Abstract
The 1000 Genomes Project aims to provide a deep characterization of human genome sequence variation as a foundation for investigating the relationship between genotype and phenotype. Here we present results of the pilot phase of the project, designed to develop and compare different strategies for genome-wide sequencing with high-throughput platforms. We undertook three projects: low-coverage whole-genome sequencing of 179 individuals from four populations; high-coverage sequencing of two mother–father–child trios; and exon-targeted sequencing of 697 individuals from seven populations. We describe the location, allele frequency and local haplotype structure of approximately 15 million single nucleotide polymorphisms, 1 million short insertions and deletions, and 20,000 structural variants, most of which were previously undescribed. We show that, because we have catalogued the vast majority of common variation, over 95% of the currently accessible variants found in any individual are present in this data set. On average, each person is found to carry approximately 250 to 300 loss-of-function variants in annotated genes and 50 to 100 variants previously implicated in inherited disorders. We demonstrate how these results can be used to inform association and functional studies. From the two trios, we directly estimate the rate of de novo germline base substitution mutations to be approximately 10−8 per base pair per generation. We explore the data with regard to signatures of natural selection, and identify a marked reduction of genetic variation in the neighbourhood of genes, due to selection at linked sites. These methods and public data will support the next phase of human genetic research.
Similar content being viewed by others
Main
Understanding the relationship between genotype and phenotype is one of the central goals in biology and medicine. The reference human genome sequence1 provides a foundation for the study of human genetics, but systematic investigation of human variation requires full knowledge of DNA sequence variation across the entire spectrum of allele frequencies and types of DNA differences. Substantial progress has already been made. By 2008 the public catalogue of variant sites (dbSNP 129) contained approximately 11 million single nucleotide polymorphisms (SNPs) and 3 million short insertions and deletions (indels)2,3,4. Databases of structural variants (for example, dbVAR) indexed the locations of large genomic variants. The International HapMap Project catalogued both allele frequencies and the correlation patterns between nearby variants, a phenomenon known as linkage disequilibrium (LD), across several populations for 3.5 million SNPs3,4.
These resources have driven disease gene discovery in the first generation of genome-wide association studies (GWAS), wherein genotypes at several hundred thousand variant sites, combined with the knowledge of LD structure, allow the vast majority of common variants (here, those with >5% minor allele frequency (MAF)) to be tested for association4 with disease. Over the past 5 years association studies have identified more than a thousand genomic regions associated with disease susceptibility and other common traits5. Genome-wide collections of both common and rare structural variants have similarly been tested for association with disease6.
Despite these successes, much work is still needed to achieve a deep understanding of the genetic contribution to human phenotypes7. Once a region has been identified as harbouring a risk locus, detailed study of all genetic variants in the locus is required to discover the causal variant(s), to quantify their contribution to disease susceptibility, and to elucidate their roles in functional pathways. Low-frequency and rare variants (here defined as 0.5% to 5% MAF, and below 0.5% MAF, respectively) vastly outnumber common variants and also contribute significantly to the genetic architecture of disease, but it has not yet been possible to study them systematically7,8,9. Meanwhile, advances in DNA sequencing technology have enabled the sequencing of individual genomes10,11,12,13, illuminating the gaps in the first generation of databases that contain mostly common variant sites. A much more complete catalogue of human DNA variation is a prerequisite to understand fully the role of common and low-frequency variants in human phenotypic variation.
The aim of the 1000 Genomes Project is to discover, genotype and provide accurate haplotype information on all forms of human DNA polymorphism in multiple human populations. Specifically, the goal is to characterize over 95% of variants that are in genomic regions accessible to current high-throughput sequencing technologies and that have allele frequency of 1% or higher (the classical definition of polymorphism) in each of five major population groups (populations in or with ancestry from Europe, East Asia, South Asia, West Africa and the Americas). Because functional alleles are often found in coding regions and have reduced allele frequencies, lower frequency alleles (down towards 0.1%) will also be catalogued in such regions.
Here we report the results of the pilot phase of the project, the aim of which was to develop and compare different strategies for genome-wide sequencing with high-throughput platforms. To this end we undertook three projects: low-coverage sequencing of 179 individuals; deep sequencing of six individuals in two trios; and exon sequencing of 8,140 exons in 697 individuals (Box 1). The results give us a much deeper, more uniform picture of human genetic variation than was previously available, providing new insights into the landscapes of functional variation, genetic association and natural selection in humans.
Data generation, alignment and variant discovery
A total of 4.9 terabases of DNA sequence was generated in nine sequencing centres using three sequencing technologies, from DNA obtained from immortalized lymphoblastoid cell lines (Table 1 and Supplementary Table 1). All sequenced individuals provided informed consent and explicitly agreed to public dissemination of their variation data, as part of the HapMap Project (see Supplementary Information for details of informed consent and data release). The heterogeneity of the sequence data (read lengths from 25 to several hundred base pairs (bp); single and paired end) reflects the diversity and rapid evolution of the underlying technologies during the project. All primary sequence data were confirmed to have come from the correct individual by comparison to HapMap SNP genotype data.
Analysis to detect and genotype sequence variants differed among variant types and the three projects, but all workflows shared the following four features. (1) Discovery: alignment of sequence reads to the reference genome and identification of candidate sites or regions at which one or more samples differ from the reference sequence; (2) filtering: use of quality control measures to remove candidate sites that were probably false positives; (3) genotyping: estimation of the alleles present in each individual at variant sites or regions; (4) validation: assaying a subset of newly discovered variants using an independent technology, enabling the estimation of the false discovery rate (FDR). Independent data sources were used to estimate the accuracy of inferred genotypes.
All primary sequence reads, mapped reads, variant calls, inferred genotypes, estimated haplotypes and new independent validation data are publicly available through the project website (http://www.1000genomes.org); filtered sets of variants, allele frequencies and genotypes were also deposited in dbSNP (http://www.ncbi.nlm.nih.gov/snp).
Alignment and the ‘accessible genome’
Sequencing reads were aligned to the NCBI36 reference genome (details in Supplementary Information) and made available in the BAM file format14, an early innovation of the project for storing and sharing high-throughput sequencing data. Accurate identification of genetic variation depends on alignment of the sequence data to the correct genomic location. We restricted most variant calling to the ‘accessible genome’, defined as that portion of the reference sequence that remains after excluding regions with many ambiguously placed reads or unexpectedly high or low numbers of aligned reads (Supplementary Information). This approach balances the need to reduce incorrect alignments and false-positive detection of variants against maximizing the proportion of the genome that can be interrogated.
For the low-coverage analysis, the accessible genome contains approximately 85% of the reference sequence and 93% of the coding sequences. Over 99% of sites genotyped in the second generation haplotype map (HapMap II)4 are included. Of inaccessible sites, over 97% are annotated as high-copy repeats or segmental duplications. However, only one-quarter of previously discovered repeats and segmental duplications were inaccessible (Supplementary Table 2). Much of the data for the trio project were collected before technical improvements in our ability to map sequence reads robustly to some of the repeated regions of the genome (primarily longer, paired reads). For these reasons, stringent alignment was more difficult and a smaller portion of the genome was accessible in the trio project: 80% of the reference, 85% of coding sequence and 97% of HapMap II sites (Table 1).
