Abstract
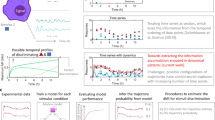
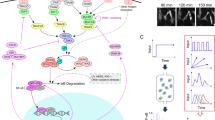
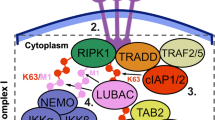
Cells operate in dynamic environments using extraordinary communication capabilities that emerge from the interactions of genetic circuitry. The mammalian immune response is a striking example of the coordination of different cell types1. Cell-to-cell communication is primarily mediated by signalling molecules that form spatiotemporal concentration gradients, requiring cells to respond to a wide range of signal intensities2. Here we use high-throughput microfluidic cell culture3 and fluorescence microscopy, quantitative gene expression analysis and mathematical modelling to investigate how single mammalian cells respond to different concentrations of the signalling molecule tumour-necrosis factor (TNF)-α, and relay information to the gene expression programs by means of the transcription factor nuclear factor (NF)-κB. We measured NF-κB activity in thousands of live cells under TNF-α doses covering four orders of magnitude. We find, in contrast to population-level studies with bulk assays2, that the activation is heterogeneous and is a digital process at the single-cell level with fewer cells responding at lower doses. Cells also encode a subtle set of analogue parameters to modulate the outcome; these parameters include NF-κB peak intensity, response time and number of oscillations. We developed a stochastic mathematical model that reproduces both the digital and analogue dynamics as well as most gene expression profiles at all measured conditions, constituting a broadly applicable model for TNF-α-induced NF-κB signalling in various types of cells. These results highlight the value of high-throughput quantitative measurements with single-cell resolution in understanding how biological systems operate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hayden, M. S., West, A. P. & Ghosh, S. NF-κB and the immune response. Oncogene 25, 6758–6780 (2006)
Cheong, R. et al. Transient IκB kinase activity mediates temporal NF-κB dynamics in response to a wide range of tumor necrosis factor-α doses. J. Biol. Chem. 281, 2945–2950 (2006)
Gómez-Sjöberg, R., Leyrat, A. A., Pirone, D. M., Chen, C. S. & Quake, S. R. Versatile, fully automated, microfluidic cell culture system. Anal. Chem. 79, 8557–8563 (2007)
Batchelor, E., Loewer, A. & Lahav, G. The ups and downs of p53: understanding protein dynamics in single cells. Nature Rev. Cancer 9, 371–377 (2009)
Spencer, S. L., Gaudet, S., Albeck, J. G., Burke, J. M. & Sorger, P. K. Non-genetic origins of cell-to-cell variability in TRAIL-induced apoptosis. Nature 459, 428–432 (2009)
Lahav, G. et al. Dynamics of the p53-mdm2 feedback loop in individual cells. Nature Genet. 36, 147–150 (2004)
Covert, M. W., Leung, T. H., Gaston, J. E. & Baltimore, D. Achieving stability of lipopolysaccharide-induced NF-κB activation. Science 309, 1854–1857 (2005)
Lee, T. K. et al. A noisy paracrine signal determines the cellular NF-κB response to LPS. Sci. Signal. 2, 93 (2009)
Cohen, A. A. et al. Dynamic proteomics of individual cancer cells in response to a drug. Science 322, 1511–1516 (2008)
Hoffmann, A. & Baltimore, D. Circuitry of nuclear factor κB signaling. Immunol. Rev. 210, 171–186 (2006)
Courtois, G. & Gilmore, T. D. Mutations in the NF-κB signaling pathway: implications for human disease. Oncogene 25, 6831–6843 (2006)
Nelson, D. E. et al. Oscillations in NF-κB signaling control the dynamics of gene expression. Science 306, 704–708 (2004)
Ashall, L. et al. Pulsatile stimulation determines timing and specificity of NF-κB-dependent transcription. Science 324, 242–246 (2009)
St, Pierre, F. & Endy, D. Determination of cell-fate selection during phage lambda infection. Proc. Natl Acad. Sci. USA 105, 20705–20710 (2008)
Snijder, B. et al. Population context determines cell-to-cell variability in endocytosis and virus infection. Nature 461, 520–523 (2009)
Elowitz, M., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Hoffmann, A., Levchenko, A., Scott, M. L. & Baltimore, D. The IκB-NF-κB signaling module: temporal control and selective gene activation. Science 298, 1241–1245 (2002)
Hao, S. & Baltimore, D. The stability of mRNA influences the temporal order of the induction of genes encoding inflammatory molecules. Nature Immunol. 10, 281–288 (2009)
Giorgetti, L. et al. Noncooperative interactions between transcription factors and clustered DNA binding sites enable graded transcriptional responses to environmental inputs. Mol. Cell 37, 418–428 (2010)
Bhat, S., Hermann, J., Armishaw, P., Corbisier, P. & Emslie, K. R. Single molecule detection in nanofluidic digital array allows accurate measurement of DNA copy number. Anal. Bioanal. Chem. 394, 457–467 (2009)
Wilson, J. W., Catherine, B. & Christopher, S. (eds) in Apoptosis Genes (Springer, 1999)
Lee, E. G. et al. Failure to regulate TNF-α-induced NF-κB and cell death responses in A20-deficient mice. Science 289, 2350–2354 (2000)
