Abstract
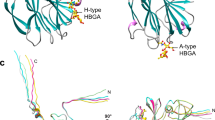
As with many other viruses, the initial cell attachment of rotaviruses, which are the major causative agent of infantile gastroenteritis, is mediated by interactions with specific cellular glycans1,2,3,4. The distally located VP8* domain of the rotavirus spike protein VP4 (ref. 5) mediates such interactions. The existing paradigm is that ‘sialidase-sensitive’ animal rotavirus strains bind to glycans with terminal sialic acid (Sia), whereas ‘sialidase-insensitive’ human rotavirus strains bind to glycans with internal Sia such as GM1 (ref. 3). Although the involvement of Sia in the animal strains is firmly supported by crystallographic studies1,3,6,7, it is not yet known how VP8* of human rotaviruses interacts with Sia and whether their cell attachment necessarily involves sialoglycans. Here we show that VP8* of a human rotavirus strain specifically recognizes A-type histo-blood group antigen (HBGA) using a glycan array screen comprised of 511 glycans, and that virus infectivity in HT-29 cells is abrogated by anti-A-type antibodies as well as significantly enhanced in Chinese hamster ovary cells genetically modified to express the A-type HBGA, providing a novel paradigm for initial cell attachment of human rotavirus. HBGAs are genetically determined glycoconjugates present in mucosal secretions, epithelia and on red blood cells8, and are recognized as susceptibility and cell attachment factors for gastric pathogens like Helicobacter pylori9 and noroviruses10. Our crystallographic studies show that the A-type HBGA binds to the human rotavirus VP8* at the same location as the Sia in the VP8* of animal rotavirus, and suggest how subtle changes within the same structural framework allow for such receptor switching. These results raise the possibility that host susceptibility to specific human rotavirus strains and pathogenesis are influenced by genetically controlled expression of different HBGAs among the world’s population.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The coordinates and structure factors for the P[14] VP8* structures are deposited in the Protein Data Bank under accession numbers 4DRR (apo), 4DRV (with A-type trisaccharide) and 4DS0 (with A-type tetrasaccharide). Raw glycan array data are available at http://www.functionalglycomics.org/glycomics/ublicdata/selectedScreens.jsp.
References
Blanchard, H., Yu, X., Coulson, B. S. & von Itzstein, M. Insight into host cell carbohydrate-recognition by human and porcine rotavirus from crystal structures of the virion spike associated carbohydrate-binding domain (VP8*). J. Mol. Biol. 367, 1215–1226 (2007)
Dormitzer, P. R. et al. Specificity and affinity of sialic acid binding by the rhesus rotavirus VP8* core. J. Virol. 76, 10512–10517 (2002)
Haselhorst, T. et al. Sialic acid dependence in rotavirus host cell invasion. Nature Chem. Biol. 5, 91–93 (2009)
Lopez, S. & Arias, C. F. Early steps in rotavirus cell entry. Curr. Top. Microbiol. Immunol. 309, 39–66 (2006)
Settembre, E. C., Chen, J. Z., Dormitzer, P. R., Grigorieff, N. & Harrison, S. C. Atomic model of an infectious rotavirus particle. EMBO J. 30, 408–416 (2011)
Dormitzer, P. R., Sun, Z. Y., Wagner, G. & Harrison, S. C. The rhesus rotavirus VP4 sialic acid binding domain has a galectin fold with a novel carbohydrate binding site. EMBO J. 21, 885–897 (2002)
Kraschnefski, M. J. et al. Effects on sialic acid recognition of amino acid mutations in the carbohydrate-binding cleft of the rotavirus spike protein. Glycobiology 19, 194–200 (2009)
Marionneau, S. et al. ABH and Lewis histo-blood group antigens, a model for the meaning of oligosaccharide diversity in the face of a changing world. Biochimie 83, 565–573 (2001)
Ilver, D. et al. Helicobacter pylori adhesin binding fucosylated histo-blood group antigens revealed by retagging. Science 279, 373–377 (1998)
Glass, R. I., Parashar, U. D. & Estes, M. K. Norovirus gastroenteritis. N. Engl. J. Med. 361, 1776–1785 (2009)
Matthijnssens, J. et al. Uniformity of rotavirus strain nomenclature proposed by the Rotavirus Classification Working Group (RCWG). Arch. Virol. 156, 1397–1413 (2011)
Monnier, N. et al. High-resolution molecular and antigen structure of the VP8* core of a sialic acid-independent human rotavirus strain. J. Virol. 80, 1513–1523 (2006)
Gerna, G. et al. Identification of a new VP4 serotype of human rotaviruses. Virology 200, 66–71 (1994)
Ciarlet, M. & Estes, M. K. Human and most animal rotavirus strains do not require the presence of sialic acid on the cell surface for efficient infectivity. J. Gen. Virol. 80, 943–948 (1999)
Chitambar, S. D., Arora, R., Kolpe, A. B., Yadav, M. M. & Raut, C. G. Molecular characterization of unusual bovine group A rotavirus G8P[14] strains identified in western India: emergence of P[14] genotype. Vet. Microbiol. 148, 384–388 (2011)
Fukai, K., Saito, T., Inoue, K. & Sato, M. Molecular characterization of novel P[14],G8 bovine group A rotavirus, Sun9, isolated in Japan. Virus Res. 105, 101–106 (2004)
Matthijnssens, J. et al. Are human P[14] rotavirus strains the result of interspecies transmissions from sheep or other ungulates that belong to the mammalian order Artiodactyla? J. Virol. 83, 2917–2929 (2009)
Byres, E. et al. Incorporation of a non-human glycan mediates human susceptibility to a bacterial toxin. Nature 456, 648–652 (2008)
