of oestrogen response elements23. Two mutations (N104S and N140*) were also identified in CBFB, the binding partner of RUNX1. Additional mutations included 3 missense (2 K700E and 1 K666Q), in SF3B1, a splicing factor implicated in myelodysplasia24 and chronic lymphocytic leukaemia25. One missense mutation, one nonsense mutation and two indels were found in the MYH9 gene, involved in hereditary macrothrombocytopenia26 as well as being observed in an ALK translocation in anaplastic large cell lymphoma27.
Somatic status of all mutations was obtained by Sanger sequencing of PCR products or Illumina sequencing of targeted capture products. The locations of conserved protein domains are highlighted. Each nonsynonymous substitution, splice site mutation or indel is designated with a circle at the representative protein position with colour to indicate translation effects of the mutation. Asterisk, nonsense mutations that cause truncation of the open reading frame.
We also identified three significantly mutated genes (LDLRAP1, AGTR2 and STMN2) not previously implicated in cancer. A missense and a nonsense mutation were observed in LDLRAP1, a gene associated with familial hypercholesterolaemia28. AGTR2, angiotensin II receptor type 2, harboured two missense mutations (V184I and R251H). Angiotensin signalling and oestrogen receptor intersect in models of tissue fibrosis29. STMN2, a gene activated by JNK family kinases30,31 and therefore regulated by MAP3K1 and MAP2K4, harboured one frameshift deletion and one missense mutation. Three deletions and one point mutation (Supplementary Fig. 3) were identified in a large, infrequently spliced non-coding (lnc) RNA gene, MALAT1 (metastasis associated lung adenocarcinoma transcript 1), that regulates alternative splicing by modulating the phosphorylation of SR splicing factor32. Translocations and point mutations of MALAT1 have been reported in sarcoma33 and colorectal cancer cell lines34. Five additional MALAT1 mutations were found in the recurrent screening set (Supplementary Table 5d). The locations of these mutations clustered in a region of species homology (F1 and 2 domains) that could mediate interactions with SRSF1 (ref. 32, Supplementary Fig. 4). Non-coding mutation clusters were found in ATR, GPR126 and NRG3 (Supplementary Information and Supplementary Table 7).
Correlating mutations with clinical data
To study clinical correlations, mutation recurrence screening was conducted on an additional 240 cases (Supplementary Table 8 and Supplementary Fig. 1). By combining WGS, exome and recurrence screening data, we determined the mutation frequency in PIK3CA to be 41.3% (131 of 317 tumours) (Supplementary Table 5a–d and Supplementary Fig. 3). TP53 was mutated in 51 of 317 tumours (16.1%) (Supplementary Table 5a–d and Supplementary Fig. 3). Additionally, 52 nonsynonymous MAP3K1 mutations in 39 tumours and 10 mutations in its substrate MAP2K4 were observed, representing a combined case frequency of 15.5% (Supplementary Table 5a–d and Fig. 3). Of note, 52 of the 62 non-silent mutations in MAP3K1 and MAP2K4 were scattered indels or other protein-truncating events strongly suggesting functional inactivation. In addition, 13 tumours harboured two non-silent MAP3K1 mutations, indicative of bi-allelic loss and reinforcing the conclusion that this gene is a tumour suppressor. Twenty nine tumours harboured a total of 30 mutations in GATA3, consisting of 25 truncation events, one in-frame insertion, and 4 missense mutations including 3 recurrent mutations at M294K (Supplementary Table 5a–d and Supplementary Fig. 3). BRC8 harboured a chromosome 10 deletion that includes GATA3. CDH1 mutation data were available for 169 samples and, as expected, its mutation status was strongly associated with lobular breast cancer13 (Table 2a). We applied a permutation-based approach in MuSiC45 to ascertain relationships between mutated genes. Negative correlations were found between mutations in gene pairs such as GATA3 and PIK3CA (P = 0.0026), CDH1 and GATA3 (P = 0.015), and CDH1 and TP53 (P = 0.022). MAP3K1 and MAP2K4 mutations were mutually exclusive, albeit without reaching statistical significance (P = 0.3). In contrast, a positive correlation between MAP3K1/MAP2K4 and PIK3CA mutations was highly significant (P = 0.0002) (Supplementary Table 9).