Calibration, local realignment and assembly
The quality of variant calls is influenced by many factors including the quantification of base-calling error rates in sequence reads, the accuracy of local read alignment and the method by which individual genotypes are defined. The project introduced key innovations in each of these areas (see Supplementary Information). First, base quality scores reported by the image processing software were empirically recalibrated by tallying the proportion that mismatched the reference sequence (at non-dbSNP sites) as a function of the reported quality score, position in read and other characteristics. Second, at potential variant sites, local realignment of all reads was performed jointly across all samples, allowing for alternative alleles that contained indels. This realignment step substantially reduced errors, because local misalignment, particularly around indels, can be a major source of error in variant calling. Finally, by initially analysing the data with multiple genotype and variant calling algorithms and then generating a consensus of these results, the project reduced genotyping error rates by 30–50% compared to those currently achievable using any one of the methods alone (Supplementary Fig. 1 and Supplementary Table 12).
We also used local realignment to generate candidate alternative haplotypes in the process of calling short (1–50-bp) indels15, as well as local de novo assembly to resolve breakpoints for deletions greater than 50 bp. The latter resulted in a doubling of the number of large (>1 kb) structural variants delineated with base-pair resolution16. Full genome de novo assembly was also performed (Supplementary Information), resulting in the identification of 3.7 megabases (Mb) of novel sequence not matching the reference at a high threshold for assembly quality and novelty. All novel sequence matched other human and great ape sequences in the public databases.
Rates of variant discovery
In the trio project, with an average mapped sequence coverage of 42× per individual across six individuals and 2.3 gigabases (Gb) of accessible genome, we identified 5.9 million SNPs, 650,000 short indels (of 1–50 bp in length), and over 14,000 larger structural variants. In the low-coverage project, with average mapped coverage of 3.6× per individual across 179 individuals (Supplementary Fig. 2) and 2.4 Gb of accessible genome, we identified 14.4 million SNPs, 1.3 million short indels and over 20,000 larger structural variants. In the exon project, with an average mapped sequence coverage of 56× per individual across 697 individuals and a target of 1.4 Mb, we identified 12,758 SNPs and 96 indels.
Experimental validation was used to estimate and control the FDR for novel variants (Supplementary Table 3). The FDR for each complete call set was controlled to be less than 5% for SNPs and short indels, and less than 10% for structural variants. Because in an initial test almost all of the sites that we called that were already in dbSNP were validated (285 out of 286), in most subsequent validation experiments we tested only novel variants and extrapolated to obtain the overall FDR. This process will underestimate the true FDR if more SNPs listed in dbSNP are false positives for some call sets. The FDR for novel variants was 2.6% for trio SNPs, 10.9% for low-coverage SNPs, and 1.7% for low-coverage indels (Supplementary Information and Supplementary Tables 3 and 4a, b).
Variation detected by the project is not evenly distributed across the genome: certain regions, such as the human leukocyte antigen (HLA) and subtelomeric regions, show high rates of variation, whereas others, for example a 5-Mb gene-dense and highly conserved region around 3p21, show very low levels of variation (Supplementary Fig. 3a). At the chromosomal scale we see strong correlation between different forms of variation, particularly between SNPs and indels (Supplementary Fig. 3b). However, we also find heterogeneity particular to types of structural variant, for example structural variants resulting from non-allelic homologous recombination are apparently enriched in the HLA and subtelomeric regions (Supplementary Fig. 3b, top).
Variant novelty
As expected, the vast majority of sites variant in any given individual were already present in dbSNP; the proportion newly discovered differed substantially among populations, variant types and allele frequencies (Fig. 1). Novel SNPs had a strong tendency to be found only in one analysis panel (set of related populations; Fig. 1a). For SNPs also present in dbSNP version 129 (the last release before 1000 Genomes Project data), only 25% were specific to a single low-coverage analysis panel and 56% were found in all panels. On the other hand, 84% of newly discovered SNPs were specific to a single analysis panel whereas only 4% were found in all analysis panels. In the exon project, where increased depth of coverage and sample size resulted in a higher fraction of low-frequency variants among discovered sites, 96% of novel variants were restricted to samples from a single analysis panel. In contrast, many novel structural variants were identified in all analysis panels, reflecting the lower degree of previous characterization (Supplementary Fig. 4).
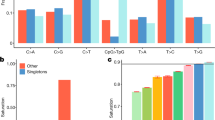
a, Venn diagrams showing the numbers of SNPs identified in each pilot project in each population or analysis panel, subdivided according to whether the SNP was present in dbSNP release 129 (Known) or not (Novel). Exon analysis panel AFR is YRI+LWK, ASN is CHB+CHD+JPT, and EUR is CEU+TSI. Note that the scale for the exon project column is much larger than for the other pilots. b, The number of variants per megabase (Mb) at different allele frequencies divided by the expectation under the neutral coalescent (1/i, where i is the variant allele count), thus giving an estimate of theta per megabase. Blue, low-coverage SNPs; red, low-coverage indels; black, low-coverage genotyped large deletions; green, exon SNPs. The spikes at the right ends of the lines correspond to excess variants for which all samples differed from the reference (approximately 1 per 30 kb), consistent with errors in the reference sequence. c, Fraction of variants in each allele frequency class that were novel. Novelty was determined by comparison to dbSNP release 129 for SNPs and small indels, dbVar (June 2010) for deletions, and two published genomes10,11 for larger indels. LC, low coverage; EX, exon. d, Size distribution and novelty of variants discovered in the low-coverage project. SNPs are shown in blue, deletions with respect to the reference sequence in red, and insertions or duplications with respect to the reference in green. The fraction of variants in each size bin that were novel is shown by the purple line, and is defined relative to dbSNP (SNPs and indels), dbVar (deletions, duplications, mobile element insertions), dbRIP and other studies47 (mobile element insertions), J. C. Venter and J. Watson genomes10,11 (short indels and large deletions), and short indels from split capillary reads48. To account for ambiguous placement of many indels, discovered indels were deemed to match known indels if they were within 25 bp of a known indel of the same size. To account for imprecise knowledge of the location of most deletions and duplications, discovered variants were deemed to match known variants if they had >50% reciprocal overlap.
Populations with African ancestry contributed the largest number of variants and contained the highest fraction of novel variants, reflecting the greater diversity in African populations. For example, 63% of novel SNPs in the low-coverage project and 44% in the exon project were discovered in the African populations, compared to 33% and 22% in the European ancestry populations.
The larger sample sizes in the exon and low-coverage projects allowed us to detect a large number of low-frequency variants (MAF <5%, Fig. 1b). Compared to the distribution expected from population genetic theory (the neutral coalescent with constant population size), we saw an excess of lower frequency variants in the exon project, reflecting purifying selection against weakly deleterious mutations and recent population growth. There are signs of a similar excess in the low-coverage project SNPs, truncated below 5% variant allele frequency by reduction in power of our call set to discover variants in this range, as discussed below.
As expected, nearly all of the high-frequency SNPs discovered here were already present in dbSNP; this was particularly true in coding regions (Fig. 1c). The public databases were much less complete for SNPs at low frequencies, for short indels and for structural variants (Fig. 1d). For example, in contrast to coding SNPs (91% of common coding SNPs described here were already present in dbSNP), approximately 50% of common short indels observed in this project were novel. These results are expected given the sample sizes used in the sequencing efforts that discovered most of the SNPs previously in dbSNP, and the more limited, and lower resolution, efforts to characterize indels and larger structural variation across the genome.
