Abstract
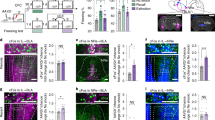
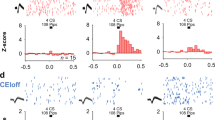
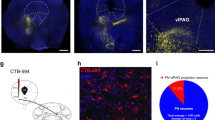
The role of different amygdala nuclei (neuroanatomical subdivisions) in processing Pavlovian conditioned fear has been studied extensively, but the function of the heterogeneous neuronal subtypes within these nuclei remains poorly understood. Here we use molecular genetic approaches to map the functional connectivity of a subpopulation of GABA-containing neurons, located in the lateral subdivision of the central amygdala (CEl), which express protein kinase C-δ (PKC-δ). Channelrhodopsin-2-assisted circuit mapping in amygdala slices and cell-specific viral tracing indicate that PKC-δ+ neurons inhibit output neurons in the medial central amygdala (CEm), and also make reciprocal inhibitory synapses with PKC-δ− neurons in CEl. Electrical silencing of PKC-δ+ neurons in vivo suggests that they correspond to physiologically identified units that are inhibited by the conditioned stimulus, called CEloff units. This correspondence, together with behavioural data, defines an inhibitory microcircuit in CEl that gates CEm output to control the level of conditioned freezing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Davis, M., Walker, D. L. & Myers, K. M. Role of the amygdala in fear extinction measured with potentiated startle. Ann. NY Acad. Sci. 985, 218–232 (2003)
LeDoux, J. E. Emotion circuits in the brain. Annu. Rev. Neurosci. 23, 155–184 (2000)
Paré, D., Quirk, G. J. & LeDoux, J. E. New vistas on amygdala networks in conditioned fear. J. Neurophysiol. 92, 1–9 (2004)
Pitkänen, A., Savander, V. & LeDoux, J. E. Organization of intra-amygdaloid circuitries in the rat: an emerging framework for understanding functions of the amygdala. Trends Neurosci. 20, 517–523 (1997)
Maren, S. & Quirk, G. J. Neuronal signalling of fear memory. Nature Rev. Neurosci. 5, 844–852 (2004)
Medina, J. F., Repa, J. C., Mauk, M. D. & LeDoux, J. E. Parallels between cerebellum- and amygdala-dependent conditioning. Nature Rev. Neurosci. 3, 122–131 (2002)
Cassell, M. D., Freedman, L. J. & Shi, C. The intrinsic organization of the central extended amygdala. Ann. NY Acad. Sci. 877, 217–241 (1999)
Cassell, M. D., Gray, T. S. & Kiss, J. Z. Neuronal architecture in the rat central nucleus of the amygdala: a cytological, hodological, and immunocytochemical study. J. Comp. Neurol. 246, 478–499 (1986)
Day, H. E. W., Curran, E. J., Watson, S. J. & Akil, H. Distinct neurochemical populations in the rat central nucleus of the amygdala and bed nucleus of the stria terminalis: evidence for their selective activation by interleukin-1β. J. Comp. Neurol. 413, 113–128 (1999)
Marchant, N. J., Densmore, V. S. & Osborne, P. B. Coexpression of prodynorphin and corticotrophin-releasing hormone in the rat central amygdala: evidence of two distinct endogenous opioid systems in the lateral division. J. Comp. Neurol. 504, 702–715 (2007)
Ehrlich, I. et al. Amygdala inhibitory circuits and the control of fear memory. Neuron 62, 757–771 (2009)
Huber, D., Veinante, P. & Stoop, R. Vasopressin and oxytocin excite distinct neuronal populations in the central amygdala. Science 308, 245–248 (2005)
Wilensky, A. E., Schafe, G. E., Kristensen, M. P. & LeDoux, J. E. Rethinking the fear circuit: the central nucleus of the amygdala is required for the acquisition, consolidation, and expression of Pavlovian fear conditioning. J. Neurosci. 26, 12387–12396 (2006)
Lerchner, W. et al. Reversible silencing of neuronal excitability in behaving mice by a genetically targeted, ivermectin-gated Cl− channel. Neuron 54, 35–49 (2007)
Slimko, E. M., McKinney, S., Anderson, D. J., Davidson, N. & Lester, H. A. Selective electrical silencing of mammalian neurons in vitro by the use of invertebrate ligand-gated chloride channels. J. Neurosci. 22, 7373–7379 (2002)
Ciocchi, S. et al. Encoding of conditioned fear in central amygdala inhibitory circuits. Nature doi:10.1038/nature09559 (this issue).