One MAP3K1 deletion in BRC49 and one MAP2K4 deletion in BRC47, and one ELP3-NRG1 fusion in BRC49 identified using Illumina paired-end reads from whole-genome sequence data. Arcs represent multiple breakpoint-spanning read pairs with sequence coverage depth plotted in black across the region. Chr, chromosome.
Two independent mutation data sets, designated ‘Set 1’ (discovery cohort) and ‘Set 2’ (validation cohort), from these clinical trial samples were analysed separately and then in combination, with a false discovery rate (FDR)-corrected P value to gauge the overall strength and consistency of genotype–phenotype relationships (Table 2a, b and Supplementary Fig. 1). TP53 mutations in both data sets correlated with significantly higher Ki67 levels, both at baseline (P = 0.0003) and at surgery (P = 0.001). Furthermore, TP53 mutations were significantly enriched in luminal B tumours (P = 0.04) and in higher histological grade tumours (P = 0.02). In contrast, MAP3K1 mutations were more frequent in luminal A tumours (P = 0.02), in grade 1 tumours (P = 0.005) and in tumours with lower Ki67 at baseline (P = 0.001) with consistent findings across both data sets. GATA3 mutation did not influence baseline Ki67 levels but was enriched in samples exhibiting greater percentage Ki67 decline (P = 0.01). This finding requires further verification because it was significant in Set 1 (uncorrected P value 0.003) but was a marginal finding in Set 2 (P = 0.08). However, it suggests GATA3 mutation may be a positive predictive marker for aromatase inhibitor response.
Structural variation and DNA repair mechanisms
Analysis of copy number alterations (CNAs) revealed arm-level gains for 1q, 5p, 8q, 16p, 17q, 20p and 20q and arm-level losses for 1p, 8p, 16q, and 17p in the 46 WGS tumour genomes (Supplementary Fig. 5). A total of 773 structural variants (579 deletions, 189 translocations and 5 inversions) identified by WGS were validated as somatic in 46 breast cancer genomes by capture validation. No recurrent translocations were detected but six in-frame fusion genes were validated by reverse transcription followed by PCR (Supplementary Information and Supplementary Tables 10–13). Seven tumours had multiple complex translocations with breakpoints suggestive of a catastrophic mitotic event (‘chromothripsis’; Supplementary Table 11). Analysis of the structural variant genomic breakpoints shows the spectra of putative chromothripsis-related events are the same as seen for other somatic events, with the majority of structural variants arising from non-homologous end-joining. We classified somatic (mitotic) and germline (meiotic) structural variants into four groups: variable number tandem repeat (VNTR), non-allelic homologous recombination (NAHR), microhomology-mediated end joining (MMEJ), and non-homologous end joining (NHEJ), according to criteria described in Supplementary Information. The fraction of each classification is shown for germline and somatic (mitotic) events (Supplementary Table 14). There were significantly more somatic NHEJ events in tumour genomes than the other three types (P < 2.2 × 10−16).
Pathways relevant to aromatase inhibitor response
Pathscan35 analysis (Supplementary Table 15 and Supplementary Information) indicated that somatic mutations detected in the 77 discovery cases affect a number of pathways, including caspase cascade/apoptosis, ErbB signalling, Akt/PI3K/mTOR signalling, TP53/RB signalling and MAPK/JNK pathways (Fig. 4a). To discern the pathways relevant to aromatase inhibitor sensitivity, we conducted separate pathway analyses for aromatase-inhibitor-sensitive versus aromatase-inhibitor-resistant tumours. Whereas the majority of top altered pathways (FDR ≤ 0.15) in each group are shared, several pathways were enriched in the aromatase-inhibitor-resistant group, including the TP53 signalling pathway, DNA replication, and mismatch repair. Specifically, 38% of the aromatase-inhibitor-resistant group (11 of 29 tumours) have mutations in the TP53 pathway with three having double or triple hits involving TP53, ATR, APAF1 or THBS1. In contrast, only 16.6% (8 of 48 tumours) of the Ki67 low group had mutations in the TP53 signalling pathway, each with only a single hit in genes TP53, ATR, CCNE2 or IGF1. (Supplementary Table 16).