The number of structural variants that we observed declined rapidly with increasing variant length (Fig. 1d), with notable peaks corresponding to Alus and long interspersed nuclear elements (LINEs). The proportion of larger structural variants that was novel depended markedly on allele size, with variants 10 bp to 5 kb in size most likely to be novel (Fig. 1d). This is expected, as large (>5 kb) deletions and duplications were previously discovered using array-based approaches17,18, whereas smaller structural variants (apart from polymorphic Alu insertions) had been less well ascertained before this study.
Mitochondrial and Y chromosome sequences
Deep coverage of the mitochondrial genome allowed us to manually curate sequences for 163 samples (Supplementary Information). Although variants that were fixed within an individual were consistent with the known phylogeny of the mitochondrial genome (Supplementary Fig. 5), we found a considerable amount of variation within individuals (heteroplasmy). For example, length heteroplasmy was detected in 79% of individuals compared with 52% using capillary sequencing19, largely in the control region (Supplementary Fig. 6a). Base-substitution heteroplasmy was observed in 45% of samples, seven times higher than reported in the control region alone19, and was spread throughout the molecule (Supplementary Fig. 6b). The extent to which this heteroplasmy arose in cell culture remains unknown, but appears low (Supplementary Information).
The Y chromosome was sequenced at an average depth of 1.8× in the 77 males in the low-coverage project, and 15.2× depth in the two trio fathers. Using customized analysis methods (Supplementary Information), we identified 2,870 variable sites, 74% novel, with 55 out of 56 passing independent validation. The Y chromosome phylogeny derived from the new variants identified novel, well supported clades within some of the 12 major haplogroups represented among the samples (for example, O2b in China and Japan; Supplementary Fig. 7). A striking pattern indicative of a recent rapid expansion specific to haplogroup R1b was observed, consistent with the postulated Neolithic origin of this haplogroup in Europe20.
Power to detect variants
The ability of sequencing to detect a site that is segregating in the population is dominated by two factors: whether the non-reference allele is present among the individuals chosen for sequencing, and the number of high-quality and well-mapped reads that overlap the variant site in individuals who carry it. Simple models show that for a given total amount of sequencing, the number of variants discovered is maximized by sequencing many samples at low coverage21,22. This is because high coverage of a few genomes, although providing the highest sensitivity and accuracy in genotyping a single individual, involves considerable redundancy and misses variation not represented by those samples. The low-coverage project provides us with an empirical view of the power of low-coverage sequencing to detect variants of different types and frequencies.
Figure 2a shows the rate of discovery of variants in the CEU (see Box 1 for definitions of this and other populations) samples of the low-coverage project as assessed by comparison to external data sources: HapMap and the exon project for SNPs and array CGH data18 for large deletions. We estimate that although the low-coverage project had only ∼25% power to detect singleton SNPs, power to detect SNPs present five times in the 120 sampled chromosomes was ∼90% (depending on the comparator), and power was essentially complete for those present ten or more times. Similar results were seen in the YRI and CHB+JPT analysis panels at high allele counts, but slightly worse performance for variants present five times (∼85% and 75%, respectively, at HapMap II sites; Supplementary Fig. 8). These results indicate that SNP discovery is less affected by the extent of LD (which is lowest in the YRI) than by sequencing coverage (which was lowest in the CHB and JPT panels).
a, Rates of low-coverage variant detection by allele frequency in CEU. Lines show the fraction of variants seen in overlapping samples in independent studies that were also found to be polymorphic in the low-coverage project (in the same overlapping samples), as a function of allele count in the 60 low-coverage samples. Note that we plot power against expected allele count in 60 samples; for example, a variant present in, say, 2 copies in an overlap of 30 samples is expected to be present 4 times in 60 samples. The crosses on the right represent the average discovery fraction for all variants having more than 10 copies in the sample. Red, HapMap II sites, excluding sites also in HapMap 3 (43 overlapping samples); blue, exon project sites (57 overlapping samples); green, deletions from ref. 18 (60 overlapping samples; deletions were classified as ‘found’ if there was any overlap). Error bars show 95% confidence interval. b, Estimated rates of discovery of variants at different frequencies in the CEU (blue), a population related to the CEU with Fst = 1% (green), and across Europe as a whole (light blue). Inset: cartoon of the statistical model for population history and thus allele frequencies in related populations where an ancestral population gave rise to many equally related populations, one of which (blue circle) has samples sequenced. c, SNP genotype accuracy by allele frequency in the CEU low-coverage project, measured by comparison to HapMap II genotypes at sites present in both call sets, excluding sites that were also in HapMap 3. Lines represent the average accuracy of homozygote reference (red), heterozygote (green) and homozygote alternative calls (blue) as a function of the alternative allele count in the overlapping set of 43 samples, and the overall genotype error rate (grey, at bottom of plot). Inset: number of each genotype class as a function of alternative allele count. d, Coverage and accuracy for the low-coverage and exon projects as a function of depth threshold. For 41 CEU samples sequenced in both the exon and low-coverage projects, on the x axis is shown the number of non-reference SNP genotype calls at HapMap II sites not in HapMap 3 that were called in the exon project target region, and on the y axis is shown the number of these calls that were not variant (that is, are reference homozygote and thus incorrectly were called as variant) according to HapMap II. Each point plotted corresponds to a minimum depth threshold for called sites. Grey lines show constant error rates. The exon project calls (red) were made independently per sample, whereas the low-coverage calls (blue), which were only slightly less accurate, were made using LD information that combined partial information across samples and sites in an imputation-based algorithm. The additional data added from point ‘1’ to point ‘0’ (upper right in the figure) for the low-coverage project were completely imputed.
For deletions larger than 500 bp, power was approximately 40% for singletons and reached 90% for variants present ten times or more in the sample set. Our use of several algorithms for structural variant discovery ensured that all major mechanistic subclasses of deletions were found in our analyses (Supplementary Fig. 9). The lack of appropriate comparator data sets for short indels and larger structural variants other than deletions prevented a detailed assessment of the power to detect these types of variants. However, power to detect short indels was approximately 70% for variants present at least five times in the sample, based on the rediscovery of indels in samples overlapping with the SeattleSNPs project23. Extrapolating from comparisons to Alu insertions discovered in the J. C. Venter genome24 indicated an average sensitivity for common mobile element insertions of about 75%. Analysis of a set of duplications18 indicated that only 30–40% of common duplications were discovered here, mostly as deletions with respect to the reference. Methods capable of discovering inversions and novel sequence insertions in low-coverage data with comparable specificity remain to be developed.
In summary, low-coverage shotgun sequencing provided modest power for singletons in each sample (∼25–40%), and very good power for variants seen five or more times in the samples sequenced. We estimate that there was approximately 95% power to find SNPs with 5% allele frequency in the sequenced samples, and nearly 90% power to find SNPs with 5% allele frequency in populations related by 1% divergence (Fig. 2b). Thus, we believe that the projects found almost all accessible common variation in the sequenced populations and the vast majority of common variants in closely related populations.
