Abstract
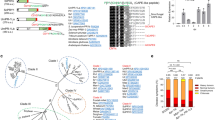
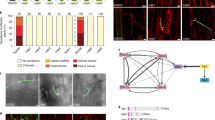
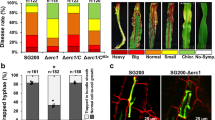
Maize smut caused by the fungus Ustilago maydis is a widespread disease characterized by the development of large plant tumours. U. maydis is a biotrophic pathogen that requires living plant tissue for its development and establishes an intimate interaction zone between fungal hyphae and the plant plasma membrane. U. maydis actively suppresses plant defence responses by secreted protein effectors1,2. Its effector repertoire comprises at least 386 genes mostly encoding proteins of unknown function1,3,4 and expressed exclusively during the biotrophic stage3. The U. maydis secretome also contains about 150 proteins with probable roles in fungal nutrition, fungal cell wall modification and host penetration as well as proteins unlikely to act in the fungal-host interface4 like a chorismate mutase. Chorismate mutases are key enzymes of the shikimate pathway and catalyse the conversion of chorismate to prephenate, the precursor for tyrosine and phenylalanine synthesis. Root-knot nematodes inject a secreted chorismate mutase into plant cells likely to affect development5,6. Here we show that the chorismate mutase Cmu1 secreted by U. maydis is a virulence factor. The enzyme is taken up by plant cells, can spread to neighbouring cells and changes the metabolic status of these cells through metabolic priming. Secreted chorismate mutases are found in many plant-associated microbes and might serve as general tools for host manipulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Doehlemann, G. et al. Pep1, a secreted effector protein of Ustilago maydis, is required for successful invasion of plant cells. PLoS Pathog. 5, e1000290 (2009)
Doehlemann, G. et al. Reprogramming a maize plant: transcriptional and metabolic changes induced by the fungal biotroph Ustilago maydis . Plant J. 56, 181–195 (2008)
Kamper, J. et al. Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis . Nature 444, 97–101 (2006)
Mueller, O. et al. The secretome of the maize pathogen Ustilago maydis . Fungal Genet. Biol. 45 (suppl. 1). S63–S70 (2008)
Bekal, S., Niblack, T. L. & Lambert, K. N. A chorismate mutase from the soybean cyst nematode Heterodera glycines shows polymorphisms that correlate with virulence. Mol. Plant Microbe Interact. 16, 439–446 (2003)
Doyle, E. A. & Lambert, K. N. Meloidogyne javanica chorismate mutase 1 alters plant cell development. Mol. Plant Microbe Interact. 16, 123–131 (2003)
Guermeur, Y., Geourjon, C., Gallinari, P. & Deleage, G. Improved performance in protein secondary structure prediction by inhomogeneous score combination. Bioinformatics 15, 413–421 (1999)
Sasso, S., Ramakrishnan, C., Gamper, M., Hilvert, D. & Kast, P. Characterization of the secreted chorismate mutase from the pathogen Mycobacterium tuberculosis . FEBS J. 272, 375–389 (2005)
Eberhard, J., Raesecke, H.-R., Schmid, J. & Amrhein, N. Cloning and expression in yeast of a higher plant chorismate mutase Molecular cloning, sequencing of the cDNA and characterization of the Arabidopsis thaliana enzyme expressed in yeast. FEBS Lett. 334, 233–236 (1993)
Krappmann, S. et al. The aroC gene of Aspergillus nidulans codes for a monofunctional, allosterically regulated chorismate mutase. J. Biol. Chem. 274, 22275–22282 (1999)
Mobley, E. M., Kunkel, B. N. & Keith, B. Identification, characterization and comparative analysis of a novel chorismate mutase gene in Arabidopsis thaliana . Gene 240, 115–123 (1999)
Basse, C. W., Kolb, S. & Kahmann, R. A maize-specifically expressed gene cluster in Ustilago maydis . Mol. Microbiol. 43, 75–93 (2002)