a, Only genetic alterations identified in 46 WGS cases are shown. Alterations were discovered in key genes in the TP53/RB, MAPK, PI3K/AKT/mTOR pathways. Genes coloured blue and red are predicted to be functionally inactivated and activated, respectively, through focused mutations including point mutations and small indels (M), copy number deletions (C), or other structural changes (S) that affect the gene. The inter-connectedness of this network (several pathways) shows that there are many different ways to perturb a pathway. b, Eight interaction networks from canonical maps are significantly over-represented by mutations in 77 luminal breast tumours (46 WGS and 31 exome cases). In the concentric circle diagram, tumours are arranged as radial spokes and categorized by their mutation status in each network (concentric ring colour) and significantly mutated gene mutation status (black dots). Tumour classification by pathway analysis shows many tumours unaffected by a given significantly mutated gene often harbour other mutations in the same network. For full annotation, see Supplementary Information and Supplementary Fig. 6. PLC, phospholipase C; SMG, significantly mutated gene.
GeneGo pathway analysis of MetaCore interacting network objects was used to identify genes in the 77 luminal breast cancers with low-frequency mutations that cluster into pathway maps. Eight networks assembled from significant maps encompassed mutations from 71 (92%) of the tumours (Fig. 4b). Many of the network objects shared pathways with significantly mutated genes such as TP53, MAP3K1, PIK3CA and CDH1. GeneGo analysis also revealed that several genes with low-frequency mutations were actually subunits of complexes, resulting in higher mutation rates for that object, for example, the condensin complex (4 mutations in 4 genes) and the MRN complex (4 mutations in 3 genes). Several pathways without multiple significantly mutated genes, such as the apoptotic cascade, calcium/phospholipase signalling and G-protein-coupled receptors, were significantly affected by low-frequency mutations. Grouping tumours by significantly mutated genes and pathway mutation status showed that whereas 55 (71%) of the tumours contained significantly mutated genes in significant pathways, an additional 16 (21%) contained only non-significantly mutated genes in these pathways. Thus, tumours without a given significantly mutated gene often had other mutations in the same relevant pathway (Fig. 4b, Supplementary Fig. 6, Supplementary Table 17 and Supplementary Information).
We also applied PARADIGM36 to infer pathway-informed gene activities using gene expression and copy-number data to identify several ‘hubs’ of activity (Supplementary Fig. 7, Supplementary Fig. 8 and Supplementary Information). As expected, ESR1 and FOXA1 were among the hubs activated cohort-wide while other hubs exhibited high but differential changes in aromatase-inhibitor-resistant tumours including MYC, FOXM1 and MYB (Supplementary Fig. 8). The concordance among the 104 MetaCore maps from GeneGo analysis described above is significant, with 75 (72%) matching one of the PARADIGM subnetworks at the 0.05 significance level after multiple test correction (P < 4.4 × 10−6; Bonferroni-adjusted hypergeometric test) (Supplementary Fig. 9). We identified significant subnetworks associated with Ki67 biomarker status (Supplementary Fig. 10 and Supplementary Information) involving transcription factors controlling large regulons.
