Abstract
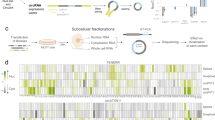
Small (<200 nucleotide) RNA (sRNA) profiling of human cells using various technologies demonstrates unexpected complexity of sRNAs with hundreds of thousands of sRNA species present1,2,3,4. Genetic and in vitro studies show that these RNAs are not merely degradation products of longer transcripts but could indeed have a function1,2,5. Furthermore, profiling of RNAs, including the sRNAs, can reveal not only novel transcripts, but also make clear predictions about the existence and properties of novel biochemical pathways operating in a cell. For example, sRNA profiling in human cells indicated the existence of an unknown capping mechanism operating on cleaved RNA2, a biochemical component of which was later identified6. Here we show that human cells contain a novel type of sRNA that has non-genomically encoded 5′ poly(U) tails. The presence of these RNAs at the termini of genes, specifically at the very 3′ ends of known mRNAs, strongly argues for the presence of a yet uncharacterized endogenous biochemical pathway in cells that can copy RNA. We show that this pathway can operate on multiple genes, with specific enrichment towards transcript-encoding components of the translational machinery. Finally, we show that genes are also flanked by sense, 3′ polyadenylated sRNAs that are likely to be capped.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Data deposits
Sequencing datasets described in this study have been deposited at the National Center for Biotechnology Information (NCBI) Short Read Archive (SRA), accession no SRA012676.
References
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007)
Affymetrix/Cold Spring Harbor Laboratory ENCODE Transcriptome Project. Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457, 1028–1032 (2009)
Taft, R. J. et al. Tiny RNAs associated with transcription start sites in animals. Nature Genet. 41, 572–578 (2009)
Seila, A. C. et al. Divergent transcription from active promoters. Science 322, 1849–1851 (2008)
Rassoulzadegan, M. et al. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 441, 469–474 (2006)
Otsuka, Y., Kedersha, N. L. & Schoenberg, D. R. Identification of a cytoplasmic complex that adds a cap onto 5′-monophosphate RNA. Mol. Cell. Biol. 29, 2155–2167 (2009)
Mardis, E. R. The impact of next-generation sequencing technology on genetics. Trends Genet. 24, 133–141 (2008)
Harris, T. D. et al. Single-molecule DNA sequencing of a viral genome. Science 320, 106–109 (2008)
Aspegren, A., Hinas, A., Larsson, P., Larsson, A. & Soderbom, F. Novel non-coding RNAs in Dictyostelium discoideum and their expression during development. Nucleic Acids Res. 32, 4646–4656 (2004)
Lipson, D. et al. Quantification of the yeast transcriptome by single-molecule sequencing. Nature Biotechnol. 27, 652–658 (2009)
Kim, S. W. et al. A sensitive non-radioactive northern blot method to detect small RNAs. Nucleic Acids Res. 38, e98 (2010)
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nature Genet. 38, 626–635 (2006)
Fodor, E., Mikulasova, A., Mingay, L. J., Poon, L. L. & Brownlee, G. G. Messenger RNAs that are not synthesized by RNA polymerase II can be 3′ end cleaved and polyadenylated. EMBO Rep. 1, 513–518 (2000)
Volloch, V., Schweitzer, B. & Rits, S. Antisense globin RNA in mouse erythroid tissues: structure, origin, and possible function. Proc. Natl Acad. Sci. USA 93, 2476–2481 (1996)
Maida, Y. et al. An RNA-dependent RNA polymerase formed by TERT and the RMRP RNA. Nature 461, 230–235 (2009)
Lai, M. M. RNA replication without RNA-dependent RNA polymerase: surprises from hepatitis delta virus. J. Virol. 79, 7951–7958 (2005)
Lehmann, E., Brueckner, F. & Cramer, P. Molecular basis of RNA-dependent RNA polymerase II activity. Nature 450, 445–449 (2007)
Ahlquist, P. RNA-dependent RNA polymerases, viruses, and RNA silencing. Science 296, 1270–1273 (2002)
Giladi, E. et al. Error tolerant indexing and alignment of short reads with covering template families. J. Comput. Biol. (in the press)
Acknowledgements
We wish to thank A. Willingham, J. Thompson and Z. Li for discussions and help in the preparation of the manuscript. S.E.A. is supported by the Swiss National Science Foundation, B.J. by the NIH (GM079756) and American Cancer Society (RSG0905401), and A.P.M. by the NIH (MH60774). S.F. acknowledges support by a grant from the Comunidad de Madrid and European Union (Exp. 11/2009).
Author information
Authors and Affiliations
Contributions
P.K., F.O. and P.M.M. designed the study, performed the experiments and the data analysis. S.F. and D.L. performed additional bioinformatics analyses. C.H. and S.R. assisted with the development of the Helicos analysis pipeline. S.W.K., A.P.M. and B.J. performed the northern blot and RPA validation experiments. C.B. and S.E.A. contributed to the validation experiments.
Corresponding authors
Ethics declarations
Competing interests
P.K., F.O., D.L., C.H. and P.M.M. are employees and stockholders of Helicos BioSciences Corporation. S.F. is an employee of Integromics, S.L.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary Methods and Materials, References, Supplementary Tables S1 –S2, Supplementary Figures S1-S8 and Supplementary Data for Human Expressed Sequence Tags. (PDF 2599 kb)
Rights and permissions
About this article
Cite this article
Kapranov, P., Ozsolak, F., Kim, S. et al. New class of gene-termini-associated human RNAs suggests a novel RNA copying mechanism. Nature 466, 642–646 (2010). https://doi.org/10.1038/nature09190
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature09190
This article is cited by
-
Strand-specific RNA-Seq reveals widespread and developmentally regulated transcription of natural antisense transcripts in Plasmodium falciparum
BMC Genomics (2014)
-
Gene regulation by antisense transcription
Nature Reviews Genetics (2013)
-
MicroRNA profiling: approaches and considerations
Nature Reviews Genetics (2012)
-
Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease
Nature Reviews Neuroscience (2012)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.