Day, H. E., Nebel, S., Sasse, S. & Campeau, S. Inhibition of the central extended amygdala by loud noise and restraint stress. Eur. J. Neurosci. 21, 441–454 (2005)
Zirlinger, M. & Anderson, D. Molecular dissection of the amygdala and its relevance to autism. Genes Brain Behav. 2, 282–294 (2003)
Zirlinger, M., Kreiman, G. & Anderson, D. J. Amygdala-enriched genes identified by microarray technology are restricted to specific amygdaloid subnuclei. Proc. Natl Acad. Sci. USA 98, 5270–5275 (2001)
Callaway, E. M. A molecular and genetic arsenal for systems neuroscience. Trends Neurosci. 28, 196–201 (2005)
Luo, L., Callaway, E. M. & Svoboda, K. Genetic dissection of neural circuits. Neuron 57, 634–660 (2008)
Zhang, F., Aravanis, A. M., Adamantidis, A., de Lecea, L. & Deisseroth, K. Circuit-breakers: optical technologies for probing neural signals and systems. Nature Rev. Neurosci. 8, 577–581 (2007)
Chieng, B. C., Christie, M. J. & Osborne, P. B. Characterization of neurons in the rat central nucleus of the amygdala: cellular physiology, morphology, and opioid sensitivity. J. Comp. Neurol. 497, 910–927 (2006)
Schiess, M. C., Callahan, P. M. & Zheng, H. Characterization of the electrophysiological and morphological properties of rat central amygdala neurons in vitro . J. Neurosci. Res. 58, 663–673 (1999)
Lopez de Armentia, M. & Sah, P. Firing properties and connectivity of neurons in the rat lateral central nucleus of the amygdala. J. Neurophysiol. 92, 1285–1294 (2004)
Gong, S. et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425, 917–925 (2003)
Li, P., Slimko, E. M. & Lester, H. A. Selective elimination of glutamate activation and introduction of fluorescent proteins into a Caenorhabditis elegans chloride channel. FEBS Lett. 528, 77–82 (2002)
Wagstaff, M. J. et al. Gene transfer using a disabled herpes virus vector containing the EMCV IRES allows multiple gene expression in vitro and in vivo . Gene Ther. 5, 1566–1570 (1998)
Veinante, P. & Freund-Mercier, M. J. Branching patterns of central amygdaloid nucleus afferents in the rat: single axon reconstructions. Ann. NY Acad. Sci. 985, 552–553 (2003)
Sun, N., Yi, H. & Cassell, M. D. Evidence for a GABAergic interface between cortical afferents and brainstem projection neurons in the rat central extended amygdala. J. Comp. Neurol. 340, 43–64 (1994)
Gautron, L., Lazarus, M., Scott, M. M., Saper, C. B. & Elmquist, J. K. Identifying the efferent projections of leptin-responsive neurons in the dorsomedial hypothalamus using a novel conditional tracing approach. J. Comp. Neurol. 518, 2090–2108 (2010)
De Oca, B. M., De Cola, J. P., Maren, S. & Fanselow, M. S. Distinct regions of the periaqueductal gray are involved in the acquisition and expression of defensive responses. J. Neurosci. 18, 3426–3432 (1998)
Kim, J. J., Rison, R. A. & Fanselow, M. S. Effects of amygdala, hippocampus, and periaqueductal grady lesions on short- and long-term contextual fear. Behav. Neurosci. 107, 1093–1098 (1993)
LeDoux, J. E., Iwata, J., Cicchetti, P. & Reis, D. J. Different projections of the central amygdaloid nucleus mediate autonomic and behavioral correlates of conditioned fear. J. Neurosci. 8, 2517–2529 (1988)
Kravitz, A. V. et al. Regulation of parkinsonian motor behaviours by optogenetic control of basal ganglia circuitry. Nature 466, 622–626 (2010)
Zhang, F. et al. Multimodal fast optical interrogation of neural circuitry. Nature 446, 633–639 (2007)
Cardin, J. A. et al. Targeted optogenetic stimulation and recording of neurons in vivo using cell-type-specific expression of Channelrhodopsin-2. Nature Protocols 5, 247–254 (2010)
Petreanu, L., Huber, D., Sobczyk, A. & Svoboda, K. Channelrhodopsin-2-assisted circuit mapping of long-range callosal projections. Nature Neurosci. 10, 663–668 (2007)
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007)
Wickersham, I. R., Finke, S., Conzelmann, K. K. & Callaway, E. M. Retrograde neuronal tracing with a deletion-mutant rabies virus. Nature Methods 4, 47–49 (2007)
Slimko, E. M. & Lester, H. A. Codon optimization of Caenorhabditis elegans GluCl ion channel genes for mammalian cells dramatically improves expression levels. J. Neurosci. Methods 124, 75–81 (2003)
Edwards, A. L. Experimental Design in Psychological Research 4th edn, 249–251 (Holt, Reinhard & Winston, 1972)
Wickens, J. R., Arbuthnott, G. W. & Shindou, T. Simulation of GABA function in the basal ganglia: computational models of GABAergic mechanisms in basal ganglia function. Prog. Brain Res. 160, 313–329 (2007)
Gozzi, A. et al. A neural switch for active and passive fear. Neuron 67, 656–666 (2010)
Tsetsenis, T., Ma, X. H., Lo Iacono, L., Beck, S. G. & Gross, C. Suppression of conditioning to ambiguous cues by pharmacogenetic inhibition of the dentate gyrus. Nature Neurosci. 10, 896–902 (2007)