The PARADIGM-inferred pathway signatures were further used to derive a map of the genetic mechanisms that may underlie treatment response. A subnetwork was constructed in which interactions were retained only if they connected two features with higher than average absolute association with Ki67 biomarker status (Supplementary Figs 10 and 11 and Supplementary Information). Consistent with the PathScan results, among the largest of the hubs in the identified network were a central DNA damage hub with the second highest connectivity (55 regulatory interactions; 1% of the network) and TP53 with the 14th highest connectivity (26 connections; 0.5% of the network). Additional highly connected hubs identified in order of connectivity were MYC with 79 connections (1.4%), FYN with 45 (0.8%), MAPK3 with 43, JUN with 40, HDAC1 with 40, SHC1 with 39, and HIF1A/ARNT complex with 39 (Supplementary Fig. 11).
To identify higher-level connections between mutations and clinical features, we compared the samples on the basis of pathway-derived signatures. For each clinical attribute and each significantly mutated gene, we dichotomized the discovery samples into a positive and a negative group to derive pathway signatures that discriminated between the groups (see details in Supplementary Information). We then computed all pair-wise Pearson correlations between pathway signatures and clustered the resulting correlations (Fig. 5). The entire process was repeated using validated mutations and signatures derived from the validation set (Supplementary Fig. 12). In line with expectation, PIK3CA, MAP3K1, MAP2K4, and low risk preoperative endocrine prognostic index (PEPI) scores (PEPI is an index of recurrence risk post neoadjuvant aromatase inhibitor therapy4) cluster with the luminal A subtypes and with each other, and are supported by the validation set analysis. The luminal B-like signatures included TP53, RB1, RUNX1 and MALAT1, which also associated with other poor outcome features such as high baseline and surgical Ki67 levels, high grade histology and high PEPI scores. The TP53 and MALAT1 associations in the discovery set also were supported by the validation set analysis.
PARADIGM-based pathway signatures were derived for tumour feature dichotomies including mutation driven gene signatures (mutant versus non-mutant), histopathology type (lobular versus ductal), preoperative endocrine prognostic index (PEPI) score (PEPI = 0 favourable versus PEPI >0 unfavourable), PAM50 (50-gene intrinsic breast cancer subtype classifier) luminal A subtype (luminal A versus luminal B) and the reverse (luminal B versus luminal A), histopathology grade (grades II and III versus I), baseline Ki67 levels (≥ 14% versus < 14%), and end-of-treatment Ki67 levels (≥ 10% versus < 10%) and overall PEPI score (higher than mean unfavourable versus lower than mean favourable). Pearson correlations were computed between all pair-wise signatures; positive correlations, red; negative correlations, blue; column features ordered identically as rows. Correlation analysis on the 77 samples in the discovery set is shown. Asterisk: Ki67 < 2.7%, oestrogen-receptor-positive, node negative and tumour size ≤ 5 cm.
Druggable gene analysis
We defined mutations in druggable tyrosine kinase domains including in ERBB2 (a V777L and a 755–759LRENT in-frame deletion homologous to gefitinib-sensitizing EGFR mutations in lung cancer37), as well as in DDR1 (A829V, R611C), DDR2 (E583D), CSF1R (D735H, M875L), and PDGFRA (E924K). In addition, pleckstrin homology domain mutations were observed in AKT1 (C77F) and AKT2 (S11F) and a kinase domain mutation was identified in RPS6KB1 (S375F) (Supplementary Table 18).
Discussion
The low frequency of many significantly mutated genes presents an enormous challenge for correlative analysis, but several statistically significant patterns were identified, including the relationship between MAP3K1 mutation, luminal A subtype, low tumour grade and low Ki67 proliferation index. On this basis, for patients with MAP3K1 mutant luminal tumours, neoadjuvant aromatase inhibitor could provide a favourable option. In contrast, tumours with TP53 mutations, which are mostly aromatase inhibitor resistant, would be more appropriately treated with other modalities. MAP3K1 activates the ERK family, thus, loss of ERK signalling could explain the indolent nature of MAP3K1-deficient tumours20. However, MAP3K1 also activates JNK through MAP2K4, which also can be mutated38. Loss of JNK signalling produces a defect in apoptosis in response to stress, which would hypothetically explain why these mutations accumulate39,40. PIK3CA harboured the most mutations (41.3%) but was neither associated with clinical nor Ki67 response, confirming our earlier report41. However, the positive association between MAP3K1/MAP2K4 mutations and PIK3CA mutation at both the mutation and pathway levels suggests cooperativity (Fig. 4a).