Doehlemann, G., Reissmann, S., Assmann, D., Fleckenstein, M. & Kahmann, R. Two linked genes encoding a secreted effector and a membrane protein are essential for Ustilago maydis-induced tumour formation. Mol. Microbiol. 81, 751–766 (2011)
Skibbe, D. S., Doehlemann, G., Fernandes, J. & Walbot, V. Maize tumors caused by Ustilago maydis require organ-specific genes in host and pathogen. Science 328, 89–92 (2010)
Bölker, M., Genin, S., Lehmler, C. & Kahmann, R. Genetic regulation of mating and dimorphism in Ustilago maydis . Can. J. Bot. 73, 320–325 (1995)
Di Stasio, M., Brefort, T., Mendoza-Mendoza, A., Munch, K. & Kahmann, R. The dual specificity phosphatase Rok1 negatively regulates mating and pathogenicity in Ustilago maydis . Mol. Microbiol. 73, 73–88 (2009)
Wille, A. C. &. Lucas W. J. Ultrastructural and histochemical studies on guard cells. Planta 160, 129–142 (1984)
Mobley, E. M., Kunkel, B. N. & Keith, B. Identification, characterization and comparative analysis of a novel chorismate mutase gene in Arabidopsis thaliana . Gene 240, 115–123 (1999)
Eberhard, J. et al. Cytosolic and plastidic chorismate mutase isozymes from Arabidopsis thaliana: molecular characterization and enzymatic properties. Plant J. 10, 815–821 (1996)
Schnappauf, G., Lipscomb, W. N. & Braus, G. H. Separation of inhibition and activation of the allosteric yeast chorismate mutase. Proc. Natl Acad. Sci. USA 95, 2868–2873 (1998)
Schnappauf, G., Strater, N., Lipscomb, W. N. & Braus, G. H. A glutamate residue in the catalytic center of the yeast chorismate mutase restricts enzyme activity to acidic conditions. Proc. Natl Acad. Sci. USA 94, 8491–8496 (1997)
Horst, R. J. et al. Ustilago maydis infection strongly alters organic nitrogen allocation in maize and stimulates productivity of systemic source leaves. Plant Physiol. 152, 293–308 (2010)
Glazebrook, J. Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu. Rev. Phytopathol. 43, 205–227 (2005)
Sambrook, J. F. E. & Maniatis, T. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, 1200 (1989)
Loubradou, G., Brachmann, A., Feldbrugge, M. & Kahmann, R. A homologue of the transcriptional repressor Ssn6p antagonizes cAMP signalling in Ustilago maydis . Mol. Microbiol. 40, 719–730 (2001)
Bohlmann, R. Isolierung und Charakterisierung von filamentspezifisch exprimierten Genen aus Ustilago maydis. PhD thesis, Ludwig-Maximilian-Univ. München. (1996)
Gilchrist, D. G. C. & Conelly, J. A. Chorismate mutase from mung bean and sorghum. Methods Enzymol. 142, 450–463 (1987)
Brachmann, A., Weinzierl, G., Kamper, J. & Kahmann, R. Identification of genes in the bW/bE regulatory cascade in Ustilago maydis . Mol. Microbiol. 42, 1047–1063 (2001)
Ueki, S., Lacroix, B., Krichevsky, A., Lazarowitz, S. G. & Citovsky, V. Functional transient genetic transformation of Arabidopsis leaves by biolistic bombardment. Nature Protocols 4, 71–77 (2008)
Doehlemann, G. et al. Establishment of compatibility in the Ustilago maydis/maize pathosystem. J. Plant Physiol. 165, 29–40 (2008)
Shimada, Y., Ichinose, S., Sadr, A., Burrow, M. F. & Tagami, J. Localization of matrix metalloproteinases (MMPs-2, 8, 9 and 20) in normal and carious dentine. Aust. Dent. J. 54, 347–354 (2009)
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 49, 1137–1146 (2008)
Kaever, A. et al. MarVis: a tool for clustering and visualization of metabolic biomarkers. BMC Bioinformatics 10, 92 (2009)
Pommerrenig, B. et al. Phloem-specific expression of yang cycle genes and identification of novel yang cycle enzymes in plantago and Arabidopsis . Plant Cell 23, 1904–1919 (2011)