Genotype accuracy
Genotypes, and, where possible, haplotypes, were inferred for most variants in each project (see Supplementary Information and Table 1). For the low-coverage data, statistically phased SNP genotypes were derived by using LD structure in addition to sequence information at each site, in part guided by the HapMap 3 phased haplotypes. SNP genotype accuracy varied considerably between projects (trio, low coverage and exon), and as a function of coverage and allele frequency. In the low-coverage project, the overall genotype error rate (based on a consensus of multiple methods) was 1–3% (Fig. 2c and Supplementary Fig. 10). The use of HapMap 3 data greatly assisted phasing of the CEU and YRI samples, for which the HapMap 3 genotypes were phased by transmission, but had a more modest effect on genotype accuracy away from HapMap 3 sites (for further details see Supplementary Information).
The accuracy at heterozygous sites, a more sensitive measure than overall accuracy, was approximately 90% for the lowest frequency variants, increased to over 95% for intermediate frequencies, and dropped to 70–80% for the highest frequency variants (that is, those where the reference allele is the rare allele). We note that these numbers are derived from sites that can be genotyped using array technology, and performance may be lower in harder to access regions of the genome. We find only minor differences in genotype accuracy between populations, reflecting differences in coverage as well as haplotype diversity and extent of LD.
The accuracy of genotypes for large deletions was assessed against previous array-based analyses18 (Supplementary Fig. 11). The genotype error rate across all allele frequencies and genotypes was <1%, with the accuracy of heterozygous genotypes at low (MAF <3%), intermediate (MAF ∼50%) and high-frequency (MAF >97%) variants estimated at 86%, 97% and 83%, respectively. The greater apparent genotype accuracy of structural variants compared to SNPs in the low-coverage project reflects the increased number of informative reads per individual for variants of large size and a bias in the known large deletion genotype set for larger, easier to genotype variants.
For calling genotypes in the low-coverage samples, the utility of using LD information in addition to sequence data at each site was demonstrated by comparison to genotypes of the exon project, which were derived independently for each site using high-coverage data. Figure 2d shows the SNP genotype error rate as a function of depth at the genotyped sites in CEU. A similar number of variants was called, and at comparable accuracy, using minimum 4× depth in the low-coverage project as was obtained with minimum 15× depth in the exon project. To genotype a high fraction of sites both projects needed to make calls at sites with low coverage, and the LD-based calling strategy for the low-coverage project used imputation to make calls at nearly 15% more sites with only a modest increase in error rate.
The accuracy and completeness of the individual genome sequences in the low-coverage project could be estimated from the trio mothers, each of whom was sequenced to high coverage, and for whom data subsampled to 4× were included in the low-coverage analysis. Comparison of the SNP genotypes in the two projects showed that where the CEU mother had at least one variant allele according to the trio analysis, in 96.9% of cases the variant was also identified in the low-coverage project and in 93.8% of cases the genotype was accurately inferred. For the YRI trio mother the equivalent figures are 95.0% and 88.4%, respectively (note that false positives in the trio calls will lead to underestimates of the accuracy).
Putative functional variants
An individual’s genome contains many variants of functional consequence, ranging from the beneficial to the highly deleterious. We estimated that an individual typically differs from the reference human genome sequence at 10,000–11,000 non-synonymous sites (sequence differences that lead to differences in the protein sequence) in addition to 10,000–12,000 synonymous sites (differences in coding exons that do not lead to differences in the protein sequence; Table 2). We found a much smaller number of variants likely to have greater functional impact: 190–210 in-frame indels, 80–100 premature stop codons, 40–50 splice-site-disrupting variants and 220–250 deletions that shift reading frame, in each individual. We estimated that each genome is heterozygous for 50–100 variants classified by the Human Gene Mutation Database (HGMD) as causing inherited disorders (HGMD-DM). Estimates from the different pilot projects were consistent with each other, taking into consideration differences in power to detect low-frequency variants, fraction of the accessible genome and population differences (Table 2), as well as with previous observations based on personal genome sequences10,11. Collectively, we refer to the 340–400 premature stops, splice-site disruptions and frame shifts, affecting 250–300 genes per individual, as putative loss-of-function (LOF) variants.
In total, we found 68,300 non-synonymous SNPs, 34,161 of which were novel (Table 2). In an early analysis, 21,657 non-synonymous SNPs were validated as polymorphic in 620 samples using a custom genotyping array (Supplementary Information). The mean minor allele frequency in the array data was 2.2% for 4,573 novel variants, and 26.2% for previously discovered variants.
Overall we rediscovered 671 (1.3%) of the 50,361 coding single nucleotide variants in HGMD-DM (Supplementary Table 5). The types of disease for which variants were identified were biased towards certain categories (Supplementary Fig. 12), with diseases associated with the eye and reproduction significantly over represented and diseases of the nervous system significantly under represented. These biases reflect multiple factors including differences in the fitness effects of the variants, the extent of medical genetics research and differences in the false reporting rate among ‘disease causing’ variants.
As expected, and consistent with purifying selection, putative functional variants had an allele frequency spectrum depleted at higher allele frequencies, with putative LOF variants showing this effect more strongly (Supplementary Fig. 13). Of the low-coverage non-synonymous, stop-introducing, splice-disrupting and HGMD-DM variants, 67.3%, 77.3%, 82.2% and 84.7% were private to single populations, compared to 61.1% for synonymous variants. Across these same functional classes, 15.8%, 25.9%, 21.6% and 19.9% of variants were found in only a single individual, compared to 11.8% of synonymous variants.
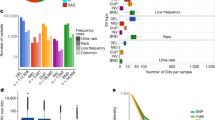
The tendency for deleterious functional variants to have lower allele frequencies has consequences for the discovery and analysis of this type of variation. In the deeply sequenced CEU trio father, who was not included in the low-coverage project, 97.8% of all single base variants had been found in the low-coverage project, but only 95% of non-synonymous, 88% of stop-inducing and 85% of HGMD-DM variants. The missed variants correspond to 389 non-synonymous, 11 stop-inducing and 13 HGMD-DM variants. As sample size increases, the number of novel variants per sequenced individual will decrease, but only slowly. Analyses based on the exon project data (Fig. 3) showed that, on average, 99% of the synonymous variants in an individual would be found in 100 deeply sequenced samples, whereas 250 samples would be required to find 99% of non-synonymous variants and 320 samples would still find only 97.4% of the LOF variants present in an individual. Using detection power data from Fig. 2a, we estimated that 250 samples sequenced at low coverage would be needed to find 99% of the synonymous variants in an individual, and with 320 sequenced samples 98.5% of non-synonymous and 96.3% of LOF variants would be found.
The fraction of variants present in an individual that would not have been found in a sequenced reference panel, as a function of reference panel size and the sequencing strategy. The lines represent predictions for synonymous (Synon.), non-synonymous (Non-synon.) and loss-of-function (LOF) variant classes, broken down by sequencing category: full sequencing as for exons (Full) and low-coverage sequencing (LC). The values were calculated from observed distributions of variants of each class in 321 East Asian samples (CHB, CHD and JPT populations) in the exon data, and power to detect variants at low allele counts in the reference panel from Fig. 2a.