The finding of multiple significantly mutated genes linked previously to benign and malignant haematopoietic disorders suggests that breast cancer, like leukaemia, can be viewed as a stem-cell disorder that produces indolent or aggressive tumours that display varying phenotypes depending on differentiation blocks generated by different mutation repertoires42. Whereas only MLL3 showed statistical significance in the analysis of 46 WGS cases, multiple mutations in genes related to histone modification and chromatin remodelling are worth noting (Supplementary Table 19). An array of coding mutations and structural variations was discovered in methyltransferases (MLL2, MLL3, MLL4 and MLL5), demethyltransferases (KDM6A, KDM4A, KDM5B and KDM5C), and acetyltransferases (MYST1, MYST3 and MYST4). Furthermore, our analysis identified several adenine-thymine (AT)-rich interactive domain-containing protein genes (ARID1A, ARID2, ARID3B and ARID4B) that harboured mutations and large deletions, reinforcing the role of members from the SNF/SWI family in breast cancer.
Pathway analysis enables the evaluation of mutations with low recurrence frequency where statistical comparisons are conventionally underpowered. For example, the eight samples with MAP2K4 mutations were sufficient to derive a reliable pathway-based gene signature in PARADIGM that aligns with MAP3K1. This approach also pointed to a putative connection between MALAT1 and the TP53 pathway. Finally, we provide evidence that transcriptional associations to Ki67 response reside in a connected network under the control of several key ‘hub’ genes including MYC, FYN and MAP kinases, among others. Targeting these hubs in resistant tumours could produce therapeutic advances. In conclusion, the genomic information derived from unbiased sequencing is a logical new starting point for clinical investigation, where the mutation status of an individual patient is determined in advance and treatment decisions are driven by therapeutic hypotheses that stem from knowledge of the genomic sequence and its possible consequences. However, the accrual of large numbers of patients and the use of comprehensive sequencing and gene expression approaches will be required because of the extreme genomic heterogeneity documented by this investigation.
Methods Summary
Clinical trial samples were accessed from the preoperative letrozole phase 2 study (NCT00084396)2 that investigated the effect of letrozole for 16 to 24 weeks on surgical outcomes and from the American College of Surgeons Oncology Group (ACOSOG) Z1031 study (NCT00265759)3 that compared anastrozole with exemestane or letrozole for 16 to 18 weeks before surgery (REMARK flow charts, Supplementary Fig. 1). Baseline snap-frozen biopsy samples with greater than 70% tumour content (by nuclei) underwent DNA extraction and were paired with a peripheral blood DNA sample. Two formalin-fixed biopsies were obtained at baseline and at surgery, and were used to conduct oestrogen receptor and Ki67 immunohistochemistry as previously published4. Paired end Illumina reads from tumours and normal samples were aligned to NCBI build36 using BWA. Somatic point mutations were identified using SomaticSniper43, and indels were identified by combining results from a modified version of the Samtools indel caller (http://samtools.sourceforge.net/), GATK and Pindel. Structural variations were identified using BreakDancer5 and SquareDancer (unpublished). All putative somatic events found in 46 cases were validated by targeted custom capture arrays (Nimblegen)/Illumina sequencing and all tier 1 mutations for 46 WGS cases also were validated using PCR/454 sequencing. All statistical analyses, including significantly mutated gene, mutation relation and clinical correlation were done using the MuSiC package45 and/or by standard statistical tests (Supplementary Information). Pathway analysis was performed with PathScan, GeneGo Metacore (http://www.genego.com/metacore.php) and PARADIGM. A complete description of the materials and methods used to generate this data set and results is provided in the Supplementary Methods section.
