Abstract
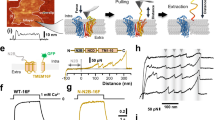
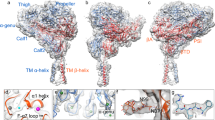
Side chains of Lys/Arg near transmembrane domain (TMD)1,2,3 membrane–water interfaces can ‘snorkel’, placing their positive charge near negatively charged phospholipid head groups4,5,6; however, snorkelling’s functional effects are obscure. Integrin β TMDs have such conserved basic amino acids. Here we use NMR spectroscopy7,8 to show that integrin β3(Lys 716) helps determine β3 TMD topography. The αΙΙbβ3 TMD structure indicates that precise β3 TMD crossing angles enable the assembly of outer and inner membrane ‘clasps’ that hold the αβ TMD together to limit transmembrane signalling9. Mutation of β3(Lys 716) caused dissociation of αΙΙbβ3 TMDs and integrin activation. To confirm that altered topography of β3(Lys 716) mutants activated αΙΙbβ3, we used directed evolution of β3(K716A) to identify substitutions restoring default state. Introduction of Pro(711) at the midpoint of β3 TMD (A711P) increased αΙΙbβ3 TMD association and inactivated integrin αΙΙbβ3(A711P,K716A). β3(Pro 711) introduced a TMD kink of 30 ± 1° precisely at the border of the outer and inner membrane clasps, thereby decoupling the tilt between these segments. Thus, widely occurring snorkelling residues in TMDs can help maintain TMD topography and membrane-embedding, thereby regulating transmembrane signalling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Killian, J. A. & von Heijne, G. How proteins adapt to a membrane–water interface. Trends Biochem. Sci. 25, 429–434 (2000)
von Heijne, G. Membrane proteins: from sequence to structure. Annu. Rev. Biophys. Biomol. Struct. 23, 167–192 (1994)
Sipos, L. & von Heijne, G. Predicting the topology of eukaryotic membrane proteins. Eur. J. Biochem. 213, 1333–1340 (1993)
Krishnakumar, S. S. & London, E. The control of transmembrane helix transverse position in membranes by hydrophilic residues. J. Mol. Biol. 374, 1251–1269 (2007)
Strandberg, E. & Killian, J. A. Snorkeling of lysine side chains in transmembrane helices: how easy can it get? FEBS Lett. 544, 69–73 (2003)
Strandberg, E. et al. Lipid dependence of membrane anchoring properties and snorkeling behavior of aromatic and charged residues in transmembrane peptides. Biochemistry 41, 7190–7198 (2002)
Lau, T. L., Partridge, A. W., Ginsberg, M. H. & Ulmer, T. S. Structure of the integrin beta3 transmembrane segment in phospholipid bicelles and detergent micelles. Biochemistry 47, 4008–4016 (2008)
Lau, T. L., Dua, V. & Ulmer, T. S. Structure of the integrin αIIb transmembrane segment. J. Biol. Chem. 283, 16162–16168 (2008)
Lau, T. L., Kim, C., Ginsberg, M. H. & Ulmer, T. S. The structure of the integrin αIIbβ3 transmembrane complex explains integrin transmembrane signalling. EMBO J. 28, 1351–1361 (2009)
Arnaout, M. A., Mahalingam, B. & Xiong, J. P. Integrin structure, allostery, and bidirectional signaling. Annu. Rev. Cell Dev. Biol. 21, 381–410 (2005)
Shattil, S. J., Kim, C. & Ginsberg, M. H. The final steps of integrin activation: the end game. Nature Rev. Mol. Cell Biol. 11, 288–300 (2010)
Ginsberg, M. H., Partridge, A. & Shattil, S. J. Integrin regulation. Curr. Opin. Cell Biol. 17, 509–516 (2005)
Stefansson, A., Armulik, A., Nilsson, I., von Heijne, G. & Johansson, S. Determination of N- and C-terminal borders of the transmembrane domain of integrin subunits. J. Biol. Chem. 279, 21200–21205 (2004)
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002)
Zhu, J. et al. Requirement of α and β subunit transmembrane helix separation for integrin outside-in signaling. Blood 110, 2475–2483 (2007)
Kim, C., Lau, T. L., Ulmer, T. S. & Ginsberg, M. H. Interactions of platelet integrin αIIb and β3 transmembrane domains in mammalian cell membranes and their role in integrin activation. Blood 113, 4747–4753 (2009)
Shattil, S. J., Hoxie, J. A., Cunningham, M. & Brass, L. F. Changes in the platelet membrane glycoprotein IIb.IIIa complex during platelet activation. J. Biol. Chem. 260, 11107–11114 (1985)
Zhu, J. et al. The structure of a receptor with two associating transmembrane domains on the cell surface: integrin αIIbβ3. Mol. Cell 34, 234–249 (2009)
Frankel, A. D. & Young, J. A. HIV-1: fifteen proteins and an RNA. Annu. Rev. Biochem. 67, 1–25 (1998)
Tadokoro, S. et al. Talin binding to integrin β tails: a final common step in integrin activation. Science 302, 103–106 (2003)
Anthis, N. J. et al. The structure of an integrin/talin complex reveals the basis of inside-out signal transduction. EMBO J. 28, 3623–3632 (2009)
Senes, A., Engel, D. E. & DeGrado, W. F. Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs. Curr. Opin. Struct. Biol. 14, 465–479 (2004)
Berger, B. W. et al. Consensus motif for integrin transmembrane helix association. Proc. Natl Acad. Sci. USA 107, 703–708 (2010)
Ye, F. et al. Recreation of the terminal events in physiological integrin activation. J. Cell Biol. 188, 157–173 (2010)
Han, J. et al. Reconstructing and deconstructing agonist-induced activation of integrin αIIbβ3. Curr. Biol. 16, 1796–1806 (2006)
Acknowledgements
This work was supported by grants from the National Institutes of Health of the USA. T.S.U. acknowledges support from the National Institutes of Health (HL089726) and M.H.G. was supported by HL078784, HL57900 and AR27214. C.K. is a recipient of a postdoctoral fellowship from the American Institute for Cancer Research.
Author information
Authors and Affiliations
Contributions
The project was conceived by C.K. and M.H.G. All experiments with the exception of the NMR studies were performed by C.K. The NMR studies were conducted by T.S. under the supervision of T.S.U. E.C. and F.Y. contributed reagents. M.H.G. and C.K. wrote the paper, which was edited by T.S. and T.S.U.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-10 with legends, and Supplementary Tables 1-3. (PDF 5072 kb)
Rights and permissions
About this article
Cite this article
Kim, C., Schmidt, T., Cho, EG. et al. Basic amino-acid side chains regulate transmembrane integrin signalling. Nature 481, 209–213 (2012). https://doi.org/10.1038/nature10697
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10697
This article is cited by
-
Functional Analysis of Conserved Transmembrane Charged Residues and a Yeast Specific Extracellular Loop of the Plasma Membrane Na+/H+ Antiporter of Schizosaccharomyces pombe
Scientific Reports (2019)
-
The membrane-distal regions of integrin α cytoplasmic domains contribute differently to integrin inside-out activation
Scientific Reports (2018)
-
On the contributing role of the transmembrane domain for subunit-specific sensitivity of integrin activation
Scientific Reports (2018)
-
Differential Binding of Active and Inactive Integrin to Talin
The Protein Journal (2018)
-
Preferential selection of Arginine at the lipid-water-interface of TRPV1 during vertebrate evolution correlates with its snorkeling behaviour and cholesterol interaction
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.