Naranjo, M. A. et al. Lithium treatment induces a hypersensitive-like response in tobacco. Planta 217, 417–424 (2003)
Banuett, F. & Herskowitz, I. Different a alleles of Ustilago maydis are necessary for maintenance of filamentous growth but not for meiosis. Proc. Natl Acad. Sci. USA 86, 5878–5882 (1989)
De Wit, P. J. G. M. & Spikman, G. Evidence for the occurence of race and cultivar-specific elicitors of necrosis in intercellular fluids of compatible interactions of Cladosporium fulvum and tomato. Physiol. Plant Pathol. 21, 1–11 (1982)
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nature Protoc. 1, 2856–2860 (2006)
Olsen, J. V. & Mann, M. Improved peptide identification in proteomics by two consecutive stages of mass spectrometric fragmentation. Proc Natl Acad Sci USA 101, 13417–13422 (2004)
Mortensen, P. et al. MSQuant, an open source platform for mass spectrometry-based quantitative proteomics. J. Proteome Res. 9, 393–403 (2010)
Acknowledgements
We are thankful to N. Amrhein and H.-U. Mösch for their comments on the manuscript. We thank B. Valent and C. H. Khang for alerting us to the fact that Magnaporthe grisea possesses a secreted chorismate mutase, and are grateful to M. Dickman for allowing us to cite his unpublished results. We thank T. Brefort, E. Mörschel, A. Kaever and M. Landesfeind for experimental support. We acknowledge advice by P. Kast, thank P. Schulze-Lefert for the Gateway-compatible plant transformation vectors, and D. Sicker for providing DIBOA and DIMBOA standards. We acknowledge technical assistance by R. Wissel, S. Löser, D. Vogel, F. Raths, G. Sowa, K. Bolte, M. Johannsen and P. Meyer. Our work was supported through DFG project DJ64/1-1, the collaborative research Center SFB593, and the LOEWE program of the State of Hesse.
Author information
Authors and Affiliations
Contributions
A.D., K.S., F.R., V.V., J.K., S.O., T.T., K.F., P.M., Y.-D.S., H.S., A.G. and B.M. designed and performed the wet bench experiments. All authors contributed to data analysis. R.K., A.D. and K.S. wrote the manuscript with input from all co-authors. R.K. directed the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1- 24 with legends, Supplementary References and Supplementary Tables 1 and 3-8 (see separate file for Supplementary Table 2). (PDF 3189 kb)
Supplementary Table 2
This table shows the data matrix of 810 high quality marker candidates (ANOVA pVal<1x10-5) identified by metabolite fingerprinting by UPLC-ESI TOF-MS analysis in leaves of Zea maize 8 days post U. maydis infection. (XLS 632 kb)
Rights and permissions
About this article
Cite this article
Djamei, A., Schipper, K., Rabe, F. et al. Metabolic priming by a secreted fungal effector. Nature 478, 395–398 (2011). https://doi.org/10.1038/nature10454
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10454
This article is cited by
-
Three new discovery effector proteins from Candidatus Liberibacter asiaticus psy62 inhibit plant defense through interaction with AtCAT3 and AtGAPA
Plant Cell Reports (2024)
-
Long noncoding RNAs emerge from transposon-derived antisense sequences and may contribute to infection stage-specific transposon regulation in a fungal phytopathogen
Mobile DNA (2023)
-
Mechanisms controlling plant proteases and their substrates
Cell Death & Differentiation (2023)
-
A transcriptional activator effector of Ustilago maydis regulates hyperplasia in maize during pathogen-induced tumor formation
Nature Communications (2023)
-
Magnaporthe oryzae encoded effector protein AvrPi54 interacts in vivo with rice encoded cognate resistance protein Pi54 at the host plasma membrane
Journal of Plant Biochemistry and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.