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
InformationDNA sequence data are deposited in the restricted access portal at dbGaP, accession number phs000472.v1.p1. Gene expression array data used in the Paradigm training set is deposited in GEO, accession number GSE29442, and a Superseries that covers both the Agilent gene expression data and the Agilent array CGH data used for the Paradigm test set is deposited in GEO, accession number GSE35191.
References
Chia, Y. H., Ellis, M. J. & Ma, C. X. Neoadjuvant endocrine therapy in primary breast cancer: indications and use as a research tool. Br. J. Cancer 103, 759–764 (2010)
Olson, J. A., Jr et al. Improved surgical outcomes for breast cancer patients receiving neoadjuvant aromatase inhibitor therapy: results from a multicenter phase II trial. J. Am. Coll. Surg. 208, 906–914; discussion. 915–906 (2009)
Ellis, M. J. et al. Randomized phase II neoadjuvant comparison between letrozole, anastrozole, and exemestane for postmenopausal women with estrogen receptor-rich stage 2 to 3 breast cancer: clinical and biomarker outcomes and predictive value of the baseline PAM50-based intrinsic subtype—ACOSOG Z1031. J. Clin. Oncol. 29, 2342–2349 (2011)
Ellis, M. J. et al. Outcome prediction for estrogen receptor-positive breast cancer based on postneoadjuvant endocrine therapy tumor characteristics. J. Natl. Cancer Inst. 100, 1380–1388 (2008)
Chen, K. et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nature Methods 6, 677–681 (2009)
Mardis, E. R. et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N. Engl. J. Med. 361, 1058–1066 (2009)
Ley, T. J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, 66–72 (2008)
Totoki, Y. et al. High-resolution characterization of a hepatocellular carcinoma genome. Nature Genet. 43, 464–469 (2011)
Pleasance, E. D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, 191–196 (2010)
Pleasance, E. D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, 184–190 (2010)
Lee, W. et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature 465, 473–477 (2010)
Usary, J. et al. Mutation of GATA3 in human breast tumors. Oncogene 23, 7669–7678 (2004)
Berx, G. et al. E-cadherin is a tumour/invasion suppressor gene mutated in human lobular breast cancers. EMBO J. 14, 6107–6115 (1995)
Samuels, Y. et al. High frequency of mutations of the PIK3CA gene in human cancers. Science 304, 554 (2004)
Prosser, J., Thompson, A. M., Cranston, G. & Evans, H. J. Evidence that p53 behaves as a tumour suppressor gene in sporadic breast tumours. Oncogene 5, 1573–1579 (1990)
T’Ang, A., Varley, J. M., Chakraborty, S., Murphree, A. L. & Fung, Y. K. Structural rearrangement of the retinoblastoma gene in human breast carcinoma. Science 242, 263–266 (1988)
Wang, X. X. et al. Somatic mutations of the mixed-lineage leukemia 3 (MLL3) gene in primary breast cancers. Pathol. Oncol. Res. 17, 429–433 (2011)
Kan, Z. et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466, 869–873 (2010)
Spirin, K. S. et al. p27/Kip1 mutation found in breast cancer. Cancer Res. 56, 2400–2404 (1996)
Fanger, G. R., Johnson, N. L. & Johnson, G. L. MEK kinases are regulated by EGF and selectively interact with Rac/Cdc42. EMBO J. 16, 4961–4972 (1997)
Fillmore, C. M. et al. Estrogen expands breast cancer stem-like cells through paracrine FGF/Tbx3 signaling. Proc. Natl Acad. Sci. USA 107, 21737–21742 (2010)
Mao, S., Frank, R. C., Zhang, J., Miyazaki, Y. & Nimer, S. D. Functional and physical interactions between AML1 proteins and an ETS protein, MEF: implications for the pathogenesis of t(8;21)-positive leukemias. Mol. Cell. Biol. 19, 3635–3644 (1999)
Stender, J. D. et al. Genome-wide analysis of estrogen receptor α DNA binding and tethering mechanisms identifies Runx1 as a novel tethering factor in receptor-mediated transcriptional activation. Mol. Cell. Biol. 30, 3943–3955 (2010)