Application to association studies
Whole-genome sequencing enables all genetic variants present in a sample set to be tested directly for association with a given disease or trait. To quantify the benefit of having more complete ascertainment of genetic variation beyond that achievable with genotyping arrays, we carried out expression quantitative trait loci (eQTL) association tests on the 142 low-coverage samples for which expression data are available in the cell lines25. When association analysis (Spearman rank correlation, FDR <5%, eQTLs within 50 kb of probe) was performed using all sites discovered in the low-coverage project, a larger number of significant eQTLs (increase of ∼20% to 50%) was observed as compared to association analysis restricted to sites present on the Illumina 1M chip (Supplementary Table 6). The increase was lower in the CHB+JPT and CEU samples, where greater LD exists between previously examined and newly discovered variants, and higher in the YRI samples, where there are more novel variants and less LD. These results indicate that, while modern genotyping arrays capture most of the common variation, there remain substantial additional contributions to phenotypic variation from the variants not well captured by the arrays.
Population sequencing of large phenotyped cohorts will allow direct association tests for low-frequency variants, with a resolution determined by the LD structure. An alternative that is less expensive, albeit less accurate, is to impute variants from a sequenced reference panel into previously genotyped samples26,27. We evaluated the accuracy of imputation that uses the current low-coverage project haplotypes as the reference panel. Specifically, we compared genotypes derived by deep sequencing of one individual in each trio (the fathers) with genotypes derived using the HapMap 3 genotype data (which combined data from the Affymetrix 6.0 and Illumina 1M arrays) in those same two individuals and imputation based on the low-coverage project haplotypes to fill in their missing genotypes. At variant sites (that is, where the father was not homozygous for the reference sequence), imputation accuracy was highest for SNPs at which the minor allele was observed at least six times in our low-coverage samples, with an error rate of ∼4% in CEU and ∼10% in YRI, and became progressively worse for rarer SNPs, with error rates of 35% for sites where the minor allele was observed only twice in the low-coverage samples (Fig. 4a).
a, Accuracy of imputing variant genotypes using HapMap 3 sites to impute sites from the low-coverage (LC) project into the trio fathers as a function of allele frequency. Accuracy of imputing genotypes from the HapMap II reference panels4 is also shown. Imputation accuracy for common variants was generally a few per cent worse from the low-coverage project than from HapMap, although error rates increase for less common variants. b, An example of imputation in a cis-eQTL for TIMM22, for which the original Ilumina 300K genotype data gave a weak signal28. Imputation using HapMap data made a small improvement, and imputation using low-coverage haplotypes provided a much stronger signal.
Although the ability to impute rare variants accurately from the 1000 Genomes Project resource is currently limited, the completeness of the resource nevertheless increases power to detect association signals. To demonstrate the utility of imputation in disease samples, we imputed into an eQTL study of ∼400 children of European ancestry28 using the low-coverage pilot data and HapMap II as reference panels. By comparison to directly genotyped sites we estimated that the effective sample size at variants imputed from the pilot CEU low-coverage data set is 91% of the true sample size for variants with allele frequencies above 10%, 76% in the allele frequency range 4–6%, and 54% in the range 1–2%. Imputing over 6 million variants from the low-coverage project data increased the number of detected cis-eQTLs by ∼16%, compared to a 9% increase with imputing from HapMap II (FDR 5%, signal within 50 kb of transcript; for an example see Fig. 4b).
In addition to this modest increase in the number of discoveries, testing almost all common variants allows identification of many additional candidate variants that might underlie each association. For example, we find that rs11078928, a variant in a splice site for GSDMB, is in strong LD with SNPs near ORMDL3, previously associated with asthma, Crohn’s disease, type 1 diabetes and rheumatoid arthritis, thus leading to the hypothesis that GSDMB could be the causative gene in these associations. Although rs11078928 is not newly discovered, it was not included in HapMap or on commercial SNP arrays, and thus could not have been identified as associated with these diseases before this project. Similarly, a recent study29 used project data to show that coding variants in APOL1 probably underlie a major risk for kidney disease in African-Americans previously attributed (at a lower effect size) to MYH9. These examples demonstrate the value of having much more complete information on LD, the almost complete set of common variants, and putative functional variants in known association intervals.
Testing almost all common variants also allows us to examine general properties of genetic association signals. The NHGRI GWAS catalogue (http://www.genome.gov/gwastudies, accessed 15 July 2010) described 1,227 unique SNPs associated with one or more traits (P < 5 × 10−8). Of these, 1,185 (96.5%) are present in the low-coverage CEU data set. Under 30% of these are either annotated as non-synonymous variants (77, 6.5%) or in substantial LD (r2 > 0.5) with a non-synonymous variant (272, 23%). In the latter group, only 93 (8.4%) are in strong LD (r2 > 0.9) with a non-synonymous variant. Because we tested ∼95% of common variation, these results indicate that no more than one-third of complex trait association signals are likely to be caused by common coding variation. Although it remains to be seen whether reported associations are better explained through weak LD to coding variants with strong effects, these results are consistent with the view that most contributions of common variation to complex traits are regulatory in nature.
Mutation, recombination and natural selection
Project sequence data allowed us to investigate fundamental processes that shape human genetic variation including mutation, recombination and natural selection.
Detecting de novo mutations in trio samples
Deep sequencing of individuals within a pedigree offers the potential to detect de novo germline mutation events. Our approach was to allow a relatively high FDR in an initial screen to capture a large fraction of true events and then use a second technology to rule out false-positive mutations.
In the CEU and YRI trios, respectively, 3,236 and 2,750 candidate de novo germline single-base mutations were selected for further study, based on their presence in the child but not the parents. Of these, 1,001 (CEU) and 669 (YRI) were validated by re-sequencing the cell line DNA. When these were tested for segregation to offspring (CEU) or in non-clonal DNA from whole blood (YRI), only 49 CEU and 35 YRI candidates were confirmed as true germline mutations. Correcting for the fraction of the genome accessible to this analysis provided an estimate of the per generation base pair mutation rate of 1.2 × 10−8 and 1.0 × 10−8 in the CEU and YRI trios, respectively. These values are similar to estimates obtained from indirect evolutionary comparisons30, direct studies based on pathogenic mutations31, and a recent analysis of a single family32.
We infer that the remaining vast majority (952 CEU and 634 YRI) of the validated variants were somatic or cell line mutations. The greater number of these validated non-germline mutations in the CEU cell line perhaps reflects the greater age of the CEU cell culture. Across the two trio offspring, we observed a single, synonymous, coding germline mutation, and 17 coding non-germline mutations of which 16 were non-synonymous, perhaps indicative of selection during cell culture.
Although the number of non-germline variants found per individual is a very small fraction of the total number of variants per individual (∼0.03% for the CEU child and ∼0.02% for the YRI child), these variants will not be shared between samples. Assuming that the number of non-germline mutations in these two trios is representative of all cell line DNA we analysed, we estimate that non-germline mutations might constitute 0.36% and 2.4% of all variants, and 0.61% and 3.1% of functional variants, in the low-coverage and exon pilots, respectively. In larger samples, of thousands, the overall false-positive rates from cell line mutations would become significant, and confound interpretation, indicating that large-scale studies should use DNA from primary tissue, such as blood, where possible.