Hutti, J. E. et al. IκB kinase beta phosphorylates the K63 deubiquitinase A20 to cause feedback inhibition of the NF-κB pathway. Mol. Cell. Biol. 27, 7451–7461 (2007)
Lipniacki, T., Paszek, P., Brasier, A. R., Luxon, B. & Kimmel, M. Mathematical model of NF-κB regulatory module. J. Theor. Biol. 228, 195–215 (2004)
Lipniacki, T., Puszynski, K., Paszek, P., Brasier, A. R. & Kimmel, M. Single TNF-α trimers mediating NF-κB activation: Stochastic robustness of NF-κB signaling. BMC Bioinformatics 8, 376 (2007)
Delhase, M., Hayakawa, M., Chen, Y. & Karin, M. Positive and negative regulation of IκB kinase activity through IKK subunit phosphorylation. Science 284, 309–313 (1998)
Chen, Y.-M. et al. Dual regulation of TNF-α induced CCL2/monocyte chemoattractant protein-1 expression in vascular smooth muscle cells by NF-κB and AP-1: modulation by type III phosphodiesterase inhibition. J. Pharmacol. Exp. Ther. 103, 06262 (2004)
Toepke, M. W. & Beebe, D. J. PDMS absorption of small molecules and consequences in microfluidic applications. Lab Chip 6, 1484–1486 (2006)
Acknowledgements
We thank A. Leyrat and R. Gomez-Sjoberg for development of the automated cell culture system, assistance with the software, and for contributions to experimental design and preliminary data. This research was supported in part by an NIH Director’s Pioneer Award (to S.R.Q.), an NCI Pathway to Independence Award (K99CA125994) (to M.W.C.), the Foundation for Polish Science (TEAM 2009-3/6) and NSF/NIH grant no. R01-GM086885 (to T.L.), a Stanford Graduate Fellowship (to T.K.L.), and a Stanford Bio-X Graduate Fellowship (to J.J.H.).
Author information
Authors and Affiliations
Contributions
S.T. and J.J.H. performed the experiments, S.T. and T.L. developed the mathematical models and performed the simulations, T.K.L. developed the image processing methods and all authors contributed to analysis of the data and to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.R.Q. is a founder, shareholder and consultant for Fluidigm.
Supplementary information
Supplementary Information 1
This file contains Supplementary Figures 1-13 with legends and Supplementary Tables 1-3. (PDF 1436 kb)
Supplementary Information 2
This file contains Supplementary Mathematical Methods and Data, Supplementary Figures M1-M6 with legends, Supplementary Tables 1-3 and References. (PDF 3192 kb)
Supplementary Movie 1
This file contains a time-lapse fluorescent microscopy video of a single microfluidic chamber during stimulation with 10 ng/ml TNF-α, showing p65-DsRed nuclear localization oscillations in single-cells. (AVI 10231 kb)
Supplementary Movie 2
This file contains a time-lapse fluorescent microscopy video of a single microfluidic chamber during stimulation with 0.1 ng/ml TNF-α, showing p65-DsRed nuclear localization oscillations in single-cells. (AVI 6166 kb)
Supplementary Information 3
This zipped file contains the stochastic and deterministic model files. All code was written in Matlab. Deterministic model files are denoted with D, i.e. ModelD.m. The files include the following: (1) Main file that calls appropriate modeling functions (Mainfile.m, MainfileD.m). Start here to perform simulations, as this file calls all the other functions in the model. This file contains some commonly changed parameters such as the TNF-α dose; (2) Parameters file (Parameters.m, ParametersD.m). This file contains most of the biochemical parameters used in the simulations; (3) Model equations (Model.m, ModelD.m). ODE's corresponding to biochemical reactions; (4) Status change file (StatusChange.m): Calculates receptor and gene binding propensities in the stochastic version of the model; (5) Plotting files: AllCellPlotting.m and AllCellPlottingD.m plot individual cell traces and related parameters, while AverageCellPlotting.m and AverageCellPlottingD.m plot the population averages of the same simulated cells. (ZIP 17 kb)
Rights and permissions
About this article
Cite this article
Tay, S., Hughey, J., Lee, T. et al. Single-cell NF-κB dynamics reveal digital activation and analogue information processing. Nature 466, 267–271 (2010). https://doi.org/10.1038/nature09145
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09145
This article is cited by
-
The population context is a driver of the heterogeneous response of epithelial cells to interferons
Molecular Systems Biology (2024)
-
Processing stimulus dynamics by the NF-κB network in single cells
Experimental & Molecular Medicine (2023)
-
Modulation of transcription factor dynamics allows versatile information transmission
Scientific Reports (2023)
-
The effect of Alnus incana (L.) Moench extracts in ameliorating iron overload-induced hepatotoxicity in male albino rats
Scientific Reports (2023)
-
Simplifying biochemical signal transport in steady flows within a single-cell-trapping microchannel by linear low-order systems
Microfluidics and Nanofluidics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.