Papaemmanuil, E. et al. Somatic SF3B1 mutation in myelodysplasia with ring sideroblasts. N. Engl. J. Med. 365, 1384–1395 (2011)
Wang, L. et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N. Engl. J. Med. 365, 2497–2506 (2011)
Chen, Z. et al. The May-Hegglin anomaly gene MYH9 is a negative regulator of platelet biogenesis modulated by the Rho-ROCK pathway. Blood 110, 171–179 (2007)
Lamant, L. et al. Non-muscle myosin heavy chain (MYH9): a new partner fused to ALK in anaplastic large cell lymphoma. Genes Chromosom. Cancer 37, 427–432 (2003)
Wilund, K. R. et al. Molecular mechanisms of autosomal recessive hypercholesterolemia. Hum. Mol. Genet. 11, 3019–3030 (2002)
Dellê, H. et al. Antifibrotic effect of tamoxifen in a model of progressive renal disease. J. Am. Soc. Nephrol. 23, 37–48 (2012)
Tararuk, T. et al. JNK1 phosphorylation of SCG10 determines microtubule dynamics and axodendritic length. J. Cell Biol. 173, 265–277 (2006)
Westerlund, N. et al. Phosphorylation of SCG10/stathmin-2 determines multipolar stage exit and neuronal migration rate. Nature Neurosci. 14, 305–313 (2011)
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 39, 925–938 (2010)
Rajaram, V., Knezevich, S., Bove, K. E., Perry, A. & Pfeifer, J. D. DNA sequence of the translocation breakpoints in undifferentiated embryonal sarcoma arising in mesenchymal hamartoma of the liver harboring the t(11;19)(q11;q13.4) translocation. Genes Chromosom. Cancer 46, 508–513 (2007)
Xu, C., Yang, M., Tian, J., Wang, X. & Li, Z. MALAT-1: a long non-coding RNA and its important 3′ end functional motif in colorectal cancer metastasis. Int. J. Oncol. 39, 169–175 (2011)
Wendl, M. C. et al. PathScan: a tool for discerning mutational significance in groups of putative cancer genes. Bioinformatics 27, 1595–1602 (2011)
Vaske, C. J. et al. Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics 26, i237–i245 (2010)
Lynch, T. J. et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 350, 2129–2139 (2004)
Johnson, G. L. & Lapadat, R. Mitogen-activated protein kinase pathways mediated by ERK, JNK, and p38 protein kinases. Science 298, 1911–1912 (2002)
Widmann, C., Johnson, N. L., Gardner, A. M., Smith, R. J. & Johnson, G. L. Potentiation of apoptosis by low dose stress stimuli in cells expressing activated MEK kinase 1. Oncogene 15, 2439–2447 (1997)
Wagner, E. F. & Nebreda, A. R. Signal integration by JNK and p38 MAPK pathways in cancer development. Nature Rev. Cancer 9, 537–549 (2009)
Ellis, M. J. et al. Phosphatidyl-inositol-3-kinase alpha catalytic subunit mutation and response to neoadjuvant endocrine therapy for estrogen receptor positive breast cancer. Breast Cancer Res. Treat. 119, 379–390 (2010)
Prat, A. & Perou, C. M. Mammary development meets cancer genomics. Nature Med. 15, 842–844 (2009)
Larson, D. E. et al. SomaticSniper: identification of somatic point mutations in whole genome sequencing data. Bioinformatics 28, 311–317 (2011)
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009)
Dees, N. et al. MuSiC: Identifying mutational significance in cancer genomes. Genome Res (in the press)
Acknowledgements
This article is dedicated to the memory of Evelyn Lauder in recognition of her efforts to eradicate breast cancer. We would like to thank the participating patients and their families, clinical investigators and their support staffs, and J. A. Zujewski and the Cancer Therapy Evaluation Program at the US National Cancer Institute. We would like to acknowledge the efforts of the following people and groups at The Genome Institute for their contributions to this manuscript: the Analysis Pipeline group for developing the automated analysis pipelines that generated alignments and somatic variants; the LIMS group for developing tools to manage validation array ordering, capture