The effects of selection on local variation
Natural selection can affect levels of DNA variation around genes in several ways: strongly deleterious mutations will be rapidly eliminated by natural selection, weakly deleterious mutations may segregate in populations but rarely become fixed, and selection at nearby sites (both purifying and adaptive) reduces genetic variation through background selection33 and the hitch-hiking effect34. The effect of these different forces on genetic variation can be disentangled by examining patterns of diversity and divergence within and around known functional elements. The low-coverage data enables, for the first time, genome-wide analysis of such patterns in multiple populations. Figure 5a (top panel) shows the pattern of diversity relative to genic regions measured by aggregating estimates of heterozygosity around protein-coding genes. Within genes, exons harbour the least diversity (about 50% of that of introns) and 5′ and 3′ UTRs harbour slightly less diversity than immediate flanking regions and introns. However, this variation in diversity is fully explained by the level of divergence (Fig. 5a, bottom panel), consistent with the common part of the allele frequency spectrum being dominated by effectively neutral variants, and weakly deleterious variants contributing only to the rare end of the frequency spectrum.
a, Diversity in genes calculated from the CEU low-coverage genotype calls (top) and diversity divided by divergence between humans and rhesus macaque (bottom). Within each element averaged diversity is shown for the first and last 25 bp, with the remaining 150 positions sampled at fixed distances across the element (elements shorter than 150 bp were not considered). Note that estimates of diversity will be reduced compared to the true population value due to the reduced power for rare variants, but relative values should be little affected. b, Average autosomal diversity divided by divergence, as a function of genetic distance from coding transcripts, calculated at putatively neutral sites, that is, excluding phastcons, conserved non-coding sequences and all sites in coding exons but fourfold degenerate sites. c, Numbers of SNPs showing increasingly high levels of differentiation in allele frequency between the CEU and CHB+JPT (red), CEU and YRI (green) and CHB+JPT and YRI (blue). Lines indicate synonymous variants (dashed), non-synonymous variants (dotted) and other variants (solid lines). The most highly differentiated genic SNPs were enriched for non-synonymous variants, indicating local adaptation. d, The decay of population differentiation around genic SNPs showing extreme allele frequency differences between populations (difference in frequency of at least 0.8 between populations, thinned so there is no more than one per gene considered; Supplementary Table 8). For all such SNPs the highest allele frequency difference in bins of 0.01 cM away from the variant was recorded and averaged.
In contrast, diversity in the immediate vicinity of genes (scaled by divergence) is reduced by approximately 10% relative to sites distant from any gene (Fig. 5b). Although a similar reduction has been seen previously in gene-dense regions35, project data enable the scale of the effect to be determined. We find that the reduction extends up to 0.1 cM away from genes, typically 85 kb, indicating that selection at linked sites restricts variation relative to neutral levels across the majority of the human genome.
Population differentiation and positive selection
Previous inferences about demographic history and the role of local adaptation in shaping human genetic variation made from genome-wide genotype data4,36,37 have been limited by the partial and complex ascertainment of SNPs on genotyping arrays. Although data from the 1000 Genomes Project pilots are neither fully comprehensive nor fully free of ascertainment bias (issues include low power for rare variants, noise in allele frequency estimates, some false positives, non-random data collection across samples, platforms and populations, and the use of imputed genotypes), they can be used to address key questions about the extent of differentiation among populations, the presence of highly differentiated variants and the ability to fine-map signals of local adaptation.
Although the average level of population differentiation is low (at sites genotyped in all populations the mean value of Wright’s Fst is 0.071 between CEU and YRI, 0.083 between YRI and CHB+JPT, and 0.052 between CHB+JPT and CEU), we find several hundred thousand SNPs with large allele frequency differences in each population comparison (Fig. 5c). As seen in previous studies4,37, the most highly differentiated sites were enriched for non-synonymous variants, indicative of the action of local adaptation. The completeness of common variant discovery in the low-coverage resource enables new perspectives in the search for local adaptation. First, it provides a more comprehensive catalogue of fixed differences between populations, of which there are very few: two between CEU and CHB+JPT (including the A111T missense variant in SLC24A5 (ref. 38) contributing to light skin colour), four between CEU and YRI (including the −46 GATA box null mutation upstream of DARC39, the Duffy O allele leading to Plasmodium vivax malaria resistance) and 72 between CHB+JPT and YRI (including 24 around the exocyst complex component gene EXOC6B); see Supplementary Table 7 for a complete list. Second, it provides new candidates for selected variants, genes and pathways. For example, we identified 139 non-synonymous variants showing large allele frequency differences (at least 0.8) between populations (Supplementary Table 8), including at least two genes involved in meiotic recombination—FANCA (ninth most extreme non-synonymous SNP in CEU versus CHB+JPT) and TEX15 (thirteenth most extreme non-synonymous SNP in CEU versus YRI, and twenty-sixth most extreme non-synonymous SNP in CHB+JPT versus YRI). Because we are finding almost all common variants in each population, these lists should contain the vast majority of the near fixed differences among these populations. Finally, it improves the fine mapping of selective sweeps (Supplementary Fig. 14) and analysis of the dynamics of location adaptation. For example, we find that the signal of population differentiation around high Fst genic SNPs drops by half within, on average, less than 0.05 cM (typically 30–50 kb; Fig. 5d). Furthermore, 51% of such variants are polymorphic in both populations. These observations indicate that much local adaptation has occurred by selection acting on existing variation rather than new mutation.
The effect of recombination on local sequence evolution
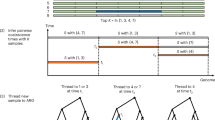
We estimated a fine-scale genetic map from the phased low-coverage genotypes. Recombination hotspots were narrower than previously estimated4 (mean hotspot width of 2.3 kb compared to 5.5 kb in HapMap II; Fig. 6a), although, unexpectedly, the estimated average peak recombination rate in hotspots is lower in YRI (13 cM Mb−1) than in CEU and CHB+JPT (20 cM Mb−1). In addition, crossover activity is less concentrated in the genome in YRI, with 70% of recombination occurring in 10% of the sequence rather than 80% of the recombination for CEU and CHB+JPT (Fig. 6b). A possible biological basis for these differences is that PRDM9, which binds a DNA motif strongly enriched in hotspots and influences the activity of LD-defined hotspots40,41,42,43, shows length variation in its DNA-binding zinc fingers within populations, and substantial differentiation between African and non-African populations, with a greater allelic diversity in Africa43. This could mean greater diversity of hotspot locations within Africa and therefore a less concentrated picture in this data set of recombination and lower usage of LD-defined hotspots (which require evidence in at least two populations and therefore will not reflect hotspots present only in Africa).
a, Improved resolution of hotspot boundaries. The average recombination rate estimated from low-coverage project data around recombination hotspots detected in HapMap II. Recombination hotspots were narrower, and in CEU (orange) and CHB+JPT (purple) more intense than previously estimated. See panel b for key. b, The concentration of recombination in a small fraction of the genome, one line per chromosome. If recombination were uniformly distributed throughout the genome, then the lines on this figure would appear along the diagonal. Instead, most recombination occurs in a small fraction of the genome. Recombination rates in YRI (green) appeared to be less concentrated in recombination hotspots than CEU (orange) or CHB+JPT (purple). HapMap II estimates are shown in black. c, The relationship between genetic variation and recombination rates in the YRI population. The top plot shows average levels of diversity, measured as mean number of segregating sites per base, surrounding occurrences of the previously described hotspot motif40 (CCTCCCTNNCCAC, red line) and a closely related, but not recombinogenic, DNA sequence (CTTCCCTNNCCAC, green line). The lighter red and green shaded areas give 95% confidence intervals on diversity levels. The bottom plot shows estimated mean recombination rates surrounding motif occurrences, with colours defined as in the top plot.