Ressler, K. J. & Mayberg, H. S. Targeting abnormal neural circuits in mood and anxiety disorders: from the laboratory to the clinic. Nature Neurosci. 10, 1116–1124 (2007)
Gong, S., Yang, X. W., Li, C. & Heintz, N. Highly efficient modification of bacterial artificial chromosomes (BACs) using novel shuttle vectors containing the R6Kγ origin of replication. Genome Res. 12, 1992–1998 (2002)
Atasoy, D., Aponte, Y., Su, H. H. & Sternson, S. M. A. FLEX switch targets Channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008)
Seidler, B. et al. A Cre-loxP-based mouse model for conditional somatic gene expression and knockdown in vivo by using avian retroviral vectors. Proc. Natl Acad. Sci. USA 105, 10137–10142 (2008)
Furler, S., Paterna, J. C., Weibel, M. & Bueler, H. Recombinant AAV vectors containing the foot and mouth disease virus 2A sequence confer efficient bicistronic gene expression in cultured cells and rat substantia nigra neurons. Gene Ther. 8, 864–873 (2001)
Szymczak, A. L. et al. Correction of multi-gene deficiency in vivo using a single ‘self-cleaving’ 2A peptide-based retroviral vector. Nature Biotechnol. 22, 589–594 (2004)
Etessami, R. et al. Spread and pathogenic characteristics of a G-deficient rabies virus recombinant: an in vitro and in vivo study. J. Gen. Virol. 81, 2147–2153 (2000)
Herry, C. et al. Switching on and off fear by distinct neuronal circuits. Nature 454, 600–606 (2008)
Nicolelis, M. A. et al. Chronic, multisite, multielectrode recordings in macaque monkeys. Proc. Natl Acad. Sci. USA 100, 11041–11046 (2003)
Acknowledgements
We thank N. Heinz and X. Gong for generating BAC transgenic mice; C. Saper for providing the Cre-dependent hrGFP AAV construct; C. Xiao for training in slice electrophysiology and preliminary experiments; H. Lester for advice on the GluCl system; L. van Tright for performing in situ hybridizations; W. Lerchner for providing a CAG-driven GluClβ construct; A. Chang for help with behavioural scoring; M. Martinez for tail genotyping; G. Mosconi for laboratory management; and J. Alex, R. Bayon and R. Sauza for animal care. This work was supported by NIH grant 1 R01 MH085082-01A1 and by funds from the Caltech ‘Conscious Mouse’ project. W.H. was supported by a fellowship of the Human Frontier Science Program and P.S.K. by the Jane Coffin Childs Memorial Fund for Medical Research. S.C. and A.L. were supported by the Novartis Research Foundation. D.J.A. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
W.H. initiated the project, generated BAC constructs, designed experiments, performed anatomical, viral injections and behavioural experiments, and wrote the manuscript. P.S.K. contributed to experimental design, performed viral injections, behavioural experiments, data analysis and interpretation. H.C. contributed to experimental design and performed viral injections and slice electrophysiology experiments, data analysis and interpretation. S.C. and A.L. designed, performed and interpreted in vivo recording experiments (Fig. 5). N.R.W. and E.M.C. performed rabies virus injection experiments. R.P. performed supplementary behavioural experiments and M.S.F. contributed to their interpretation and to statistical analysis. J.B. and H.-W.D. performed supplementary stereotaxic viral injection experiments. K.D. provided Cre-dependent ChR2 constructs and advice. D.J.A. conceived the project, contributed to experimental design and interpretation and wrote the manuscript. P.S.K., H.C. and S.C. contributed equally. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-6, Supplementary Figures 1- 11 with legends, Supplementary Notes 1-3 and additional references. (PDF 2911 kb)
Rights and permissions
About this article
Cite this article
Haubensak, W., Kunwar, P., Cai, H. et al. Genetic dissection of an amygdala microcircuit that gates conditioned fear. Nature 468, 270–276 (2010). https://doi.org/10.1038/nature09553
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09553
This article is cited by
-
Social buffering in rats reduces fear by oxytocin triggering sustained changes in central amygdala neuronal activity
Nature Communications (2024)
-
The parabrachial to central amygdala pathway is critical to injury-induced pain sensitization in mice
Neuropsychopharmacology (2024)
-
Suppressing fear in the presence of a safety cue requires infralimbic cortical signaling to central amygdala
Neuropsychopharmacology (2024)
-
Neurotechnologies, Ethics, and the Limits of Free Will
Integrative Psychological and Behavioral Science (2024)
-
Experience-dependent changes in affective valence of taste in male mice
Molecular Brain (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.