and sequencing, and J. Veizer and H. Schmidt for structural variant and recurrent screening analyses. We thank the many members of the Siteman Cancer Center at Washington University in St Louis for support, and the committed members of the American College of Surgeons Oncology Group and their patients for contributing samples to the Z1031 trial. This work was funded by grants to R.K.W. from the National Human Genome Research Institute (NHGRI U54 HG003079), grants to M.J.E. from the National Cancer Institute (NCI R01 CA095614, NCI U01 CA114722), the Susan G. Komen Breast Cancer Foundation (BCTR0707808), and the Fashion Footwear Charitable Foundation, Inc., grant awards to ACOSOG included NCI U10 CA076001, the Breast Cancer Research Foundation, and clinical trial support from Novartis and Pfizer, and a Center grant (NCI P50 CA94056) to D.P.-W. We also acknowledge institutional support in the form of the Washington University Cancer Genome Initiative (R.K.W.), and a productive partnership with Illumina, Inc. The tissue procurement core was supported by an NCI core grant to the Siteman Cancer Center (NCI 3P50 CA68438). The BRIGHT Institute is supported in part by an ATT/Emerson gift to the Siteman Cancer Center.
Author information
Authors and Affiliations
Contributions
M.J.E. led the clinical investigations, biomarker analysis and chip-based genomics. E.R.M., M.J.E., L.D., R.S.F., T.J.L. and R.K.W. designed the experiments. L.D. and M.J.E. led data analysis. D.S., J.W.W., D.C.K., C.C.H., M.D.M., K.C., C.A.Mi., F.D., W.S.S., M.C.W., R.C. and C.K. performed data analysis. D.S., C.A.Ma., J.W.W., J.F.M., C.L. and L.D. prepared figures and tables. R.S.F., L.L.F., R.D., M.H., T.L.V., J.H., L.L., R.C. and J.S. performed laboratory experiments. L.E., G.U., J.M., G.V.B., P.K.M., J.M.G., M.L., K.H. and J.O. provided samples and clinical data. V.J.S., K.B., J.L., Y.T. and C.K. provided statistical and clinical correlation analysis. D.O. oversees the ACOSOG Operations Center that provides oversight and tracking for ACOSOG clinical trials. K.D., S.McD., D.C.A. and M.W. provided pathology analysis. B.A.V.T., J.W., R.J.G., A.E., D.P.-W., H.P.-W., J.M.S., T.C.G., S.N., C.K. and M.C.W. performed pathway analysis. L.-W.C. and R.B. analysed the druggable target mutation data. D.J.D. and B.O. provided informatics support. L.D., M.J.E. and E.R.M. wrote the manuscript. T.J.L., M.C.W. and R.K.W. critically read and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Materials and Methods, Supplementary Figures and Supplementary Figure legends 1-12, Supplementary Tables 1-20 and Supplementary References. (PDF 34070 kb)
Supplementary Data
This zipped file contains the Cytoscape data files. (ZIP 1022 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
Ellis, M., Ding, L., Shen, D. et al. Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature 486, 353–360 (2012). https://doi.org/10.1038/nature11143
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11143
This article is cited by
-
Studying the connection between SF3B1 and four types of cancer by analyzing networks constructed based on published research
Scientific Reports (2023)
-
Breast cancers with high proliferation and low ER-related signalling have poor prognosis and unique molecular features with implications for therapy
British Journal of Cancer (2023)
-
LncRNA MALAT1 signaling pathway and clinical applications in overcome on cancers metastasis
Clinical and Experimental Medicine (2023)
-
Mutational landscape of nasopharyngeal carcinoma based on targeted next-generation sequencing: implications for predicting clinical outcomes
Molecular Medicine (2022)
-
Heritable genomic diversity in breast cancer driver genes and associations with risk in a Chilean population
Biological Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.