The low-coverage data also allowed us to address a long-standing debate about whether recombination has any local mutagenic effect. Direct examination of diversity around hotspots defined from LD data are potentially biased (because the detection of hotspots requires variation to be present), but we can, without bias, examine rates of SNP variation and recombination around the PRDM9 binding motif associated with hotspots. Figure 6c shows the local recombination rate and pattern of SNP variation around the motif compared to the same plots around a motif that is a single base difference away. Although the motif is associated with a sharp peak in recombination rate, there is no systematic effect on local rates of SNP variation. We infer that, although recombination may influence the fate of new mutations, for example through biased gene conversion, there is no evidence that it influences the rate at which new variants appear.
Discussion
The 1000 Genomes Project launched in 2008 with the goal of creating a public reference database for DNA polymorphism that is 95% complete at allele frequency 1%, and more complete for common variants and exonic variants, in each of multiple human population groups. The three pilot projects described here were designed to develop and evaluate methods to use high-throughput sequencing to achieve these goals. The results indicate (1) that robust protocols now exist for generating both whole-genome shotgun and targeted sequence data; (2) that algorithms to detect variants from each of these designs have been validated; and (3) that low-coverage sequencing offers an efficient approach to detect variation genome wide, whereas targeted sequencing offers an efficient approach to detect and accurately genotype rare variants in regions of functional interest (such as exons).
Data from the pilot projects are already informing medical genetic studies. As shown in our analysis of previous eQTL data sets, a more complete catalogue of genetic variation can identify signals previously missed and markedly increase the number of identified candidate functional alleles at each locus. Project data have been used to impute over 6 million genetic variants into GWAS, for traits as diverse as smoking44 and multiple sclerosis45, as an exclusionary filter in Mendelian disease studies46 and tumour sequencing studies, and to design the next generation of genotyping arrays.
The results from this study also provide a template for future genome-wide sequencing studies on larger sample sets. Our plans for achieving the 1000 Genomes Project goals are described in Box 2. Other studies using phenotyped samples are already using components of the design and analysis framework described above.
Measurement of human DNA variation is an essential prerequisite for carrying out human genetics research. The 1000 Genomes Project represents a step towards a complete description of human DNA polymorphism. The larger data set provided by the full 1000 Genomes Project will allow more accurate imputation of variants in GWAS and thus better localization of disease-associated variants. The project will provide a template for studies using genome-wide sequence data. Applications of these data, and the methods developed to generate them, will contribute to a much more comprehensive understanding of the role of inherited DNA variation in human history, evolution and disease.
Methods Summary
The Supplementary Information provides full details of samples, data generation protocols, read mapping, SNP calling, short insertion and deletion calling, structural variation calling and de novo assembly. Details of methods used in the analyses relating to imputation, mutation rate estimation, functional annotation, population genetics and extrapolation to the full project are also presented.
Change history
25 May 2011
Several corrections to the author consortium list were corrected on 25 May 2011. Please see the corrigendum at the end of the PDF for details. The corresponding author was changed on 28 July 2011
References
The International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004)
Sachidanandam, R. et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001)
The International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005)
The International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
Hindorff, L. A., Junkins, H. A., Hall, P. N., Mehta, J. P. & Manolio, T. A. A catalog of published genome-wide association studies. 〈http://www.genome.gov/gwastudies〉 (2010)
Craddock, N. et al. Genome-wide association study of CNVs in 16,000 cases of eight common diseases and 3,000 shared controls. Nature 464, 713–720 (2010)
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009)
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J. A. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324, 387–389 (2009)
Cohen, J. C., Boerwinkle, E., Mosley, T. H., Jr & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006)
Levy, S. et al. The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007)
Wheeler, D. A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008)
Bentley, D. R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008)
Wang, J. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008)
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Albers, C. et al. Dindel: Accurate indel calls from short read data. Genome Res. (in the press)
Lam, H. Y. et al. Nucleotide-resolution analysis of structural variants using BreakSeq and a breakpoint library. Nature Biotechnol. 28, 47–55 (2010)
The International HapMap 3 Consortium Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010)
Conrad, D. F. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010)
Irwin, J. A. et al. Investigation of heteroplasmy in the human mitochondrial DNA control region: a synthesis of observations from more than 5000 global population samples. J. Mol. Evol. 68, 516–527 (2009)
Balaresque, P. et al. A predominantly neolithic origin for European paternal lineages. PLoS Biol. 8, e1000285 (2010)
Wendl, M. C. & Wilson, R. K. The theory of discovering rare variants via DNA sequencing. BMC Genomics 10, 485 (2009)
Le, S. Q., Li, H. & Durbin, R. QCALL: SNP detection and genotyping from low coverage sequence data on multiple diploid samples. Genome Res. (in the press)
NHLBI Program for Genomic Applications. SeattleSNPs. 〈http://pga.gs.washington.edu/〉 (2010)
Xing, J. et al. Mobile elements create structural variation: analysis of a complete human genome. Genome Res. 19, 1516–1526 (2009)
Stranger, B. E. et al. Population genomics of human gene expression. Nature Genet. 39, 1217–1224 (2007)
Li, Y., Willer, C. J., Ding, J., Scheet, P. & Abecasis, G. R. MaCH: Using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epi. (in the press)
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nature Rev. Genet. 11, 499–511 (2010)
Dixon, A. L. et al. A genome-wide association study of global gene expression. Nature Genet. 39, 1202–1207 (2007)
Genovese, G. et al. Association of trypanolytic ApoL1 variants with kidney disease in African Americans. Science 329, 841–845 (2010)
Nachman, M. W. & Crowell, S. L. Estimate of the mutation rate per nucleotide in humans. Genetics 156, 297–304 (2000)
Kondrashov, A. S. Direct estimates of human per nucleotide mutation rates at 20 loci causing Mendelian diseases. Hum. Mutat. 21, 12–27 (2003)
Roach, J. C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010)
Charlesworth, B., Morgan, M. T. & Charlesworth, D. The effect of deleterious mutations on neutral molecular variation. Genetics 134, 1289–1303 (1993)