Stevens, J. et al. Glycan microarray analysis of the hemagglutinins from modern and pandemic influenza viruses reveals different receptor specificities. J. Mol. Biol. 355, 1143–1155 (2006)
Neu, U. et al. Structure-function analysis of the human JC polyomavirus establishes the LSTc pentasaccharide as a functional receptor motif. Cell Host Microbe 8, 309–319 (2010)
Midgley, S. E., Hjulsager, C. K., Larsen, L. E., Falkenhorst, G. & Bottiger, B. Suspected zoonotic transmission of rotavirus group A in Danish adults. Epidemiol. Infect. 10.1017/S0950268811001981 (27 September 2011)
Parashar, U. D., Gibson, C. J., Bresse, J. S. & Glass, R. I. Rotavirus and severe childhood diarrhea. Emerg. Infect. Dis. 12, 304–306 (2006)
Estes, M. K. & Kapikian, A. Z. in Fields Virology Vol. 2 (eds Knipe, D. M. & Howley, P. M. ) 1917–1974 (Lippincott Williams & Wilkins, 2007)
Gray, J. & Iturriza-Gomara, M. Rotaviruses. Methods Mol. Biol. 665, 325–355 (2011)
Angel, J., Franco, M. A. & Greenberg, H. B. Rotavirus vaccines: recent developments and future considerations. Nature Rev. Microbiol. 5, 529–539 (2007)
Gentsch, J. R. et al. Serotype diversity and reassortment between human and animal rotavirus strains: implications for rotavirus vaccine programs. J. Infect. Dis. 192 (Suppl 1). S146–S159 (2005)
Guillon, P. et al. Inhibition of the interaction between the SARS-CoV spike protein and its cellular receptor by anti-histo-blood group antibodies. Glycobiology 18, 1085–1093 (2008)
Pflugrath, J. W. The finer things in X-ray diffraction data collection. Acta Crystallogr. D 55, 1718–1725 (1999)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Morris, R. J., Perrakis, A. & Lamzin, V. S. ARP/wARP and automatic interpretation of protein electron density maps. Methods Enzymol. 374, 229–244 (2003)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Bohne, A., Lang, E. & von der Lieth, C. W. SWEET - WWW-based rapid 3D construction of oligo- and polysaccharides. Bioinformatics 15, 767–768 (1999)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Lütteke, T., Frank, M. & von der Lieth, C. W. Carbohydrate Structure Suite (CSS): analysis of carbohydrate 3D structures derived from the PDB. Nucleic Acids Res. 33, D242–D246 (2005)
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. 8, 127–134 (1995)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Smith, D. F., Song, X. & Cummings, R. D. Use of glycan microarrays to explore specificity of glycan-binding proteins. Methods Enzymol. 480, 417–444 (2010)
Haselhorst, T. et al. Sialic acid dependence in rotavirus host cell invasion. Nature Chem. Biol. 5, 91–93 (2009)
Crawford, S. E. et al. Rotavirus viremia and extraintestinal viral infection in the neonatal rat model. J. Virol. 80, 4820–4832 (2006)
Guillon, P. et al. Inhibition of the interaction between the SARS-CoV spike protein and its cellular receptor by anti-histo-blood group antibodies. Glycobiology 18, 1085–1093 (2008)
Hutson, A. M., Atmar, R. L., Marcus, D. M. & Estes, M. K. Norwalk virus-like particle hemagglutination by binding to H histo-blood group antigens. J. Virol. 77, 405–415 (2003)
Acknowledgements
We acknowledge the support from NIH grants AI36040 (to B.V.V.P.), AI 080656 and P30 DK56338 (to M.K.E.), GM62116 (to the Consortium for Functional Glycomics), and the Robert Welch foundation (Q1279) to B.V.V.P. We thank R. Atmar and S. Shanker for helpful discussions and BCM X-ray core facility for data collection.
Author information
Authors and Affiliations
Contributions
L.H. carried out expression, purification, crystallization, diffraction data collection and structure determination. L.H., S.E.C., R.C. and N.W.C.-P. contributed to virus infectivity assays in HT29, CHO cells and haemagglutination assays and data analyses. D.F.S. contributed to glycan array experiments and analysis. J.L.P. provided parental and genetically modified CHO cells and advice. M.K.E. provided supervision and advice on cell infectivity assays and analysis. L.H. and B.V.V.P. analysed and interpreted the structural data. B.V.V.P. contributed to the overall direction of the project and wrote the manuscript with input from other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-2 and Supplementary Figures 1-6. (PDF 1833 kb)
Rights and permissions
About this article
Cite this article
Hu, L., Crawford, S., Czako, R. et al. Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen. Nature 485, 256–259 (2012). https://doi.org/10.1038/nature10996
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10996
This article is cited by
-
Effective SARS-CoV-2 replication of monolayers of intestinal epithelial cells differentiated from human induced pluripotent stem cells
Scientific Reports (2023)
-
Maternal H-antigen secretor status is an early biomarker for potential preterm delivery
Journal of Perinatology (2021)
-
Highly sialylated mucin-type glycopeptide from porcine intestinal mucosa after heparin extraction: O-glycan profiling and immunological activity evaluation
Glycoconjugate Journal (2021)
-
Recent advances in rotavirus reverse genetics and its utilization in basic research and vaccine development
Archives of Virology (2021)
-
Rotavirus A shedding and HBGA host genetic susceptibility in a birth community-cohort, Rio de Janeiro, Brazil, 2014–2018
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.