Maynard Smith, J. & Haigh, J. The hitch-hiking effect of a favourable gene. Genet. Res. 23, 23–35 (1974)
Cai, J. J., Macpherson, J. M., Sella, G. & Petrov, D. A. Pervasive hitchhiking at coding and regulatory sites in humans. PLoS Genet. 5, e1000336 (2009)
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006)
Barreiro, L. B., Laval, G., Quach, H., Patin, E. & Quintana-Murci, L. Natural selection has driven population differentiation in modern humans. Nature Genet. 40, 340–345 (2008)
Lamason, R. L. et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310, 1782–1786 (2005)
Tournamille, C., Colin, Y., Cartron, J. P. & Le Van Kim, C. Disruption of a GATA motif in the Duffy gene promoter abolishes erythroid gene expression in Duffy-negative individuals. Nature Genet. 10, 224–228 (1995)
Myers, S. et al. Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science 327, 876–879 (2010)
Myers, S., Freeman, C., Auton, A., Donnelly, P. & McVean, G. A common sequence motif associated with recombination hot spots and genome instability in humans. Nature Genet. 40, 1124–1129 (2008)
Baudat, F. et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327, 836–840 (2010)
Parvanov, E. D., Petkov, P. M. & Paigen, K. Prdm9 controls activation of mammalian recombination hotspots. Science 327, 835 (2010)
Liu, J. Z. et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nature Genet. 42, 436–440 (2010)
Sanna, S. et al. Variants within the immunoregulatory CBLB gene are associated with multiple sclerosis. Nature Genet. 42, 495–497 (2010)
Musunuru, K. et al. Exome sequencing, mutations in ANGPTL3, and familial combined hypolipidemia. N. Engl. J. Med. (in the press)
Ewing, A. D. & Kazazian, H. H., Jr High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res. 20, 1262–1270 (2010)
Mills, R. E. et al. An initial map of insertion and deletion (INDEL) variation in the human genome. Genome Res. 16, 1182–1190 (2006)
Liti, G. et al. Population genomics of domestic and wild yeasts. Nature 458, 337–341 (2009)
Li, Y., Willer, C., Sanna, S. & Abecasis, G. Genotype imputation. Annu. Rev. Genomics Hum. Genet. 10, 387–406 (2009)
Acknowledgements
We thank many people who contributed to this project: K. Beal, S. Fitzgerald, G. Cochrane, V. Silventoinen, P. Jokinen, E. Birney and J. Ahringer for comments on the manuscript; T. Hunkapiller and Q. Doan for their advice and coordination; N. Kälin, F. Laplace, J. Wilde, S. Paturej, I. Kühndahl, J. Knight, C. Kodira and M. Boehnke for valuable discussions; Z. Cheng, S. Sajjadian and F. Hormozdiari for assistance in managing data sets; and D. Leja for help with the figures. We thank the Yoruba in Ibadan, Nigeria, the Han Chinese in Beijing, China, the Japanese in Tokyo, Japan, the Utah CEPH community, the Luhya in Webuye, Kenya, the Toscani in Italia, and the Chinese in Denver, Colorado, for contributing samples for research. This research was supported in part by Wellcome Trust grants WT089088/Z/09/Z to R.M.D.; WT085532AIA to P.F.; WT086084/Z/08/Z to G.A.M.; WT081407/Z/06/Z to J.S.K.; WT075491/Z/04 to G.L.; WT077009 to C.T.-S.; Medical Research Council grant G0801823 to J.L.M.; British Heart Foundation grant RG/09/012/28096 to C.A.; The Leverhulme Trust and EPSRC studentships to L.M. and A.T.; the Louis-Jeantet Foundation and Swiss National Science Foundation in support of E.T.D. and S.B.M.; NGI/EBI fellowship 050-72-436 to K.Y.; a National Basic Research Program of China (973 program no. 2011CB809200); the National Natural Science Foundation of China (30725008, 30890032, 30811130531, 30221004); the Chinese 863 program (2006AA02Z177, 2006AA02Z334, 2006AA02A302, 2009AA022707); the Shenzhen Municipal Government of China (grants JC200903190767A, JC200903190772A, ZYC200903240076A, CXB200903110066A, ZYC200903240077A, ZYC200903240076A and ZYC200903240080A); the Ole Rømer grant from the Danish Natural Science Research Council; an Emmy Noether Fellowship of the German Research Foundation (Deutsche Forschungsgemeinschaft) to J.O.K.; BMBF grant 01GS08201; BMBF grant PREDICT 0315428A to R.H.; BMBF NGFN PLUS and EU 6th framework READNA to S.S.; EU 7th framework 242257 to A.V.S.; the Max Planck Society; a grant from Genome Quebec and the Ministry of Economic Development, Innovation and Trade, PSR-SIIRI-195 to P.A.; the Intramural Research Program of the NIH; the National Library of Medicine; the National Institute of Environmental Health Sciences; and NIH grants P41HG4221 and U01HG5209 to C.L.; P41HG4222 to J.S.; R01GM59290 to L.B.J. and M.A.B.; R01GM72861 to M.P.; R01HG2651 and R01MH84698 to G.R.A.; U01HG5214 to G.R.A. and A.C.; P01HG4120 to E.E.E.; U54HG2750 to D.L.A.; U54HG2757 to A.C.; U01HG5210 to D.C.; U01HG5208 to M.J.D.; U01HG5211 to R.A.G.; R01HG3698, R01HG4719 and RC2HG5552 to G.T.M.; R01HG3229 to C.D.B. and A.G.C.; P50HG2357 to M.S.; R01HG4960 to B.L.B; P41HG2371 and U41HG4568 to D.H.; R01HG4333 to A.M.L.; U54HG3273 to R.A.G.; U54HG3067 to E.S.L.; U54HG3079 to R.K.W.; N01HG62088 to the Coriell Institute; S10RR025056 to the Translational Genomics Research Institute; Al Williams Professorship funds for M.B.G.; the BWF and Packard Foundation support for P.C.S.; the Pew Charitable Trusts support for G.R.A.; and an NSF Minority Postdoctoral Fellowship in support of R.D.H. E.E.E. is an HHMI investigator, M.P. is an HHMI Early Career Scientist, and D.M.A. is Distinguished Clinical Scholar of the Doris Duke Charitable Foundation.
Author information
Authors and Affiliations
Consortia
Contributions
Details of author contributions can be found in the author list.
Corresponding author
Ethics declarations
Competing interests
A.C. is on the Scientific Advisory Board of Affymetrix, Inc.; E.E.E. is a member of the Scientific Advisory Board for Pacific Biosciences; A.L.M. advises Ion Torrents Systems; M.S. is a member of the Scientific Advisory Boards of DNANexus and GenapSis; M.B., D.R.B., R.K.C., T.C., M.E., N.G., S.H., T.J., S.K., Z.K. P.K.-G., L.M., J.S. and M.P.S. work for Illumina; G.L.C., F.M.D.L.V., Y.F., F.C.L.H., J.K.I., C.C.L., J.M.M., K.J.M., S.F.M., H.E.P., O.S., Y.A.S. and E.F.T. work for Life Technologies; J.A., D.A., S.A., M.B., E.B., A.B., A.C., C.C., S.C., D.C., B.D., M.E., L.G., L.G., K.K., A.K., J.K., J.K., M.L., L.M., C.M., M.M., A.N., F.N., K.P., R.R., D.R., W.S., C.T., S.W. and R.W. work for Roche Applied Science.
Supplementary information
Supplementary Information
This file contains Supplementary Text 1-16 (see contents list for details), additional references and Supplementary Figures 1-16 with legends and references. Supplementary Information section 7.7 was corrected on 05 May 2011. (PDF 4175 kb)
Supplementary Tables
This file contains Supplementary Tables 1-13 (XLS 414 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010). https://doi.org/10.1038/nature09534
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09534
This article is cited by
-
Periodontitis and Sjogren’s syndrome: a bidirectional two-sample mendelian randomization study
BMC Oral Health (2024)
-
No genetic causal association between periodontitis and ankylosing spondylitis: a bidirectional two-sample mendelian randomization analysis
BMC Medical Genomics (2024)
-
SGLT1 and SGLT2 inhibition, circulating metabolites, and cerebral small vessel disease: a mediation Mendelian Randomization study
Cardiovascular Diabetology (2024)
-
Polygenic Risk Scores for Breast Cancer
Current Breast Cancer Reports (2024)
-
Genetic evidence for T-wave area from 12-lead electrocardiograms to monitor cardiovascular diseases in patients taking diabetes medications
Human Genetics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.